Lus10000392 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10000392 pacid=23170849 polypeptide=Lus10000392 locus=Lus10000392.g ID=Lus10000392.BGIv1.0 annot-version=v1.0
ATGGATAGCAACACGGAATGTGTCAAACTCGCAGTCAATATTCGTCCTCTCATTACTCCAGAGCTTCTAAATGGATGTACTGACTGTATCACTCTCGTTC
CCGGAGAACCTCAGGTGCAAATTGGGTCGCATTCTTTTACATATGATTATGTATATGGTAGTTTGGATTCCCCGTCTTCAGCCATCTATGATGATTGCGT
TGCTCCTCTGGTTGATGCGTTGTTCCATGGTTACAATGCCACTGTCCTTGCTTACGGCCAGACGGGTTCCGGTAAAACATATACTATGGGTACAAATTAT
ACTGGAGAAGGATCTAATGAAGGTGTTATTCCAAAAGTTATGGCTACCATCTTTAGAAGACTTGAGTCGATAAAACAATCGACTGAGTTCTTGATCAGAG
TGTCATTCATTGAGATATTCAAAGAAGAGGTGTTCGATTTGCTTGACACGAATACGGTAATATCTAAAGGGGAGGGTGGAAATGCCGGAAAGGCGGTTGT
GTCTCCTAGAGTTCCCATTCAAATTAGGGAAACTGTGAATGGAGGCATAACACTTGCCGGTGTGTCCGAAGCAGAAGTTCGTTGTAAAGAAGAAATGGGA
GCATACCTCTCCAAAGGTTCATTGGCGCGTGCTACTGCTAGCACTAACATGAATAAGCAATCAAGTCGGTCCCATGCTATATTTACAATTTCCATGGAGC
AGAAGAAACTTACTCCTTGTTCAAGTGGAGGAAACAATGAGGATATGTGTGACGACATATTACGAGCAAAACTACACCTTGTAGATCTTGCTGGATCTGA
GCGTGCAAAACGAACAGGCGCTGATGGCATGCGTTTCAAAGAAGGCATCCATATCAATAAGGGTTTGTTGGCCCTTGGCAACGTTATCAGTGCATTGGGG
GATGAGAAAAAGCGAAAAGATGGTGGTCACGTCCCATACCGTGATAGCAAGTTAACTCGCTTGTTACAGGATTCACTAGGAGGTAACAGCAAGACCGTGA
TGATCGCATGTGTAAGCCCTGCTGATACCAACGCGGAGGAGACGCTGAACACATTGAAGTACGCAAATCGTGCTCGAAACATCCAGAACAAAGCTGTTAT
CAACCGTGATCCTAATGCCGTTCAACTTCAAAGGATGAGGAGCCAAATTGAACAGTTGCAAGCTGAACTGTGTATTTATCGGGGAGATGCAAATGCACCT
TTTGATGAAATTCAGATTCTTAAACATAAGGTATCTTTACTTGAAGCAAGCAATGCTGAATTACAGAAAGAGCTTCAGGAGCGGCAAATGACTTTTGAGC
ATTTGAATCAACGCGCTGTTGATGCACAGGTTGAAAAGGACAAATTGATAATGCAGATTGAATTAGTTCGAAATGGTAAATCTTGGGATGAGATTGACGC
TGCTTCAGATCAGGATGCTGATTTGGTCAGAAGTTATGTGTCAAAAATTCAAGATTTGGAAGGGGAATTGCTGCGCATCAAAAACTCCAAGCTAAAACAT
ACTAGGTTTGTCAATTGTGTGGACTCTAATGATGAGACGTTCCATTCCAGTCCTGGTTTATTTCCTCCAATTGACGGTTTTGCTCCTCCAGATACAAAAT
CCTTAGTGATTGCAGATGAGGCTGAAGATGAGGCTGAAGATGAGGAAAAGGAGCTAGAGCACTCATCTATGCAAGAGAAACTTGACAGAGAGTTAAAAGA
