External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G43940 130 / 7e-35
PAR2, ATGSNOR1, HOT5, GSNOR, ADH2
PARAQUAT RESISTANT 2, sensitive to hot temperatures 5, S-NITROSOGLUTATHIONE REDUCTASE, ALCOHOL DEHYDROGENASE 2, GroES-like zinc-binding dehydrogenase family protein (.1.2)
AT1G77120 105 / 6e-26
ADH1, ATADH, ATADH1
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
AT1G32780 96 / 2e-22
GroES-like zinc-binding dehydrogenase family protein (.1)
AT1G64710 93 / 2e-21
GroES-like zinc-binding dehydrogenase family protein (.1.2)
AT5G42250 92 / 7e-21
Zinc-binding alcohol dehydrogenase family protein (.1)
AT5G24760 89 / 3e-20
GroES-like zinc-binding dehydrogenase family protein (.1.2.3)
AT1G22430 88 / 2e-19
GroES-like zinc-binding dehydrogenase family protein (.1.2)
AT1G22440 86 / 1e-18
Zinc-binding alcohol dehydrogenase family protein (.1)
AT4G22110 82 / 2e-17
GroES-like zinc-binding dehydrogenase family protein (.1.2)
AT5G63620 43 / 0.0002
GroES-like zinc-binding alcohol dehydrogenase family protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10000496
125 / 2e-32
AT5G43940 693 / 0.0
PARAQUAT RESISTANT 2, sensitive to hot temperatures 5, S-NITROSOGLUTATHIONE REDUCTASE, ALCOHOL DEHYDROGENASE 2, GroES-like zinc-binding dehydrogenase family protein (.1.2)
Lus10022853
120 / 2e-30
AT5G43940 700 / 0.0
PARAQUAT RESISTANT 2, sensitive to hot temperatures 5, S-NITROSOGLUTATHIONE REDUCTASE, ALCOHOL DEHYDROGENASE 2, GroES-like zinc-binding dehydrogenase family protein (.1.2)
Lus10042787
105 / 1e-25
AT1G77120 674 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Lus10029759
104 / 3e-25
AT1G77120 673 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Lus10034042
103 / 4e-25
AT1G77120 672 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Lus10010506
103 / 4e-25
AT1G77120 673 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Lus10031890
97 / 1e-22
AT5G24760 566 / 0.0
GroES-like zinc-binding dehydrogenase family protein (.1.2.3)
Lus10031319
97 / 1e-22
AT5G24760 353 / 2e-135
GroES-like zinc-binding dehydrogenase family protein (.1.2.3)
Lus10033241
93 / 4e-21
AT1G32780 511 / 0.0
GroES-like zinc-binding dehydrogenase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G254900
135 / 7e-37
AT5G43940 694 / 0.0
PARAQUAT RESISTANT 2, sensitive to hot temperatures 5, S-NITROSOGLUTATHIONE REDUCTASE, ALCOHOL DEHYDROGENASE 2, GroES-like zinc-binding dehydrogenase family protein (.1.2)
Potri.014G193800
132 / 6e-36
AT5G43940 684 / 0.0
PARAQUAT RESISTANT 2, sensitive to hot temperatures 5, S-NITROSOGLUTATHIONE REDUCTASE, ALCOHOL DEHYDROGENASE 2, GroES-like zinc-binding dehydrogenase family protein (.1.2)
