Lus10000390 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10000390 pacid=23146074 polypeptide=Lus10000390 locus=Lus10000390.g ID=Lus10000390.BGIv1.0 annot-version=v1.0
ATGGCGGGTACGGGCACTATGAGCAATCTCCTTCAACAAGGAGCAAGCTCCTCTTCGTCGTCTACGCTCCCGAAGAAGCCAAGCAGGTCGTTATGTTCGC
CTGTATTCGCCGTCGCGCCGGTGATCCCTCGCCGTGGTGGGACTCGATGCTCCGTCGCGAAGAATTCGGCAGCCGGTTTAGGGAAAAACTTGTTGCGGAC
TCACGGATCCGATAAGAAACTCGTGTGGAAATCTGACGGACCGGGGAGGTCGCCGAAGCTGCGGATGATGGTCAGGTCCGCATTGTCCGGGGTTCCGGAG
AAGCCGCTTGGACTTTACGATCCGTCTTTCGATAAGGACTCTTGTGGGGTGGGATTCGTCGCTGAGCTGTCGGGTGAAAGCAATCGCAAGACGGTAAGCG
ATGCATTGGAGATGTTAGTGCGGATGACGCATAGAGGTGCTTGTGGATGTGAAGCGAATACTGGCGATGGAGCTGGGATTCTCGTTGGTCTTCCTCACGC
CTTCTACAAAGAGGTTGCAAAGGATGCTGGGTTTGTGCTACCTCCACCGGGGGAATATGCAGTTGGCATGTTCTTTTTGCCCACCTTGGATAACCGCAGA
GAGGAAAGCAAGAATGTGTTTACCAAGGTTGCCGAGTCTCTTGGGCATACAGTTCTTGGATGGCGGTCGGTGCCGACAGATAATTCTGGTTTGGGTCAGT
CTGCTGTGCAGACTGAGCCTGTGGTTGAGCAGGTTTTCCTCACACCCACTCCTAGATCGAAAGCTGATTTCGAGCAACAGATGTACATCTTACGGAGGGT
GTCCATGGTAGCTATCCGAGCTGCTCTAAACCTTCAACATGGTGGTGTCCGAGACTTCTATATATGTTCTTTGTCATCTAGGACTGTTGTTTATAAGGGT
CAGCTGAAGCCTGTACAGTTGAGCGAGTACTACTATGCGGATCTTGGGAATGAAAGGTTTACGAGCTACATGGCCCTGGTACATTCTCGCTTTTCGACCA
ATACCTTCCCTAGCTGGGATCGTGCCCAACCTATGCGTGTCTTGGGCCACAATGGTGAAATTAACACTCTTAGGGGCAATGTGAACTGGATGAGGGCACG
CGAAGGTCTTTTGAAATGCAAGGAGCTTGGACTCTCTAAGAATGAGATGAAGAAGCTTTTGCCAATTGTTGATGCTAGTTCATCAGACTCTGGGGCTTTC
GATGGTGTCCTTGAACTTCTGGTTCGAGCTGGTAGAAGTCTCCCTGAAGCTGTCATGATGATGATTCCTGAGGCCTGGCAAAATGACAAAAATATCGACC
CACAAAGGAAAGCTTTGTACGAGTACTTCTCTTGTCTCATGGAGCCATGGGATGGGCCTGCACTTGTCTCATTCACTGATGGCCACTACCTTGGAGCTAC
ACTAGATCGCAATGGCTTGCGTCCAGGCAGGTTCTATGTCACTCGTAGTGGTAGAGTTATCATGGCCAGTGAAGTTGGTGTTGTAGATGTGCCGCCTGAA
GATGTCCTCAGGAAAGGAAGGCTTAACCCTGGAATGATGCTTCTGGTGGATTTTGAGAAGCATATTGTTGTTGATGATGAAGCCTTGAAACAGCAGTACT
CGTTGTCACAGCCTTATGGTGATTGGTTGAAGAGGCAAAAGATTGAACTAAAGGACATAATTGAGACAGTTCCTGAATCTGACCGACTTCCTCCAGGGAT