ATTGGACCGAAGACTTGAACAAAAAGAGGCTGAAATGAAACTTTTTGCAAATGGGGACACTTCCGTTCTTAAACAACACTACGAGAAGAAAGTTCACGAC
TTAGAACAAGAAAAGAGAGAATTACAGAAAGAGATTGGTGAACTGAGAAACAATATTTCTAATATCTCGTCTACTTCTGACGACGGTACAAAGAAGTCAA
AGGAACAATATCTTCAGAAGTTGACAATCCTTGAGACACAAGTGTCTGAGTTGAAGAAGAAGCAGGACGCCCAAACCCAGTTATTGAGGCAAAAACAAAA
GAGTGATGAAGCAGCCAGAAAGCTACAGGAGGAAATTCATCGGATTAAGACTCAAAAGGTTCAACTCCAGCATAAGATCAAACATGAGTCGGAACAATTT
AGGTTGTGGAAGGCTTCACGTGAAAAAGAGGTTCTTCAGCTAAAGAAAGAGGGAAGGAAAAATGAATACGAAATGCACAAGTTATTGGCCTTGAATCAGC
GCCAGAAAATGGTTCTGCAAAGGAAGACGGAAGAAGCTTCCTTGGCTACCAAAAGACTAAAGGATCTCTTAGAATCCAAAAAGGCTTCTGCACGTGAATT
GGATAATGCTGGAAATACCAATGGTGCTGGAATTCAGGCTCTGATGCAGACAGTTGAGCATGAGCTCGAAGTTAGTCTTCGAGTGCATGCAGTGCGTTCT
GAGTATGAACGTCAAATGCAAGAGAGAGGTAAAATTGCTCAGGAACTTGCACGTTTGAAGGAAGAAACCGAGATGCTTAAGAAAACTAATTCAAGTGATA
CTCCATCAACAATGTCTCCCGGTGCAAGGAATTCGAGAATGTTTGCGCTGGAAAACATGCTGACTACGTCTTCTAACACTCTTGTCTCTATGGCCTCACA
ACTCTCTGAGGCAGAAGAGAACGGAAAAGGCTTGAGTGGCAGGGGACGTTGGAATCAAGTGAGATCACTTATTGATGGAAAACACTTGTTAAATTATCTG
TTTAATCTGGCTTCTTCATCCAGGTGCATGCTGCGAGACAAAGAAGTTGACTGCAGAGAGAAGGATGGTGAAATAAGAGAGTTGAAAGGAAAGATTGTAA
ACCTCAGTGGTTTAATTAGGCATCTGGAGAAACAGAAGGCAGAGCTCAGTCACCAAGTAAAGATACAGAGGTCAGCTTTGAAGATGGAAAATGGAAGCTG
TAAATATGAGTTGCGGAAACTGCCACCACGGAGCTCCATCTATTTGATGGAAGATATGGATACATCATTATCAGATCTTTCTGATGATTCGGAGACAAAT
GATGATGATGAATGGGTGCGATCAACCAGAAAGAGAAGCAGGAAGATGAACTCAAGAATAGGAGGACGTGACTCAAGATCACCATCAGATTCGTCATCGT
CAGGAGATGGAGACGAAGGAGATTCAACAAAAAACGAAGGTGCTGCTGCAGGAGGACAGTGCTGCTCTTGCAGCAAGAAGTCATTTTGCAAAACAACAAA
ATGTGAATGCCTAGCAGCTGGTGGAATATGCGGGTCGTCTTGTGGTTGTTCGGTGAATAAGTGCAGTAATAGAGACCCTGCTACTTATGCCATGGTGAAC
TTGGATCAATCACAGATTACTGCTGATGAATGCGGTAAGAAAAATAATAATATTGACGAGCTTGCTTCCCATGGCGCCATGCTTCTGGAGAATGCCATGT
TCAGCAAGCCTGTTGCTGCTGGGGAGGGAGAGGATGATGCCTCTGAAGCAGCACAAAGGAAGGCATTGTCAGACATAGGCAATACAAAGGCGAAGTCAAA
TGCACCGAAGCCAAACCAGAGAAAGAAATGGAGAAAATCGGTAATTCAGCTTGCCCCTGTAGCACCAACGCAACCAGAAACAACAGCAACCAATCATGTT
