Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G26710 84 / 2e-19
CYP72B1, CYP734A1, BAS1
PHYB ACTIVATION TAGGED SUPPRESSOR 1, Cytochrome P450 superfamily protein (.1)
AT1G67110 65 / 1e-12
CYP735A2
"cytochrome P450, family 735, subfamily A, polypeptide 2", cytochrome P450, family 735, subfamily A, polypeptide 2 (.1)
AT5G38450 64 / 2e-12
CYP735A1
"cytochrome P450, family 735, subfamily A, polypeptide 1", cytochrome P450, family 735, subfamily A, polypeptide 1 (.1)
AT1G75130 64 / 2e-12
CYP721A1
"cytochrome P450, family 721, subfamily A, polypeptide 1", cytochrome P450, family 721, subfamily A, polypeptide 1 (.1)
AT1G17060 58 / 2e-10
SHK1, CHI2, SOB7, CYP72C1
SUPPRESSOR OF PHYB-4 7, SHRINK 1, CHIBI 2, cytochrome p450 72c1 (.1)
AT2G46950 58 / 2e-10
CYP709B2
"cytochrome P450, family 709, subfamily B, polypeptide 2", cytochrome P450, family 709, subfamily B, polypeptide 2 (.1)
AT3G14640 57 / 4e-10
CYP72A10
"cytochrome P450, family 72, subfamily A, polypeptide 10", cytochrome P450, family 72, subfamily A, polypeptide 10 (.1)
AT4G27710 56 / 8e-10
CYP709B3
"cytochrome P450, family 709, subfamily B, polypeptide 3", cytochrome P450, family 709, subfamily B, polypeptide 3 (.1)
AT3G14660 54 / 4e-09
CYP72A13
"cytochrome P450, family 72, subfamily A, polypeptide 13", cytochrome P450, family 72, subfamily A, polypeptide 13 (.1)
AT3G14690 53 / 1e-08
CYP72A15
"cytochrome P450, family 72, subfamily A, polypeptide 15", cytochrome P450, family 72, subfamily A, polypeptide 15 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10036822
268 / 8e-88
AT2G26710 410 / 4e-137
PHYB ACTIVATION TAGGED SUPPRESSOR 1, Cytochrome P450 superfamily protein (.1)
Lus10004633
137 / 1e-38
AT2G26710 378 / 4e-126
PHYB ACTIVATION TAGGED SUPPRESSOR 1, Cytochrome P450 superfamily protein (.1)
Lus10036820
110 / 8e-29
AT2G26710 397 / 2e-133
PHYB ACTIVATION TAGGED SUPPRESSOR 1, Cytochrome P450 superfamily protein (.1)
Lus10036942
108 / 8e-28
AT2G26710 375 / 2e-119
PHYB ACTIVATION TAGGED SUPPRESSOR 1, Cytochrome P450 superfamily protein (.1)
Lus10006245
104 / 2e-26
AT2G26710 345 / 4e-108
PHYB ACTIVATION TAGGED SUPPRESSOR 1, Cytochrome P450 superfamily protein (.1)
Lus10033044
82 / 1e-18
AT2G26710 803 / 0.0
PHYB ACTIVATION TAGGED SUPPRESSOR 1, Cytochrome P450 superfamily protein (.1)
Lus10019184
80 / 7e-18
AT3G14680 369 / 2e-122
