External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G04770 385 / 4e-135
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G48850 363 / 1e-126
ATSDI1
SULPHUR DEFICIENCY-INDUCED 1, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G51280 274 / 7e-90
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G20900 231 / 6e-73
TDM1, MS5
MALE-STERILE 5, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G44330 206 / 7e-63
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G22794 114 / 4e-30
unknown protein
AT1G49940 55 / 9e-09
unknown protein
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10011738
579 / 0
AT1G04770 389 / 1e-136
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10011872
379 / 7e-133
AT5G48850 392 / 2e-138
SULPHUR DEFICIENCY-INDUCED 1, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10022806
380 / 2e-126
AT5G05740 644 / 0.0
ethylene-dependent gravitropism-deficient and yellow-green-like 2 (.1.2.3)
Lus10028434
270 / 5e-88
AT3G51280 560 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10028433
271 / 6e-88
AT3G51280 569 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10041887
271 / 7e-88
AT3G51280 572 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10041886
270 / 1e-87
AT3G51280 566 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10023256
244 / 1e-76
AT4G20900 330 / 4e-108
MALE-STERILE 5, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10008852
231 / 1e-71
AT4G20900 311 / 7e-101
MALE-STERILE 5, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G177600
450 / 4e-161
AT1G04770 384 / 8e-135
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.001G050600
437 / 7e-156
AT1G04770 381 / 6e-134
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.002G240700
394 / 6e-139
AT5G48850 404 / 9e-143
SULPHUR DEFICIENCY-INDUCED 1, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.007G056500
279 / 4e-91
AT3G51280 603 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G112100
274 / 5e-89
AT3G51280 595 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.011G163500
270 / 2e-85
AT4G20900 328 / 1e-105
MALE-STERILE 5, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.001G465401
268 / 2e-84
AT4G20900 337 / 1e-108
MALE-STERILE 5, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.009G033201
56 / 7e-10
AT1G04770 53 / 2e-09
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF00515
TPR_1
Tetratricopeptide repeat
Representative CDS sequence
>Lus10000753 pacid=23148782 polypeptide=Lus10000753 locus=Lus10000753.g ID=Lus10000753.BGIv1.0 annot-version=v1.0
ATGCAAGAGACGACGAGTCAACAAAGGATATCAGAGAGGATTCAGAACTTTGATGTCGTGGCTGCTCCGTATCACGTCCTTCACAAGCTTCCTCCTGGCG
ATGGCCCTTACGTTAGAGCTAAGCACGTTCAGCTAGTTGAGAAAGATCCAGTGGGAGCAGTAGAATGGTTCCTGAAAGCAATCAATTCGGGAGATAGAGT
AGATAGTGCCCTCAAGGACATGGCGCTAGTGATGAAACAACAAGACAGATCCGAAGAAGCAATTGAAGCTATCAGGCGTTTTAGACATCTCTGCTCCAAG
CAAGCTCAAGAGTCGCTCGACAATGTCCTCATTGACCTTTACAAGAAATGTGGGAAAATGGAGGAGCAGATAGAGTTATTGAAGCAGAAGCTTTGGATGA
TTTACCAAGGAGATGCGTTCAATGGCAAGCCAACTAAGACTGCACGATCTCACGGCAGGAAGTTTCAGGTCACTGTCCAGCAAGAGACTTCTCGGATATT
GGGGAACTTGGGTTGGGCATACATGCAGCTAGGCAACTATTCGGCGGCCGAAACAGTATACGTCAAAGCACAGCTAATTGACCCAGATGCCAACAAGGCC
TGCAACTTATCCATGTGCCTTTTGAAGCAAATGCGATACGATGAAGCACGTTCTGTATTGGATGATGTGGTCAAAGGCAAGATTTTGGGATACGACGACC
CCAAGTCGAGGAATCGGATCAAGGAGCTACTTGAAGAACTGGAGATGCGCCAATCCTCTCGTGGACAATCCTCCTCTTCAGTTACATTAAGTTTGGAAGA
CGCATTCGCTGAGGGACTAGACCAGCTTATGGGACAACTGACCCCGTATCGATCTAGGAGGCTTCCGATTTTCGAAGAGATCTCTCCTGTGAGAGACCAG
TTAGCTTGTTGA
AA sequence
>Lus10000753 pacid=23148782 polypeptide=Lus10000753 locus=Lus10000753.g ID=Lus10000753.BGIv1.0 annot-version=v1.0
MQETTSQQRISERIQNFDVVAAPYHVLHKLPPGDGPYVRAKHVQLVEKDPVGAVEWFLKAINSGDRVDSALKDMALVMKQQDRSEEAIEAIRRFRHLCSK
QAQESLDNVLIDLYKKCGKMEEQIELLKQKLWMIYQGDAFNGKPTKTARSHGRKFQVTVQQETSRILGNLGWAYMQLGNYSAAETVYVKAQLIDPDANKA
CNLSMCLLKQMRYDEARSVLDDVVKGKILGYDDPKSRNRIKELLEELEMRQSSRGQSSSSVTLSLEDAFAEGLDQLMGQLTPYRSRRLPIFEEISPVRDQ
LAC
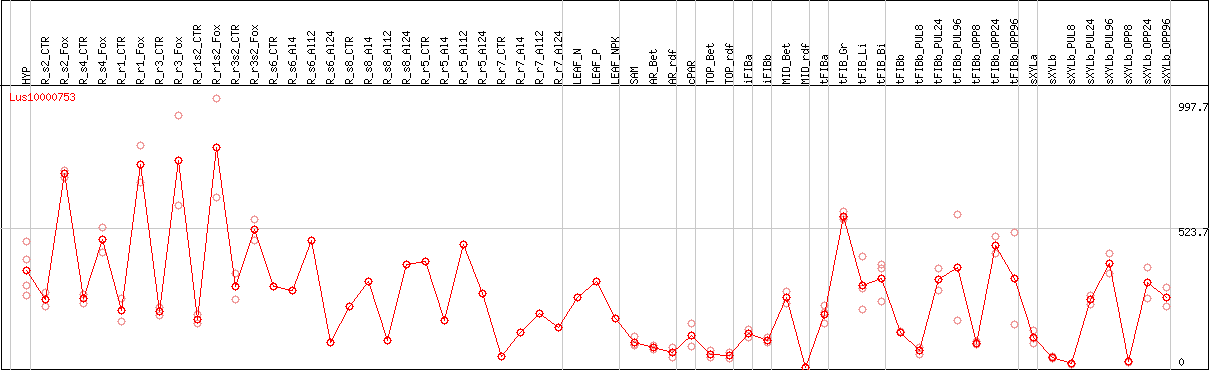
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000753 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.