External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G25810 47 / 8e-07
Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
AT2G24210 45 / 2e-06
AtTPS10
terpene synthase 10 (.1)
AT3G25830 43 / 2e-05
ATTPS-CIN
"terpene synthase-like sequence-1,8-cineole", terpene synthase-like sequence-1,8-cineole (.1)
AT3G25820 43 / 2e-05
ATTPS-CIN
"terpene synthase-like sequence-1,8-cineole", terpene synthase-like sequence-1,8-cineole (.1.2)
AT4G16730 41 / 5e-05
AtTPS02
terpene synthase 02 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10001110
66 / 2e-13
AT3G25810 362 / 1e-118
Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Lus10038179
66 / 2e-13
AT3G25810 351 / 2e-113
Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Lus10033643
64 / 5e-13
AT4G16730 356 / 2e-115
terpene synthase 02 (.1)
Lus10025922
63 / 1e-12
AT3G25810 353 / 3e-114
Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Lus10028648
56 / 7e-10
AT1G21400 645 / 0.0
Thiamin diphosphate-binding fold (THDP-binding) superfamily protein (.1)
Lus10006356
56 / 8e-10
AT3G25820 371 / 3e-121
"terpene synthase-like sequence-1,8-cineole", terpene synthase-like sequence-1,8-cineole (.1.2)
Lus10012312
53 / 7e-09
AT4G16740 363 / 3e-118
terpene synthase 03 (.1.2)
Lus10031352
49 / 8e-08
AT3G25810 236 / 2e-73
Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Lus10006355
50 / 9e-08
AT2G24210 361 / 7e-117
terpene synthase 10 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G023008
50 / 4e-08
AT3G25830 535 / 0.0
"terpene synthase-like sequence-1,8-cineole", terpene synthase-like sequence-1,8-cineole (.1)
Potri.019G023006
50 / 4e-08
AT3G25830 540 / 0.0
"terpene synthase-like sequence-1,8-cineole", terpene synthase-like sequence-1,8-cineole (.1)
Potri.019G023020
49 / 7e-08
AT3G25830 440 / 1e-151
"terpene synthase-like sequence-1,8-cineole", terpene synthase-like sequence-1,8-cineole (.1)
Potri.001G308300
49 / 9e-08
AT3G25830 507 / 4e-174
"terpene synthase-like sequence-1,8-cineole", terpene synthase-like sequence-1,8-cineole (.1)
Potri.001G308200
48 / 2e-07
AT3G25810 499 / 4e-171
Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Potri.007G119700
47 / 7e-07
AT4G16740 438 / 8e-147
terpene synthase 03 (.1.2)
Potri.019G023000
43 / 2e-05
AT4G16740 208 / 8e-63
terpene synthase 03 (.1.2)
Potri.017G041700
40 / 0.0001
AT3G25810 432 / 3e-145
Terpenoid cyclases/Protein prenyltransferases superfamily protein (.1)
Potri.011G031800
40 / 0.0001
AT3G25830 382 / 5e-126
"terpene synthase-like sequence-1,8-cineole", terpene synthase-like sequence-1,8-cineole (.1)
Potri.007G118600
39 / 0.0003
AT4G16740 462 / 3e-157
terpene synthase 03 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01397
Terpene_synth
Terpene synthase, N-terminal domain
CL0613
Terp_synthase
PF03936
Terpene_synth_C
Terpene synthase family, metal binding domain
Representative CDS sequence
>Lus10000770 pacid=23142590 polypeptide=Lus10000770 locus=Lus10000770.g ID=Lus10000770.BGIv1.0 annot-version=v1.0
ATGGATGTTCAGCCTGCTGGGATTCGGGGCATTCGTGACGAGGTCTCGGAGGACAAATTCGTGAGTACCGAAAGTGTCCTTGACAACACGATATCGGTGA
AAGAGTTGAAAAAGTACGTGAAAGGTTTGCTAGAGAAGAATGATGATGATGATCAAAAGATGATCACACCGATCAAACAGCTGGAAACGATCGACAACCT
CCAACGGCTTGGATTGGGTGAGCTAGAAAGGGGAGATGTAACGAAATCGATGGAATGTTACATGAAAGAGAAAGGGGTTTCGGAAGATGCGGCTCGAGAA
CACGTAGAGTTGTTGATCGGTGAAACATGGAAGATGGACAATCAAGACATAGGTTTCCTCCAAGCATAA
AA sequence
>Lus10000770 pacid=23142590 polypeptide=Lus10000770 locus=Lus10000770.g ID=Lus10000770.BGIv1.0 annot-version=v1.0
MDVQPAGIRGIRDEVSEDKFVSTESVLDNTISVKELKKYVKGLLEKNDDDDQKMITPIKQLETIDNLQRLGLGELERGDVTKSMECYMKEKGVSEDAARE
HVELLIGETWKMDNQDIGFLQA
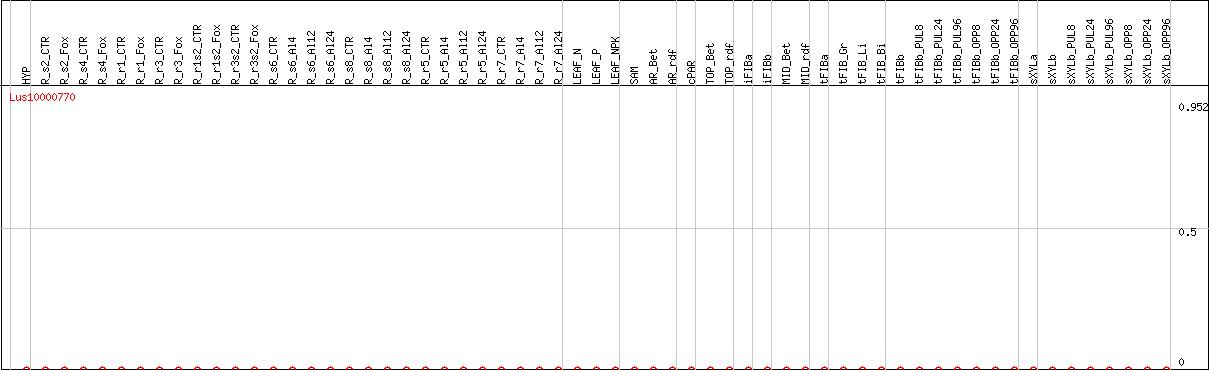
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000770 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.