"cytochrome P450, family 72, subfamily A, polypeptide 14", cytochrome P450, family 72, subfamily A, polypeptide 14 (.1)
Lus10017771
79 / 1e-17
AT2G26710 705 / 0.0
PHYB ACTIVATION TAGGED SUPPRESSOR 1, Cytochrome P450 superfamily protein (.1)
Lus10036817
77 / 8e-17
AT2G26710 382 / 3e-121
PHYB ACTIVATION TAGGED SUPPRESSOR 1, Cytochrome P450 superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G139300
112 / 1e-29
AT2G26710 372 / 1e-123
PHYB ACTIVATION TAGGED SUPPRESSOR 1, Cytochrome P450 superfamily protein (.1)
Potri.010G139200
103 / 2e-26
AT2G26710 369 / 6e-123
PHYB ACTIVATION TAGGED SUPPRESSOR 1, Cytochrome P450 superfamily protein (.1)
Potri.019G014407
102 / 1e-25
AT2G26710 389 / 3e-130
PHYB ACTIVATION TAGGED SUPPRESSOR 1, Cytochrome P450 superfamily protein (.1)
Potri.017G000600
100 / 1e-25
AT1G17060 166 / 1e-46
SUPPRESSOR OF PHYB-4 7, SHRINK 1, CHIBI 2, cytochrome p450 72c1 (.1)
Potri.010G139400
101 / 2e-25
AT2G26710 402 / 2e-135
PHYB ACTIVATION TAGGED SUPPRESSOR 1, Cytochrome P450 superfamily protein (.1)
Potri.010G139500
98 / 1e-24
AT3G14620 283 / 4e-90
"cytochrome P450, family 72, subfamily A, polypeptide 8", cytochrome P450, family 72, subfamily A, polypeptide 8 (.1)
Potri.019G071200
99 / 2e-24
AT2G26710 346 / 9e-114
PHYB ACTIVATION TAGGED SUPPRESSOR 1, Cytochrome P450 superfamily protein (.1)
Potri.010G139600
90 / 2e-21
AT2G26710 395 / 6e-133
PHYB ACTIVATION TAGGED SUPPRESSOR 1, Cytochrome P450 superfamily protein (.1)
Potri.018G070900
88 / 6e-21
AT2G26710 796 / 0.0
PHYB ACTIVATION TAGGED SUPPRESSOR 1, Cytochrome P450 superfamily protein (.1)
Potri.006G154500
87 / 3e-20
AT2G26710 789 / 0.0
PHYB ACTIVATION TAGGED SUPPRESSOR 1, Cytochrome P450 superfamily protein (.1)
Representative CDS sequence
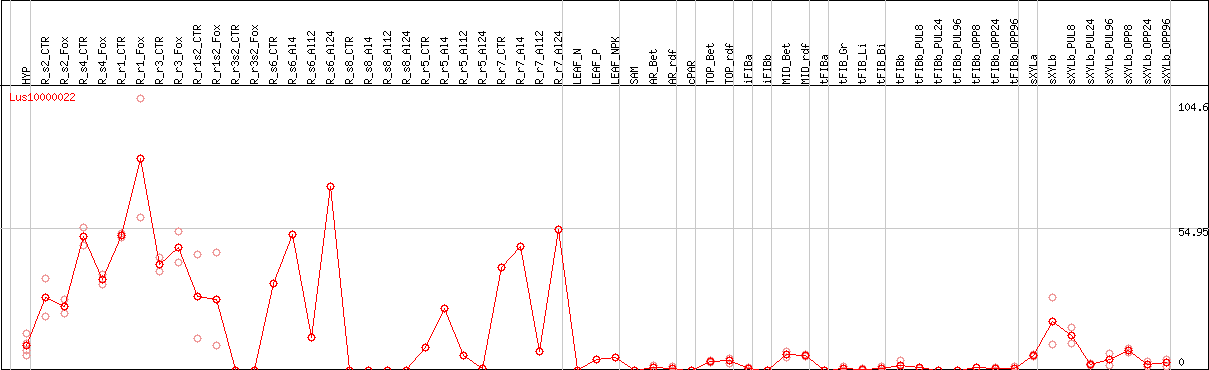
>Lus10000022 pacid=23155429 polypeptide=Lus10000022 locus=Lus10000022.g ID=Lus10000022.BGIv1.0 annot-version=v1.0
ATGGCCAACTTGACATATAATATTATGATGCTCAACATCACTCTCAAAAGTCTTCTTTATTTAATTCCACTCAACGGAAGAGATAAACTAGGTTCTACTG
ATGCGAAAGTTAGAAGAAGAAGTAGATCTAAAGAAAACCTTGAGATGGGAGACCTAATTATGATGCAACTGTTTTTCCTTACCTTGCTTTCTGTTGTCCT
TGTTAATATCCTCAGGTTCTTGAATCAAGTATGGCTGAGACCGATCCGGATTCAGTCCGAAATGACTTCTCAGGGGATTAAAGGCCCTCCTTACAGGTTT
CTTCATGGAAACACCAAAGAGATCAATGATATGATAAGTCAAGCTGTTAGCAACCCCATGGAAATTTCTCATCAGATTCTCCCCAGAATCATGCCACATG
TCCATTCCTGGACAAAGATTTACGGTACCCAGTTTCGGATTCTGGTTTTACTTCTATTGTTGCTCAGTTGCAGATAA