External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G06450 174 / 1e-51
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT1G61470 164 / 1e-48
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT1G27820 159 / 2e-46
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT3G44240 157 / 2e-46
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT2G32070 156 / 1e-45
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT1G15920 154 / 1e-44
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1.2)
AT1G80780 153 / 2e-44
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1.2.3)
AT5G10960 152 / 6e-44
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT1G27890 152 / 7e-44
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT3G44260 138 / 7e-39
AtCAF1a
CCR4- associated factor 1a, Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10035078
577 / 0
AT1G06450 168 / 2e-49
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10017905
436 / 2e-154
AT1G06450 162 / 5e-48
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10001651
429 / 1e-150
AT1G61470 167 / 4e-49
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10008043
424 / 7e-150
AT1G06450 165 / 5e-48
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10035077
411 / 7e-145
AT1G61470 162 / 6e-48
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10000561
395 / 1e-138
AT1G61470 143 / 1e-40
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10038113
324 / 7e-112
AT3G44240 140 / 3e-41
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10007771
250 / 1e-81
AT1G61470 170 / 5e-51
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10029713
238 / 4e-77
AT5G10960 144 / 4e-41
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G038700
229 / 1e-73
AT1G61470 182 / 8e-56
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.001G046700
159 / 1e-46
AT2G32070 472 / 2e-170
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.003G181100
158 / 2e-46
AT1G80780 468 / 1e-168
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1.2.3)
Potri.018G020900
149 / 7e-43
AT2G32070 441 / 2e-158
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.006G205600
149 / 1e-42
AT5G22250 302 / 4e-103
CCR4- associated factor 1b, Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.016G073000
148 / 1e-42
AT3G44260 322 / 5e-111
CCR4- associated factor 1a, Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.006G262500
145 / 2e-41
AT2G32070 443 / 5e-159
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.009G161500
144 / 3e-41
AT5G22250 387 / 1e-136
CCR4- associated factor 1b, Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.006G187200
144 / 4e-41
AT2G32070 403 / 2e-143
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.004G200400
140 / 1e-39
AT5G22250 419 / 3e-149
CCR4- associated factor 1b, Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0219
RNase_H
PF04857
CAF1
CAF1 family ribonuclease
Representative CDS sequence
>Lus10000327 pacid=23149795 polypeptide=Lus10000327 locus=Lus10000327.g ID=Lus10000327.BGIv1.0 annot-version=v1.0
ATGGGTTTTCTCGTGAGGGAAGTCTGGGACGAAAATCTCCGTCAAGAATTGAAAGCCATGGAAGAATGTCTGGCCAAGTACCCAGTTATATCAGTCGACT
CCGAATTCCCAGGCTTCCTCCGTCAAACGCCGAGATACGCTTCCGAGGAGCTCCGATTTGAGAACTTGAAGTACAACGTCGGCCAAACCTCTCTCGTTCA
ATTAGGGCTCACTCTCTCAAACACAAAAGGCACCGTCGGCGGAATCTGGCAATTCAACTTCCAATTCGATCTAGACAGCGATCTCCACTGCCCAAATTCG
ATCCAATTTTTAATCCTCCACGGCATCGATTTCAACAAGCTCAAAACCTACGGGATTGACCGGGTCAAATTTGGCACGGTGTTCGGAAGGCTGATTGCGA
GAAGGAAGAGTTCGATTACGTGGATTACTTTCCACGGCGTGTACGACCTCGCCCATATTGTGAAGGCGGTGAGTTTCCGTCCGGTTGCGGAATCGTCGGC
GGAATTCGTTGACGTTTTGGGGAGGGGGTTTGACTCGGTTGTTGATGTGAAATACATGGCTAAGTTTTATCAAGGGACAGGGAGTGAAATTGGATTGCAG
AAGTTGGCTGATAATTTAGGGGTTAAGAGAAGAGGGGAAGCTCATACGGCTGCTTCCGACAGTTTGTTGACGGCGCTGGTTTACTTCAAGCTGAAGAAGA
AGCTGAAGCAGCAGGGGATTGAGGAAAACGTTTACTTTGACTTGGTGTATGGAATTAGTTCCAGAATCTCTAGGGAGTTCATCAATGCTACTGCTGCTGC
TGCTGCTCTTGTGTATCAGGGGGATATGGTGTATGCGAATCCGTATTACAAGAGGAATTTGGCTGCCGAATTGATGATGATGGGTCATTGGGATTATTGC
TCCTCCACTACTGGTGCAACACCAGTGTCATCTCTGATCTACGGATGA
AA sequence
>Lus10000327 pacid=23149795 polypeptide=Lus10000327 locus=Lus10000327.g ID=Lus10000327.BGIv1.0 annot-version=v1.0
MGFLVREVWDENLRQELKAMEECLAKYPVISVDSEFPGFLRQTPRYASEELRFENLKYNVGQTSLVQLGLTLSNTKGTVGGIWQFNFQFDLDSDLHCPNS
IQFLILHGIDFNKLKTYGIDRVKFGTVFGRLIARRKSSITWITFHGVYDLAHIVKAVSFRPVAESSAEFVDVLGRGFDSVVDVKYMAKFYQGTGSEIGLQ
KLADNLGVKRRGEAHTAASDSLLTALVYFKLKKKLKQQGIEENVYFDLVYGISSRISREFINATAAAAALVYQGDMVYANPYYKRNLAAELMMMGHWDYC
SSTTGATPVSSLIYG
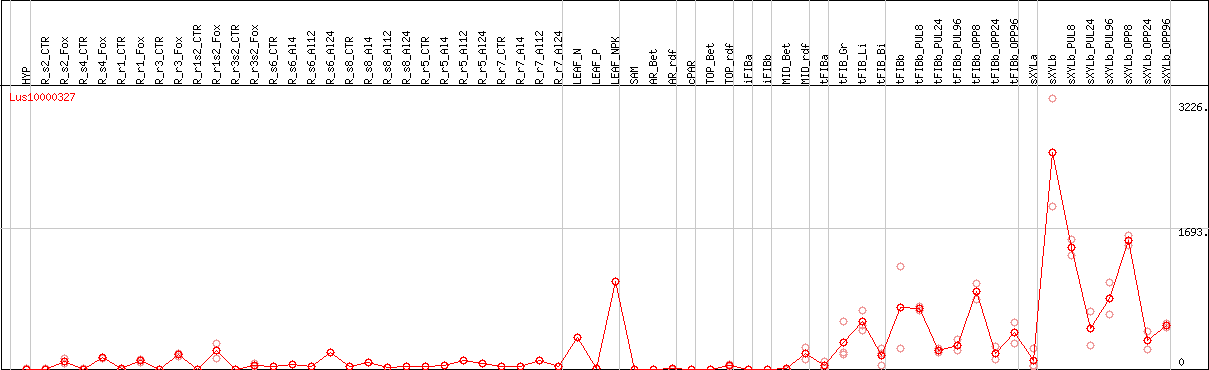
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000327 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.