GGCGGGTGTTGCTCCCGCTTCTGATGATGACGAGAACATGGAGAAAATGGGTGTTCACGGCTTGTTGGCTCCATTGAAAGCTTTTGGCTATACTGTTGAA
GCCTTGGAAATGCTGCTTCTTCCGATGGCAAAAGATGGCAGCGAGGCTCTTGGCTCAATGGGCAATGATACTCCATTGGCTGTGATGTCCGACAGAGAAA
AGCTAACTTTTGAATACTTCAAGCAAATGTTTGCCCAAGTGACAAACCCTCCAATTGACCCCATTAGAGAGAAAATTGTCACTTCAATGGAATGCATGAT
TGGTCCTGAAGGTGATCTGACAGAAACAACTGAAGATCAATGCCACCGCCTCTCACTGAAAGGTCCTCTCCTGACAATTGATGAAATACAGGCTCTTAAA
AAGATGGACTACAGGGGTTGGAGGAGCAAAGTTCTTGATATAACTTATTCAAGGGACCGTGGCAGGAAGGGACTGGAGGAAACGTTGGATAGGATTTGTG
CTGAGGCACATGCTGCTATCAAGGAAGGCTACACCTTGTTGGTTCTTTCTGACAGAGCATTCTCTCCAACAAGAGTTGCTATAAGCTCACTCTTGGCTGT
TGGTGCTGTCCATCATCACCTTGTCAGAAGGCTTGAGCGGACTCAAGTGGGCCTGATAGTGGAGTCTGCTGAACCGCGTGAAGTGCATCATTTTTGTACG
TTGGTTGGATTTGGCGCCGATGCTATATGCCCATACTTAGCTGTAGAAGCTATTTGGAGACTGCAGGTTGATGGAAAAATTCCACCAAAATCTAATGGCG
AGTTCCGCTCCAAGAATGAGTTGGTGAAAAAGTATTTCAAAGCAAGTAATTATGGTATGATGAAGGTTCTTGCCAAAATGGGGATATCAACGTTGGCGTC
ATACAAAGGGGCACAGATTTTCGAAGCCCTTGGCCTTTCATCCGAGGTGATTGATAAATGCTTTGCTGGAACTCCCAGCAGGGTTGAGGGAGCTACGTTT
GAGATGCTTGCCCATGATTATCTTCATTTACACGGGTTGGCATTTCCTACACGAGGTCTTCCTCGTCTTCCGCGTGGATGTGAAGAGGCTGTTGCACTGC
CAAATCCAGGGGATTATCATTGGCGAAAGGGTGGTGAAATCCACTTGAATGATCCTCTTGCTATTGCAAAGCTTCAAGAAGCTGCGAGAGCGAATAGTGT
AGCTGCTTATAAAGAATACTCAAAACGTATCCAGGAATTGAACAAAGCATGTAACTTGAGGGGTATCCTGAAGTTTAAAGACTCTGCCGCTAAGGTTCCG
CTGGAAGAAGTAGAGGCAGCTAGGGGAATTGTTAAACGTTTCTGCACTGGGGCTATGAGTTATGGGTCGATTTCATTGGAGGCTCACACAACACTGGCTC
TTGCGATGAATAAGATTGGAGGGAGATCTAACACAGGTGAGGGAGGCGAGCAGCCATCTCGACTGGTGCCTCTTCCAGATGGTTCCAGGAATCCGAAAAG
GAGTTCAATTAAGCAGGTTGCAAGTGGGCGTTTTGGCGTTTCAAGTTACTATCTTACAAATGCTGATGAGTTGCAGATAAAGATGGCTCAGGGTGCCAAG
CCTGGTGAAGGCGGCGAGCTACCTGGACATAAAGTTATAGGAGACATTGCAGTAACCCGAAATTCCACTGCTGGTGTTGGTCTCATCAGTCCGCCTCCTC
ATCATGATATCTACTCAATTGAAGACCTTGCCCAACTGATTCATGATCTCAAGAATGCCAATCCAGGAGCCCGGATTAGTGTCAAGTTGGTGTCTGAAGC
TGGTGTTGGTGTGATTGCTAGTGGAGTGGTGAAGGGTCATGCTGATCATGTTTTGATCTCTGGTCACGATGGAGGTACTGGAGCTTCTAGATGGACTGGC
