External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G04150 386 / 4e-127
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
AT5G12970 375 / 3e-125
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
AT1G51570 373 / 2e-124
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
AT3G57880 371 / 2e-123
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
AT4G11610 360 / 6e-117
NTRB, ATNTRB
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
AT5G06850 352 / 4e-116
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
AT5G48060 349 / 1e-112
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
AT1G22610 331 / 8e-106
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
AT4G00700 323 / 4e-103
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
AT3G61300 316 / 9e-101
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10011271
555 / 0
AT1G04150 1187 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Lus10012026
377 / 1e-125
AT3G57880 1496 / 0.0
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
Lus10016280
375 / 3e-125
AT3G57880 1499 / 0.0
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
Lus10037479
365 / 3e-124
AT5G48060 866 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Lus10021030
371 / 3e-123
AT5G06850 1383 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Lus10023823
366 / 2e-121
AT5G06850 1382 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Lus10000605
361 / 2e-117
AT4G11610 1634 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Lus10018839
345 / 3e-111
AT4G11610 1348 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Lus10001538
333 / 3e-106
AT1G22610 1425 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G254400
443 / 6e-149
AT1G04150 1096 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Potri.016G049100
372 / 4e-124
AT3G57880 1425 / 0.0
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
Potri.001G015700
368 / 2e-122
AT5G12970 1295 / 0.0
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
Potri.006G058900
368 / 2e-122
AT3G57880 1372 / 0.0
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
Potri.003G210801
367 / 3e-122
AT5G12970 1300 / 0.0
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
Potri.T085601
367 / 4e-122
AT5G12970 1315 / 0.0
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
Potri.006G058700
366 / 1e-121
AT5G06850 1321 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Potri.016G049300
362 / 8e-120
AT5G06850 1355 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Potri.009G065600
360 / 5e-119
AT5G06850 1308 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Potri.001G271400
363 / 4e-118
AT5G48060 1481 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0484
Peroxisome
PF08372
PRT_C
Plant phosphoribosyltransferase C-terminal
Representative CDS sequence
>Lus10000164 pacid=23143549 polypeptide=Lus10000164 locus=Lus10000164.g ID=Lus10000164.BGIv1.0 annot-version=v1.0
ATGCTGCATCCGACAGGGGTGAAGAAAATGGGAGAGATTCATTTGGCAGTGAGGTTCTCTTGTGCTAATGTGGCCAATATGTTGAACATGTACACTCTCC
CACTACTACCAAAGATGCACTACGTACATCCTTTGAGTGTGAACCAAGTCGAGACATTGAGGTATCAGGCGATGAATGTGTTGTCCTCGAGGCTGAGTAG
GGCCGAGCCACCGTTGGGGAGGGAGGTGGTGGAGTACATGATTGATCATGATTCTCATATGTGGAGCATGAGGAGGAGCAAAGCCAATTTCGCAAGGGTG
ATCAATGTACTCTCCGGTCTAGTTGCATTTGGAAGAATGGTGGAGGCAATGAGGAGTTGGCACAAACCAGTATACTCAGTCCTCTTCCTAGTAGTATTCT
TGTTTCTTGTACTTATGCCGGAGCTCATCATGCCATTGATGCTCGTAATGATGGCGGTAGTGGGACTATGGAGATACCATTCGTCAAGGCCACGCCATCC
GCCTCACATGGATACAAGGCTGTCCAATGCAGAGAATGTAAATCCGGACGAGCTAGATGAAGAGTTCGACTCGTTTCCAACGAGTAGGAACGCTGACATT
GTGCGGATGAGGTATGATAGGGTGAGGAGTGTGGCGGGAAGGATACAGACATTGGTGGGTGACATGGCAACTCAAGGGGAGAGATTTTATGCGCTGGTGA
GCTGGAGGGATCCGAGGGCAACGTTCCTGTTTGTGGCGGGTTGCCTTGTGGCTGCTACTGGGTTGTATGGGGTGCCTGTGAGAGTTGTGGTGGGTGTGTG
GGGGTTGTACATGATGAGGCCTCCCAGGTTCAGGAATAAACTGCCGTCCAAGGCTTCCTGCTTCTTCAAGAGGTTGCCTACTAAGGCTGATAGCTTGCTC
TAG
AA sequence
>Lus10000164 pacid=23143549 polypeptide=Lus10000164 locus=Lus10000164.g ID=Lus10000164.BGIv1.0 annot-version=v1.0
MLHPTGVKKMGEIHLAVRFSCANVANMLNMYTLPLLPKMHYVHPLSVNQVETLRYQAMNVLSSRLSRAEPPLGREVVEYMIDHDSHMWSMRRSKANFARV
INVLSGLVAFGRMVEAMRSWHKPVYSVLFLVVFLFLVLMPELIMPLMLVMMAVVGLWRYHSSRPRHPPHMDTRLSNAENVNPDELDEEFDSFPTSRNADI
VRMRYDRVRSVAGRIQTLVGDMATQGERFYALVSWRDPRATFLFVAGCLVAATGLYGVPVRVVVGVWGLYMMRPPRFRNKLPSKASCFFKRLPTKADSLL
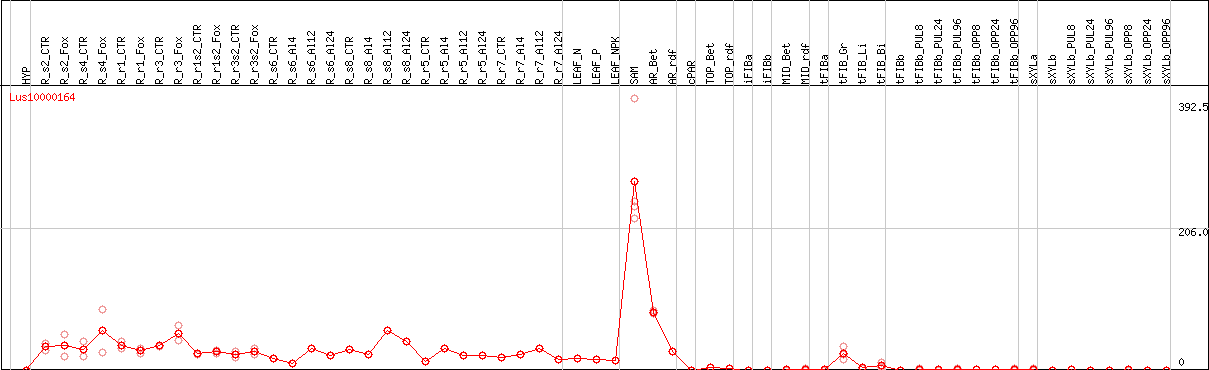
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000164 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.