External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G10720 114 / 5e-31
CKI2, AHK5
CYTOKININ INDEPENDENT 2, histidine kinase 5 (.1)
AT1G27320 51 / 8e-09
AHK3
histidine kinase 3 (.1)
AT5G26594 45 / 6e-07
ARR24
response regulator 24 (.1)
AT2G01830 44 / 2e-06
CRE1, AHK4, WOL1, WOL, ATCRE1
WOODEN LEG 1, WOODEN LEG, CYTOKININ RESPONSE 1, ARABIDOPSIS HISTIDINE KINASE 4, CHASE domain containing histidine kinase protein (.1.2.3)
AT5G35750 43 / 9e-06
AHK2
histidine kinase 2 (.1)
AT2G17820 39 / 0.0001
AHK1, ATHK1
histidine kinase 1 (.1)
AT2G47430 37 / 0.0006
CKI1
CYTOKININ-INDEPENDENT 1, Signal transduction histidine kinase (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G270500
123 / 3e-34
AT5G10720 725 / 0.0
CYTOKININ INDEPENDENT 2, histidine kinase 5 (.1)
Potri.018G011400
122 / 1e-33
AT5G10720 707 / 0.0
CYTOKININ INDEPENDENT 2, histidine kinase 5 (.1)
Potri.003G171000
56 / 3e-10
AT1G27320 1425 / 0.0
histidine kinase 3 (.1)
Potri.010G102900
51 / 8e-09
AT2G01830 1381 / 0.0
WOODEN LEG 1, WOODEN LEG, CYTOKININ RESPONSE 1, ARABIDOPSIS HISTIDINE KINASE 4, CHASE domain containing histidine kinase protein (.1.2.3)
Potri.008G137900
50 / 2e-08
AT2G01830 1390 / 0.0
WOODEN LEG 1, WOODEN LEG, CYTOKININ RESPONSE 1, ARABIDOPSIS HISTIDINE KINASE 4, CHASE domain containing histidine kinase protein (.1.2.3)
Potri.014G121500
48 / 1e-07
AT2G47430 753 / 0.0
CYTOKININ-INDEPENDENT 1, Signal transduction histidine kinase (.1)
Potri.013G009700
48 / 2e-07
AT2G47430 572 / 0.0
CYTOKININ-INDEPENDENT 1, Signal transduction histidine kinase (.1)
Potri.008G191100
47 / 2e-07
AT2G47430 746 / 0.0
CYTOKININ-INDEPENDENT 1, Signal transduction histidine kinase (.1)
Potri.001G057400
46 / 6e-07
AT1G27320 1384 / 0.0
histidine kinase 3 (.1)
Potri.007G056400
45 / 1e-06
AT2G17820 1475 / 0.0
histidine kinase 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0304
CheY
PF00072
Response_reg
Response regulator receiver domain
Representative CDS sequence
>Lus10000137 pacid=23141620 polypeptide=Lus10000137 locus=Lus10000137.g ID=Lus10000137.BGIv1.0 annot-version=v1.0
ATGGTAGCTCGATCCATGATGAAGCAGTTAGGGCACGACATAATTGTGGTGAACAATGGAGTCGAAGCAGTGAAGGCAATACAACGGGAACACTATGATC
TCGTTCTAATGGATGTATATATGCCAGTTATGGATGGCCTTCAAGCTACAAGGATAATCCGTTCCTACGAGGAAACAGGAAGGTGGGATGCAGCTGTCCA
AGCTGGAATTGAGCAATCAGCCACTTGCTTACAAAATGGGCATAGCTTCCATCCCAGTTCCAGAGATCGTATCCCCATAGTCGCGGTAAGTTAG
AA sequence
>Lus10000137 pacid=23141620 polypeptide=Lus10000137 locus=Lus10000137.g ID=Lus10000137.BGIv1.0 annot-version=v1.0
MVARSMMKQLGHDIIVVNNGVEAVKAIQREHYDLVLMDVYMPVMDGLQATRIIRSYEETGRWDAAVQAGIEQSATCLQNGHSFHPSSRDRIPIVAVS
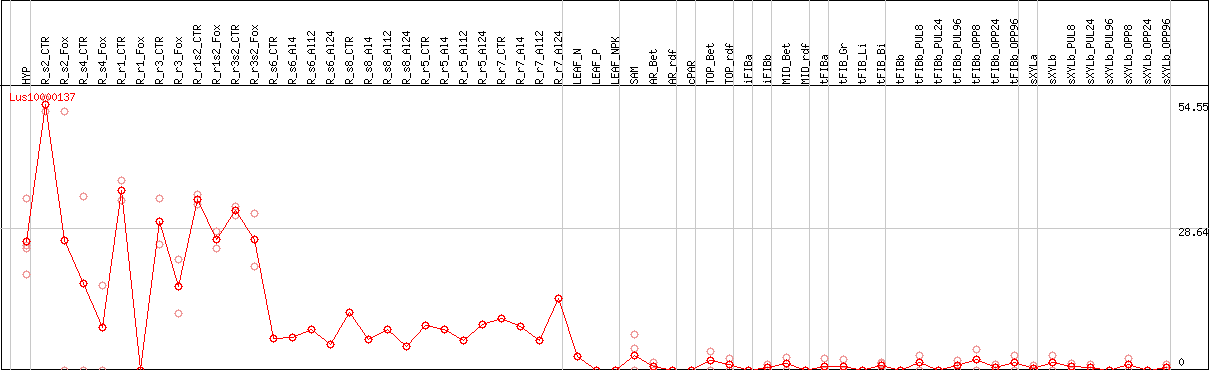
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000137 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.