ATCAAGAACGCTGGCCTGCCTTGGGAACTTGGTTTGGCCGAGACCCACCAAACTCTTGTTGCTAACAACCTTCGTGGACGGACCGTTCTTCAGACTGATG
GTCAATTGAAAACTGGAAGAGATGTAGCTGTTGCTGCACTTCTTGGTGCAGAAGAGTTTGGCTTCAGCACTGCACCTCTTATAACTCTCGGTTGTATCAT
GATGAGAAAGTGCCATAAAAATACATGCCCCGTTGGCATTGCTACCCAAGATCCAGTTTTGAGAGAGAAGTTTGCTGGGGAACCTGAACATGTCATCAAT
TTCTTATTTATGCTAGCTGAAGAACTTAGGGAAATCATGGCTGGTCTTGGTTTCCGTACTGTTAATGAGATGGTTGGACGTGCGGATATGCTCGAAGTTG
ATAAAGAGGTTGTACGAAACAATGAGAAGTTGGAGAACATTGATCTTTCGCTTTTGCTTAGACCTGCTTCTGACATCCGACCTGAGGCTGCACAGTATTG
TGTTGAGAAGCAGGATCATGGCTTAGACATGGCGCTGGACAACAAACTTATCACACTCTCAACTGCTGCCATAGAAAAGAGTATTCCTGTATACATAGAG
TCTCCTATATGCAATGTGAACCGTGCTGTTGGTACAATGCTTAGTCACGAAGTTACTAAGCGTTACAATTTGGCAGGGCTTCCTAAAGACACAATTCATG
TCAAACTCTTTGGAAGTGCTGGGCAAAGTCTTGGTGCTTTCCTTTGTCCTGGCATAACCCTGGAGCTTGAGGGTGACAGCAATGATTACGTTGGTAAAGG
ATTGTCAGGTGGAAAGATCGTAGTTTATCCACCAAAAGGAAGCCTATTTGATCCAAAAGAAAACATCGTCATTGGTAATGTGGCTCTGTATGGTGCCACC
AGTGGTGAAGCCTATTTTAATGGCATGGCAGCAGAAAGGTTTTGTGTGCGTAATTCAGGGGCAACTGCAGTTGTTGAAGGAGTCGGTGATCATGGATGTG
AGTACATGACAGGCGGAACAGTAGTTGTGCTTGGGAAGACGGGCAGGAATTTTGCTGCAGGCATGAGCGGTGGTATTGCTTATGTTCTTGACGGTGATGG
TAAATTTCATTCTCGTTGCAATCCGGAATTGGTAGATCTTGATAAAGTTGAGGAAGAAGAGGACATCATGCATCTAAGAATGATGATACAGCAGCATCAG
CGCAATACTAACAGCCAGTTAGCTAAAGAAGTGTTGGCTGATTTTGACAATCTCCTTCCGAAGTTCATCAAGGTCTTTCCAAGAGACTACAAGCGTATTC
TTGCAAGCATGAAAAGCCAGTCAGCTTCTCAGGAAGTTGCTGAGAAAATGGTTAAAGAAACAGAGGACCAAGATGAGGTAGAGTTGACTCCGAAAGATGC
TTTTGAGGAGCTCAAGAAATTAGCAGCTGCATCATTGAGTGACAAATCCATTCAGGGGCAAGAAGCTGATGAATCAAAAAGGCCAAGTCGGGTTAGTGAT
GCTGTTAAGCATCGTGGGTTTATTGCTTATGAGCGTGAGGGTGTTCACTACAGAGATCCTAATGTCCGCATGAATGACTATAATGAAGTTATGGAGGAGA
CGAAACCTGGCCCTCTTTTGAACACCCAGTCAGCTCGCTGCATGGATTGTGGTACTCCTTTCTGCCATCAGGAGCACACTGGCTGTCCTCTAGGAAACAA
AATACCTGAGTTCAACGAGCTGGTTTACCAAAATAGATGGCGTGAAGCATTAGACAGGCTGCTTGAGACGAACAACTTCCCGGAATTTACTGGCCGTGTA