CCACCAGCGACACCTGAGAATGCAGCAGCAGCTCCAAGGATGATGTCTGAGAATGCTGCTAGTAGCAACGGGGTTGGCCTGAAGCTACCAAGGGCCACGA
GAGCCCCAGGGTCAAACAAGGGAGTCCTTTTAAGAGAAAGGAATAATGCCACTACTAGTCATCCAGATGAATCTGTTGTGCACAAGGAAGGCGGTGGCAG
GGGACGAATCCAGAAGTCAAAAAGAAGTCCTGCCAGGCCGGCGAGAGCTACCGAAGAGAAGGAGAACCACAGCAGGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10000392 pacid=23170849 polypeptide=Lus10000392 locus=Lus10000392.g ID=Lus10000392.BGIv1.0 annot-version=v1.0
MDSNTECVKLAVNIRPLITPELLNGCTDCITLVPGEPQVQIGSHSFTYDYVYGSLDSPSSAIYDDCVAPLVDALFHGYNATVLAYGQTGSGKTYTMGTNY
TGEGSNEGVIPKVMATIFRRLESIKQSTEFLIRVSFIEIFKEEVFDLLDTNTVISKGEGGNAGKAVVSPRVPIQIRETVNGGITLAGVSEAEVRCKEEMG
AYLSKGSLARATASTNMNKQSSRSHAIFTISMEQKKLTPCSSGGNNEDMCDDILRAKLHLVDLAGSERAKRTGADGMRFKEGIHINKGLLALGNVISALG
DEKKRKDGGHVPYRDSKLTRLLQDSLGGNSKTVMIACVSPADTNAEETLNTLKYANRARNIQNKAVINRDPNAVQLQRMRSQIEQLQAELCIYRGDANAP
FDEIQILKHKVSLLEASNAELQKELQERQMTFEHLNQRAVDAQVEKDKLIMQIELVRNGKSWDEIDAASDQDADLVRSYVSKIQDLEGELLRIKNSKLKH
TRFVNCVDSNDETFHSSPGLFPPIDGFAPPDTKSLVIADEAEDEAEDEEKELEHSSMQEKLDRELKELDRRLEQKEAEMKLFANGDTSVLKQHYEKKVHD
LEQEKRELQKEIGELRNNISNISSTSDDGTKKSKEQYLQKLTILETQVSELKKKQDAQTQLLRQKQKSDEAARKLQEEIHRIKTQKVQLQHKIKHESEQF
RLWKASREKEVLQLKKEGRKNEYEMHKLLALNQRQKMVLQRKTEEASLATKRLKDLLESKKASARELDNAGNTNGAGIQALMQTVEHELEVSLRVHAVRS
EYERQMQERGKIAQELARLKEETEMLKKTNSSDTPSTMSPGARNSRMFALENMLTTSSNTLVSMASQLSEAEENGKGLSGRGRWNQVRSLIDGKHLLNYL
FNLASSSRCMLRDKEVDCREKDGEIRELKGKIVNLSGLIRHLEKQKAELSHQVKIQRSALKMENGSCKYELRKLPPRSSIYLMEDMDTSLSDLSDDSETN
DDDEWVRSTRKRSRKMNSRIGGRDSRSPSDSSSSGDGDEGDSTKNEGAAAGGQCCSCSKKSFCKTTKCECLAAGGICGSSCGCSVNKCSNRDPATYAMVN
LDQSQITADECGKKNNNIDELASHGAMLLENAMFSKPVAAGEGEDDASEAAQRKALSDIGNTKAKSNAPKPNQRKKWRKSVIQLAPVAPTQPETTATNHV
PPATPENAAAAPRMMSENAASSNGVGLKLPRATRAPGSNKGVLLRERNNATTSHPDESVVHKEGGGRGRIQKSKRSPARPARATEEKENHSR
|
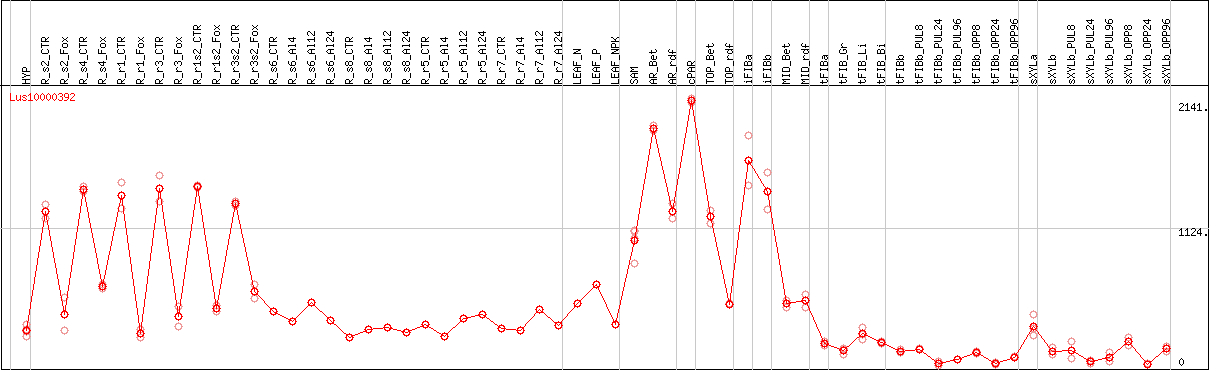
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000392 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