Potri.007G108601
109 / 2e-27
AT1G77120 634 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Potri.007G108401
109 / 3e-27
AT1G77120 630 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Potri.005G060000
109 / 3e-27
AT1G77120 658 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Potri.005G060300
108 / 5e-27
AT1G77120 626 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Potri.005G060200
108 / 5e-27
AT1G77120 627 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Potri.007G108500
107 / 1e-26
AT1G77120 626 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Potri.007G108701
104 / 2e-25
AT1G77120 656 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Potri.011G152800
103 / 6e-25
AT1G32780 540 / 0.0
GroES-like zinc-binding dehydrogenase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF00107
ADH_zinc_N
Zinc-binding dehydrogenase
Representative CDS sequence
>Lus10000497 pacid=23150912 polypeptide=Lus10000497 locus=Lus10000497.g ID=Lus10000497.BGIv1.0 annot-version=v1.0
ATGGATGGCAAAACAATTCATGTTCTTTTGTACATACTAAAAGTTACCTTTGCCTTCAACCACAAGTTAACTGCGAAATGGGTTGATTGGATGAGCATTT
GTGGTGTGTCAATCTACCAAAAAGCCTTTTCATGTACTTTTGCTAGAAGCCCCAAATCTAGGCCCATTATAATCTCGAAGCTATTCAAACTTGACCCGGA
AGGCCCAAACCGCTCACGTGCCGGATCCGGTTCTCTTGGGAGGCGGCTTGACAAACCAGACCAGTTTGACCAACCCAGCCTACCGAATCGGACCATATTA
GTTGGCCAGGAACGACTATCCGTTCATTCTTGTGTTCAACTACTTTCACTCGCCACTTCCACTCTCCTCTTCACTCTCTTTTTGCTTTCTAACGCTCGGA
TGGCGACTCAGGGCCAAGTCATCACCTGCAAAGCTGCGGTGTCGTGGGAACCTAACAAGCCGCTGGTGATCGAGGATGTTCAGGTAGCTCCGCCTCAGGC
CGGCGAGGTCCGCATCAAGATTCTCTTCACTGCTCTCTGCCACACTGATGCTTACACTTTGGAGCGGCAAGGTCTTGGAGCTGTCTGGAACACAGCTAAA
GTAGAAGCTGGGTCAAATGTTGCTATTTTTGGCCTTGATACTGTTGGCCTTGCAGTTGCGGAGGGTGCAAAAGCAGTTGGTGCTTCTCGCATTATCGGCA
TTGACATTGATAGTAAAAAGTTTGAGAGAGCAAAGGACTTTGGAGTTACAGAATTTGTGAACCCAAAAGAACACAAGAAACCGATCCAGAATGAAATAGA
GGTCGACGAGTACATCACGCATAGTCTGACACTGCCAGAGATCAATAAGGCTTTTGATCTATTGCATGAAGGAGGTTGCCTAAGGTGTGTTCTTGACTTG
CAAGTCTAA
AA sequence
>Lus10000497 pacid=23150912 polypeptide=Lus10000497 locus=Lus10000497.g ID=Lus10000497.BGIv1.0 annot-version=v1.0
MDGKTIHVLLYILKVTFAFNHKLTAKWVDWMSICGVSIYQKAFSCTFARSPKSRPIIISKLFKLDPEGPNRSRAGSGSLGRRLDKPDQFDQPSLPNRTIL
VGQERLSVHSCVQLLSLATSTLLFTLFLLSNARMATQGQVITCKAAVSWEPNKPLVIEDVQVAPPQAGEVRIKILFTALCHTDAYTLERQGLGAVWNTAK
VEAGSNVAIFGLDTVGLAVAEGAKAVGASRIIGIDIDSKKFERAKDFGVTEFVNPKEHKKPIQNEIEVDEYITHSLTLPEINKAFDLLHEGGCLRCVLDL
QV
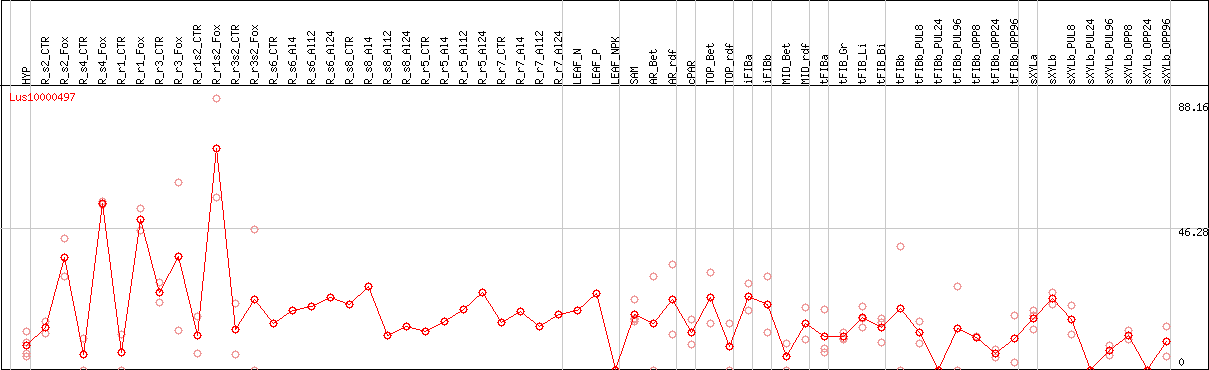
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000497 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.