TGCCCTGCTCCCTGCGAAGGTTCCTGCGTTCTTGGAATTATTGAAAATCCCGTTTCCATTAAAAGCATAGAATGTGCTATCATTGACAAGGCCTTCGAAG
AGGGGTGGATGAAACCAAGGCCTCCTCTCAATAGAACTGGGAAACGAGTTGCTATCATTGGAAGTGGACCTGCTGGTTTGGCTGCAGCTGATCAGCTAAA
TAGAATGGGTCATTTGGTGACAGTCTACGAGCGTGCTGATAGAATTGGAGGGCTGATGATGTATGGAGTCCCCAACATGAAGACTGACAAAATGGATGTA
GTTCAGCGGCGTGTGAATCTCATGGCAGAGGAAGGCATTGACTTCGTGGTCAATGCTAATGTTGGGGTTGATCCCCAGTATCATCTTGACCGTCTCCGGG
AAAATAATGACGCTGTTATTTTGGCTGTAGGAGCCACCAAACCGAGGGACCTTCCCGTACCTGGACGGGACTTGCAAGGAGTTCATTTTGCCATGGAGTT
CCTTCACGCAAACACAAAGAGTTTGCTCGATAGCAATCTTGAGGATGGCAACTACATATCAGCTAAAGGGAAAAAAGTGGTCGTCATTGGTGGAGGTGAC
ACCGGTACAGACTGCATAGGAACATCCATCCGCCATGGTTGTACCAGCATCGTAAATCTTGAGCTTCTACCTGAGCCGCCAAGGACTAGAGCACCAGGCA
ACCCTTGGCCACAGTGGCCTCGCATCTTCCGGGTAGACTATGGACACCAAGAAGCTGCAACGAAATTCGGCAAAGACCCACGATCTTACCAAGTGTTGAC
CAAACGTTTCGTGGGAGACGAAAGTGGGAAACTCAAGGGGCTTGAGATTGTCCATGTCGAATGGGAGAAGGATGCGAGTGGGAGATTCCAGTTTAAGGAG
GTCGAAGGTTCGGAGGAAATCATTGAGGCTGACCTTGTCATGCTAGCAATGGGATTCCTTGGTCCTGAGGCGAACTTGGCTGAAAAGCTGGGCTTGGAGC
AAGACGGCCGATCAAACTTCAAAGCTGATTACGGCCGATTCTCTACCAACGTGGAAGGAGTGTTTGCTGCTGGGGATTGTCGTCGAGGGCAGTCGCTTGT
CGTGTGGGCAATATCAGAGGGAAGACAAACAGCAGCACAGGTGGACAAGTATTTGATGGGAGAAGAGGATGCCAGCATCGGCACATCGGACGATGATGCC
GAGAGACAGCCAGACAAGACAAAGAAGCAGCAAGACAACAAGCACCGAGTCATGACATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10000390 pacid=23146074 polypeptide=Lus10000390 locus=Lus10000390.g ID=Lus10000390.BGIv1.0 annot-version=v1.0
MAGTGTMSNLLQQGASSSSSSTLPKKPSRSLCSPVFAVAPVIPRRGGTRCSVAKNSAAGLGKNLLRTHGSDKKLVWKSDGPGRSPKLRMMVRSALSGVPE
KPLGLYDPSFDKDSCGVGFVAELSGESNRKTVSDALEMLVRMTHRGACGCEANTGDGAGILVGLPHAFYKEVAKDAGFVLPPPGEYAVGMFFLPTLDNRR
EESKNVFTKVAESLGHTVLGWRSVPTDNSGLGQSAVQTEPVVEQVFLTPTPRSKADFEQQMYILRRVSMVAIRAALNLQHGGVRDFYICSLSSRTVVYKG
QLKPVQLSEYYYADLGNERFTSYMALVHSRFSTNTFPSWDRAQPMRVLGHNGEINTLRGNVNWMRAREGLLKCKELGLSKNEMKKLLPIVDASSSDSGAF
DGVLELLVRAGRSLPEAVMMMIPEAWQNDKNIDPQRKALYEYFSCLMEPWDGPALVSFTDGHYLGATLDRNGLRPGRFYVTRSGRVIMASEVGVVDVPPE
