External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G17720 297 / 2e-100
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT5G46870 283 / 3e-95
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT5G16840 237 / 1e-77
BPA1
binding partner of acd11 1 (.1.2.3)
AT1G67950 198 / 5e-62
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3), RNA-binding (RRM/RBD/RNP motifs) family protein (.4)
AT5G32450 178 / 2e-54
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
AT1G14340 138 / 4e-39
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT3G01210 108 / 8e-28
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10011063
444 / 4e-158
AT4G17720 333 / 2e-114
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10030952
350 / 2e-121
AT4G17720 382 / 7e-134
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10040095
345 / 2e-119
AT4G17720 379 / 8e-133
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10025743
234 / 6e-76
AT4G17720 295 / 4e-100
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10035919
186 / 2e-57
AT4G17720 226 / 3e-73
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10010660
158 / 6e-46
AT5G32450 352 / 3e-122
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10013651
158 / 9e-45
AT5G32450 343 / 5e-117
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10012843
145 / 6e-42
AT1G14340 301 / 5e-104
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10030491
144 / 3e-41
AT1G14340 300 / 1e-103
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G138400
307 / 8e-105
AT4G17720 365 / 2e-127
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.003G095400
281 / 2e-94
AT4G17720 355 / 9e-124
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.010G103600
231 / 2e-74
AT1G67950 280 / 6e-94
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3), RNA-binding (RRM/RBD/RNP motifs) family protein (.4)
Potri.013G055100
175 / 3e-53
AT5G32450 389 / 9e-138
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.019G037600
165 / 3e-49
AT5G32450 400 / 2e-142
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.017G081400
137 / 9e-39
AT1G14340 229 / 3e-75
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.017G081500
132 / 4e-37
AT1G14340 239 / 6e-80
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
PFAM info
Representative CDS sequence
>Lus10000676 pacid=23144028 polypeptide=Lus10000676 locus=Lus10000676.g ID=Lus10000676.BGIv1.0 annot-version=v1.0
ATGTCGCAGGTAAACACAATCAAAGTGAGTAATGTTTCTCTAAAGGCAACCGATAAGGACCTCAAGGAGTTCTTCTCTTTCTCTAAAGGCAACCTTTTAC
TTACTATCCCAAATATAACCCGAATCCGATCGAACTCGCCCATTTTATTATGCAGCGACGACCACCACAACTCTCAGGTGGGATACGTTACCTTCAAAGA
TTATCAAGGTGCAGAAACTGCAATTCTTCTTTCGGGAGCAACTATAGTTGATCTTGCTGTCACCATATCCTTGGATCCCTACTACAAGCTACCACAATCT
GTCCTAGCCGCAATGGAGAATGAGAGCAAGGACCCAGCAGACCCAGACTCGGCGCTACGAAAGGCAGAGGACGTGGTGAGCAGCATGCTGGCAAAGGGAT
ACATACTAGGGGAAGATGCTGTGAACAAGGCCAAGGCATTCGACGAGAAGCACCAGCTAACGTCGACAGGGTTGGCTAAGGTGGCGTCCTTCGACAAGAC
GATAGGCCTGACGGACAAGATCAGCACCAGCGTGACAGTGATGGGTGACAAAGTCCGGGAAGTGGACCAGAAGTACCAGGTCACTGAGAAGACCAAGTCT
GCACTGGCTGTTGCAGAGCAGACGGTCAGCACTGCAGGGTCTAAGATAATGAAGAACCGGTACGTGTTCACCAGCGCTGCCTGGGTGACCGGAGCCTTCA
GCAGGGTGGCCAAGGCTGCTGAGGACGTCGGGCAGAAGGCCAAAGAGAAGGCCGCCGAAGGAGAGGAGAGAAAGGAAGTTGTGGATGATGATCGTAGTGA
TAAGCAGAAGGAAGCCAAGAAGGAGGAACAGAAAGAGAAGCTCGGAGAGGGAATGAATCAGCAGTTGACTGCTGTTGATCATCCCAAGTCACCTTCTTCC
AAGTCTGCTGCACCTATTCAATAA
AA sequence
>Lus10000676 pacid=23144028 polypeptide=Lus10000676 locus=Lus10000676.g ID=Lus10000676.BGIv1.0 annot-version=v1.0
MSQVNTIKVSNVSLKATDKDLKEFFSFSKGNLLLTIPNITRIRSNSPILLCSDDHHNSQVGYVTFKDYQGAETAILLSGATIVDLAVTISLDPYYKLPQS
VLAAMENESKDPADPDSALRKAEDVVSSMLAKGYILGEDAVNKAKAFDEKHQLTSTGLAKVASFDKTIGLTDKISTSVTVMGDKVREVDQKYQVTEKTKS
ALAVAEQTVSTAGSKIMKNRYVFTSAAWVTGAFSRVAKAAEDVGQKAKEKAAEGEERKEVVDDDRSDKQKEAKKEEQKEKLGEGMNQQLTAVDHPKSPSS
KSAAPIQ
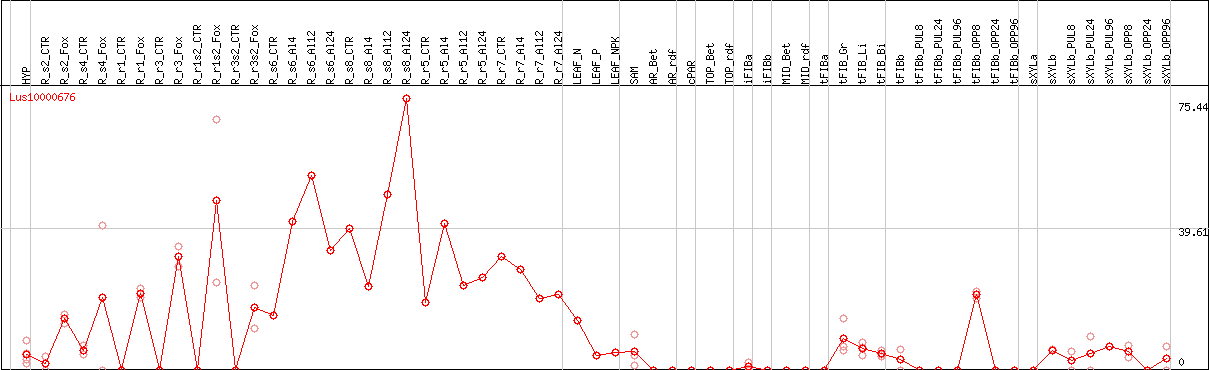
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000676 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.