External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G00050 51 / 1e-07
bHLH
bHLH016, UNE10
unfertilized embryo sac 10, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G67110 47 / 2e-06
bHLH
ALC, bHLH073
ALCATRAZ, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
AT4G36930 47 / 3e-06
bHLH
SPT, bHLH024
SPATULA, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT2G24260 45 / 9e-06
bHLH
LRL1, bHLH066
LJRHL1-like 1 (.1)
AT4G30980 44 / 2e-05
bHLH
LRL2, bHLH069
LJRHL1-like 2 (.1)
AT2G20180 44 / 3e-05
bHLH
PIF1, PIL5, bHLH015
PHY-INTERACTING FACTOR 1, phytochrome interacting factor 3-like 5 (.1.2.3)
AT1G09530 44 / 4e-05
bHLH
PIF3, POC1, PAP3
PHOTOCURRENT 1, PHYTOCHROME-ASSOCIATED PROTEIN 3, phytochrome interacting factor 3 (.1.2)
AT5G58010 43 / 6e-05
bHLH
LRL3, bHLH082
LJRHL1-like 3 (.1)
AT4G02590 40 / 0.0005
bHLH
bHLH059, UNE12
unfertilized embryo sac 12, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
AT1G03040 40 / 0.0007
bHLH
bHLH007
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015734
275 / 2e-92
AT4G00050 138 / 6e-37
unfertilized embryo sac 10, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10003471
234 / 9e-77
AT4G00050 129 / 8e-34
unfertilized embryo sac 10, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10028175
53 / 3e-08
AT4G00050 232 / 2e-71
unfertilized embryo sac 10, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10018724
47 / 2e-06
AT4G36930 126 / 2e-34
SPATULA, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10024811
47 / 4e-06
AT4G36930 148 / 3e-41
SPATULA, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10036286
45 / 1e-05
AT2G24260 214 / 2e-67
LJRHL1-like 1 (.1)
Lus10021846
45 / 1e-05
AT2G24260 275 / 3e-90
LJRHL1-like 1 (.1)
Lus10014726
43 / 5e-05
AT4G02590 278 / 8e-96
unfertilized embryo sac 12, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Lus10022833
43 / 8e-05
AT2G20180 310 / 5e-99
PHY-INTERACTING FACTOR 1, phytochrome interacting factor 3-like 5 (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G066500
50 / 3e-07
AT4G00050 303 / 5e-99
unfertilized embryo sac 10, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.002G143300
50 / 3e-07
AT4G00050 278 / 2e-89
unfertilized embryo sac 10, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.002G124400
47 / 2e-06
AT4G36930 154 / 6e-44
SPATULA, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.014G025800
47 / 2e-06
AT4G36930 148 / 1e-41
SPATULA, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.018G109500
46 / 7e-06
AT2G24260 194 / 5e-58
LJRHL1-like 1 (.1)
Potri.006G186600
46 / 7e-06
AT2G24260 198 / 2e-59
LJRHL1-like 1 (.1)
Potri.014G111400
44 / 3e-05
AT2G46970 131 / 3e-33
phytochrome interacting factor 3-like 1 (.1)
Potri.005G001800
44 / 5e-05
AT1G09530 247 / 3e-73
PHOTOCURRENT 1, PHYTOCHROME-ASSOCIATED PROTEIN 3, phytochrome interacting factor 3 (.1.2)
Potri.013G001300
44 / 5e-05
AT1G09530 224 / 6e-65
PHOTOCURRENT 1, PHYTOCHROME-ASSOCIATED PROTEIN 3, phytochrome interacting factor 3 (.1.2)
Potri.002G045400
42 / 0.0001
AT4G02590 295 / 8e-100
unfertilized embryo sac 12, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
PFAM info
Representative CDS sequence
>Lus10000208 pacid=23158593 polypeptide=Lus10000208 locus=Lus10000208.g ID=Lus10000208.BGIv1.0 annot-version=v1.0
ATGAAAGCACTCCAGAAGTTAGTCCCTAATGCAAACAAGGTATTGGGCATCACATACAAAGTTGTCTTCTTTCTGAAACAACCTTTTCAAGGGTTAACTC
AATGTTTCACCTCTGTTGATCAGACGGATAAAGCTTCGATGTTGGACGAGGTGATCGATTACTTGAGGCAACTACGAGCACAAGTTGAAATGATGAACAA
TATTAGAAGCATGAAACCCCCACAACCATCACAAATGATGATGATGGTCCCCAACATTGATCAAATGCCTCTCTTTGGTTTAGGAATGGGAATGAATCCA
TTAGCATCATCATCATCGTCGATTCACATGCGTCATCTTCTACCTCCATCAGCATTCATGACTAATACCACTCCTCCATTTCTTCTTCATCACTCTACTG
CAAATATGAACTACAACAATACTTGTAAGAGCAGTGGAGAAGCTAATTTGTCATCGTTGAATGCAGCACAGTCTTCCATGGATGTAGAGCTTTACTTGAA
GAACATGGCTGCAATGGATCAACAGCAGCAACAAGAAGTCCATAATCGTAGCACAAATGCGATCGGTGCTGCATCTCACCATTCCAGCCATACTATTCAA
AGGGACAAGTAA
AA sequence
>Lus10000208 pacid=23158593 polypeptide=Lus10000208 locus=Lus10000208.g ID=Lus10000208.BGIv1.0 annot-version=v1.0
MKALQKLVPNANKVLGITYKVVFFLKQPFQGLTQCFTSVDQTDKASMLDEVIDYLRQLRAQVEMMNNIRSMKPPQPSQMMMMVPNIDQMPLFGLGMGMNP
LASSSSSIHMRHLLPPSAFMTNTTPPFLLHHSTANMNYNNTCKSSGEANLSSLNAAQSSMDVELYLKNMAAMDQQQQQEVHNRSTNAIGAASHHSSHTIQ
RDK
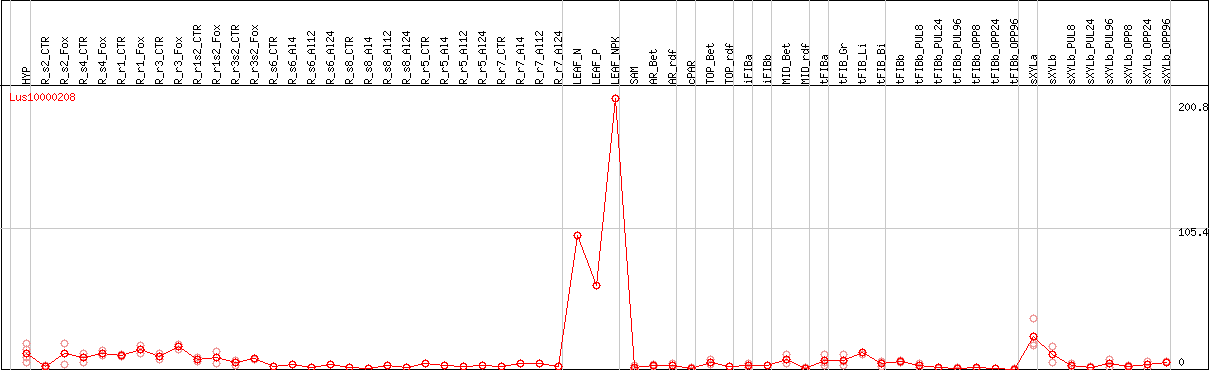
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000208 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.