DVLRKGRLNPGMMLLVDFEKHIVVDDEALKQQYSLSQPYGDWLKRQKIELKDIIETVPESDRLPPGMAGVAPASDDDENMEKMGVHGLLAPLKAFGYTVE
ALEMLLLPMAKDGSEALGSMGNDTPLAVMSDREKLTFEYFKQMFAQVTNPPIDPIREKIVTSMECMIGPEGDLTETTEDQCHRLSLKGPLLTIDEIQALK
KMDYRGWRSKVLDITYSRDRGRKGLEETLDRICAEAHAAIKEGYTLLVLSDRAFSPTRVAISSLLAVGAVHHHLVRRLERTQVGLIVESAEPREVHHFCT
LVGFGADAICPYLAVEAIWRLQVDGKIPPKSNGEFRSKNELVKKYFKASNYGMMKVLAKMGISTLASYKGAQIFEALGLSSEVIDKCFAGTPSRVEGATF
EMLAHDYLHLHGLAFPTRGLPRLPRGCEEAVALPNPGDYHWRKGGEIHLNDPLAIAKLQEAARANSVAAYKEYSKRIQELNKACNLRGILKFKDSAAKVP
LEEVEAARGIVKRFCTGAMSYGSISLEAHTTLALAMNKIGGRSNTGEGGEQPSRLVPLPDGSRNPKRSSIKQVASGRFGVSSYYLTNADELQIKMAQGAK
PGEGGELPGHKVIGDIAVTRNSTAGVGLISPPPHHDIYSIEDLAQLIHDLKNANPGARISVKLVSEAGVGVIASGVVKGHADHVLISGHDGGTGASRWTG
IKNAGLPWELGLAETHQTLVANNLRGRTVLQTDGQLKTGRDVAVAALLGAEEFGFSTAPLITLGCIMMRKCHKNTCPVGIATQDPVLREKFAGEPEHVIN
FLFMLAEELREIMAGLGFRTVNEMVGRADMLEVDKEVVRNNEKLENIDLSLLLRPASDIRPEAAQYCVEKQDHGLDMALDNKLITLSTAAIEKSIPVYIE
SPICNVNRAVGTMLSHEVTKRYNLAGLPKDTIHVKLFGSAGQSLGAFLCPGITLELEGDSNDYVGKGLSGGKIVVYPPKGSLFDPKENIVIGNVALYGAT
SGEAYFNGMAAERFCVRNSGATAVVEGVGDHGCEYMTGGTVVVLGKTGRNFAAGMSGGIAYVLDGDGKFHSRCNPELVDLDKVEEEEDIMHLRMMIQQHQ
RNTNSQLAKEVLADFDNLLPKFIKVFPRDYKRILASMKSQSASQEVAEKMVKETEDQDEVELTPKDAFEELKKLAAASLSDKSIQGQEADESKRPSRVSD
AVKHRGFIAYEREGVHYRDPNVRMNDYNEVMEETKPGPLLNTQSARCMDCGTPFCHQEHTGCPLGNKIPEFNELVYQNRWREALDRLLETNNFPEFTGRV
CPAPCEGSCVLGIIENPVSIKSIECAIIDKAFEEGWMKPRPPLNRTGKRVAIIGSGPAGLAAADQLNRMGHLVTVYERADRIGGLMMYGVPNMKTDKMDV
VQRRVNLMAEEGIDFVVNANVGVDPQYHLDRLRENNDAVILAVGATKPRDLPVPGRDLQGVHFAMEFLHANTKSLLDSNLEDGNYISAKGKKVVVIGGGD
TGTDCIGTSIRHGCTSIVNLELLPEPPRTRAPGNPWPQWPRIFRVDYGHQEAATKFGKDPRSYQVLTKRFVGDESGKLKGLEIVHVEWEKDASGRFQFKE
VEGSEEIIEADLVMLAMGFLGPEANLAEKLGLEQDGRSNFKADYGRFSTNVEGVFAAGDCRRGQSLVVWAISEGRQTAAQVDKYLMGEEDASIGTSDDDA
ERQPDKTKKQQDNKHRVMT
|
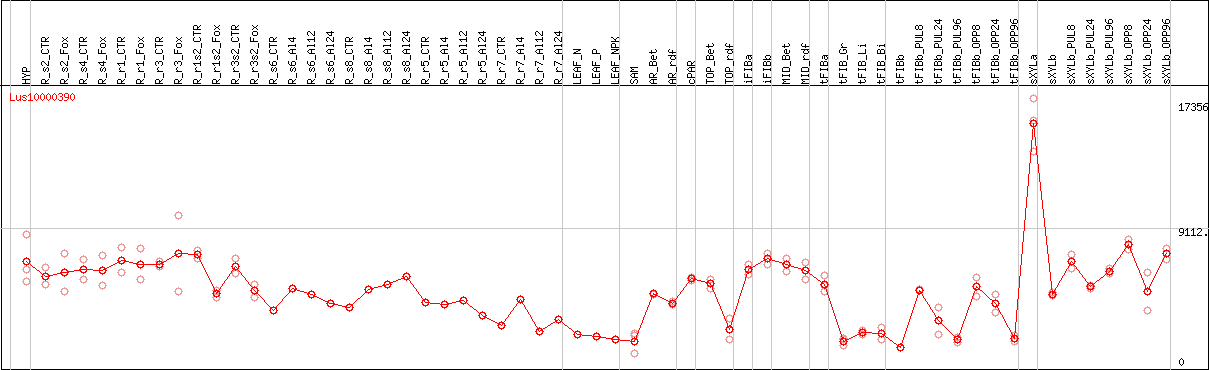
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000390 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
