External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G60600 91 / 7e-24
(AT)VAP, (AT)VAP, (AT)VAP, (AT)VAP, (AT)VAP, (AT)VAP, (AT)VAP, VAP27-1, VAP27, (AT)VAP, (AT)VAP, (A
VAMP/SYNAPTOBREVIN-ASSOCIATED PROTEIN 27-1, vesicle associated protein (.1.2.3)
AT2G45140 90 / 2e-23
PVA12
plant VAP homolog 12 (.1)
AT4G00170 84 / 5e-21
Plant VAMP (vesicle-associated membrane protein) family protein (.1)
AT2G23830 77 / 2e-19
PapD-like superfamily protein (.1)
AT1G51270 73 / 1e-16
structural molecules;transmembrane receptors;structural molecules (.1.2.3.4)
AT5G47180 66 / 2e-14
Plant VAMP (vesicle-associated membrane protein) family protein (.1), Plant VAMP (vesicle-associated membrane protein) family protein (.2)
AT1G08820 63 / 7e-13
VAP27-2
vamp/synaptobrevin-associated protein 27-2 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10000032
133 / 8e-42
AT2G45140 148 / 7e-46
plant VAP homolog 12 (.1)
Lus10008512
134 / 1e-40
AT2G45140 296 / 7e-102
plant VAP homolog 12 (.1)
Lus10007463
122 / 2e-35
AT3G60600 306 / 4e-105
VAMP/SYNAPTOBREVIN-ASSOCIATED PROTEIN 27-1, vesicle associated protein (.1.2.3)
Lus10028937
124 / 6e-35
AT3G60600 305 / 2e-101
VAMP/SYNAPTOBREVIN-ASSOCIATED PROTEIN 27-1, vesicle associated protein (.1.2.3)
Lus10010360
94 / 6e-25
AT4G00170 311 / 5e-108
Plant VAMP (vesicle-associated membrane protein) family protein (.1)
Lus10015885
87 / 2e-22
AT2G45140 307 / 2e-106
plant VAP homolog 12 (.1)
Lus10036495
87 / 3e-22
AT4G00170 316 / 4e-110
Plant VAMP (vesicle-associated membrane protein) family protein (.1)
Lus10009279
87 / 2e-21
AT4G00060 1034 / 0.0
maternal effect embryo arrest 44, Nucleotidyltransferase family protein (.1)
Lus10011004
68 / 2e-15
AT5G47180 167 / 2e-52
Plant VAMP (vesicle-associated membrane protein) family protein (.1), Plant VAMP (vesicle-associated membrane protein) family protein (.2)
PFAM info
Representative CDS sequence
>Lus10000031 pacid=23149887 polypeptide=Lus10000031 locus=Lus10000031.g ID=Lus10000031.BGIv1.0 annot-version=v1.0
ATGCAAGCGCAGAAGGAAGCTCCTCCGGATATGCAATGCAAGGACAAATTCTTGCTTCAGAGCGTAAAAGCTCCAGAGGGTTCGACTGTAAAGGATATCA
CTGCTGAGATGTTCAATAAGGAGGCTGGACATATAGTTGAGGAAACCAAGTTGAAGGTTGTTTATGTTGCCCCACCTCAACCTCCTTCTCCTGTTCCCGA
AGGTTCAGAGGAAGGATCATCTCCTCGCGGTTCTGTCTCGGACAATGGACATGTAAACGGTGCTGAGTTTGCTGCTGTGAGTGCTAACTAA
AA sequence
>Lus10000031 pacid=23149887 polypeptide=Lus10000031 locus=Lus10000031.g ID=Lus10000031.BGIv1.0 annot-version=v1.0
MQAQKEAPPDMQCKDKFLLQSVKAPEGSTVKDITAEMFNKEAGHIVEETKLKVVYVAPPQPPSPVPEGSEEGSSPRGSVSDNGHVNGAEFAAVSAN
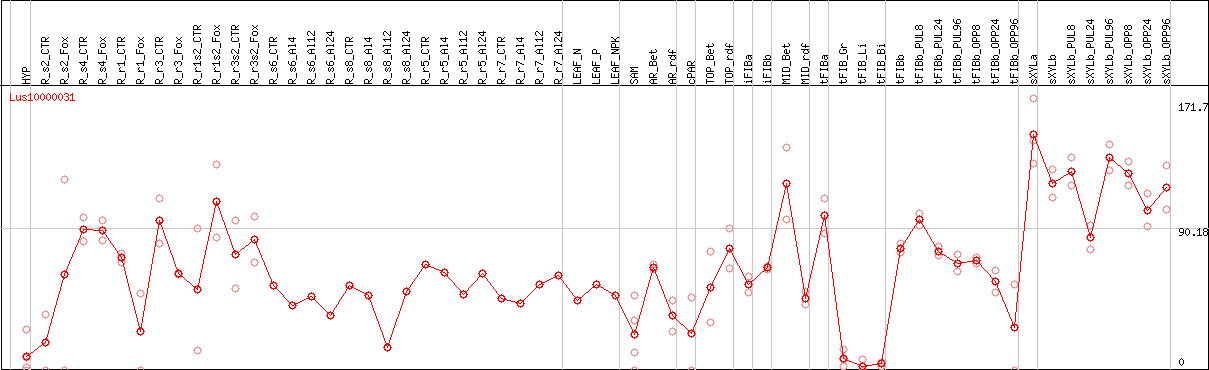
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000031 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.