External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G01260 77 / 6e-16
GeBP
DNA-binding storekeeper protein-related transcriptional regulator (.1)
AT2G01370 73 / 6e-15
GeBP
DNA-binding storekeeper protein-related transcriptional regulator (.1)
AT4G00390 71 / 9e-14
GeBP
DNA-binding storekeeper protein-related transcriptional regulator (.1)
AT1G11510 70 / 1e-13
GeBP
DNA-binding storekeeper protein-related transcriptional regulator (.1)
AT5G28040 67 / 1e-12
GeBP
DNA-binding storekeeper protein-related transcriptional regulator (.1)
AT1G66420 66 / 3e-12
GeBP
DNA-binding storekeeper protein-related transcriptional regulator (.1)
AT3G04930 66 / 5e-12
GeBP
DNA-binding storekeeper protein-related transcriptional regulator (.1.2)
AT4G00250 64 / 2e-11
GeBP
DNA-binding storekeeper protein-related transcriptional regulator (.1)
AT4G00610 62 / 9e-11
GeBP
DNA-binding storekeeper protein-related transcriptional regulator (.1)
AT4G25210 62 / 1e-10
GeBP
DNA-binding storekeeper protein-related transcriptional regulator (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10004772
113 / 1e-28
AT4G01260 95 / 1e-21
DNA-binding storekeeper protein-related transcriptional regulator (.1)
Lus10002794
103 / 4e-26
AT4G01260 96 / 4e-23
DNA-binding storekeeper protein-related transcriptional regulator (.1)
Lus10005506
103 / 3e-25
AT4G00390 108 / 9e-27
DNA-binding storekeeper protein-related transcriptional regulator (.1)
Lus10018859
88 / 3e-20
AT4G00390 115 / 9e-30
DNA-binding storekeeper protein-related transcriptional regulator (.1)
Lus10028550
84 / 3e-18
AT4G00390 122 / 1e-31
DNA-binding storekeeper protein-related transcriptional regulator (.1)
Lus10019007
75 / 4e-15
AT5G28040 256 / 1e-81
DNA-binding storekeeper protein-related transcriptional regulator (.1)
Lus10015944
75 / 6e-15
AT5G28040 264 / 2e-84
DNA-binding storekeeper protein-related transcriptional regulator (.1)
Lus10000940
70 / 3e-13
AT5G28040 272 / 2e-87
DNA-binding storekeeper protein-related transcriptional regulator (.1)
Lus10017793
54 / 2e-08
AT1G66420 72 / 6e-15
DNA-binding storekeeper protein-related transcriptional regulator (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G293900
96 / 2e-22
AT4G25210 142 / 9e-39
DNA-binding storekeeper protein-related transcriptional regulator (.1)
Potri.009G088400
95 / 3e-22
AT4G25210 133 / 2e-35
DNA-binding storekeeper protein-related transcriptional regulator (.1)
Potri.001G261700
72 / 3e-14
AT5G28040 239 / 9e-75
DNA-binding storekeeper protein-related transcriptional regulator (.1)
Potri.009G056400
69 / 5e-13
AT5G28040 229 / 3e-71
DNA-binding storekeeper protein-related transcriptional regulator (.1)
Potri.007G119800
59 / 1e-09
AT1G61730 101 / 3e-23
DNA-binding storekeeper protein-related transcriptional regulator (.1)
Potri.005G075000
47 / 1e-05
AT1G61730 75 / 8e-15
DNA-binding storekeeper protein-related transcriptional regulator (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF04504
DUF573
Protein of unknown function, DUF573
Representative CDS sequence
>Lus10000807 pacid=23175379 polypeptide=Lus10000807 locus=Lus10000807.g ID=Lus10000807.BGIv1.0 annot-version=v1.0
ATGGCTCCCCAACACCCTGAAGAAGAAGAGATATCAGCTGCCTCCAACAAGCCTGTTGACAAGTCCCATTCCGAGGGCTCTGATTCGGACGATCATTCCA
ATTCAGCCTCAGAAGAACAAGACTTAGCTCCTCCCTCCAACAAGCCACTCAATGCTGAATCTGAAGCAGAAGAATCACCTACAGGCCCAAAGATTAATAA
TGTGACTGCTCCGACCGAAACACCAAAGAATGTGACTCCTCCTCCGGCAAGGAAGAAGAGGGGCAACGAAGAAACTGAAGGTGAAGAAGGTAATAAAGCA
AAGCGAGGACGGAATGAACAAGTTGCTGTTGATGTTCAGAAATCTGCACAATCTACAAGCAACCATCTCTTTCAGAAAATATGGAGCGATGATGATGAGA
TTAAGTTGTTACAAGGACTCGTCATCAAGTATTCCCGAAAGGGAAATGGAGTTGACCCGGCTGAAATGCAAATACACTGGTCTGATTTCTATAATTCCAT
CAAGCCATGTATTAGCTTTGATGTCAATCTTTCTCAGGTCAAAGATAAAGTTAGGAAACTGAAGGCCAAGTTCCTGAACCACTTCAACAAAGGAACTGGT
GAAGGTACGAGATTCACTGAACCTCATCATCAACTGTGTTCTGATTTGTCAATGATGATTTGGGGATCTAGCAATGCAAAGACGACAACACTGAGGCCAC
GAAGAAACATCAACAACAATGAAGAGGTTAGTTGCAGAGTGCCCCTGAGAATGGACAAGAAATGGAACAACTTTTATGCGGCTAGTTTTGAGCTTATGAT
TAAGAGGTATGCCTTGTTGATTGACGAAGCCAATTTAATGCGGGCTTCCATGGACGAAGATGATTAG
AA sequence
>Lus10000807 pacid=23175379 polypeptide=Lus10000807 locus=Lus10000807.g ID=Lus10000807.BGIv1.0 annot-version=v1.0
MAPQHPEEEEISAASNKPVDKSHSEGSDSDDHSNSASEEQDLAPPSNKPLNAESEAEESPTGPKINNVTAPTETPKNVTPPPARKKRGNEETEGEEGNKA
KRGRNEQVAVDVQKSAQSTSNHLFQKIWSDDDEIKLLQGLVIKYSRKGNGVDPAEMQIHWSDFYNSIKPCISFDVNLSQVKDKVRKLKAKFLNHFNKGTG
EGTRFTEPHHQLCSDLSMMIWGSSNAKTTTLRPRRNINNNEEVSCRVPLRMDKKWNNFYAASFELMIKRYALLIDEANLMRASMDEDD
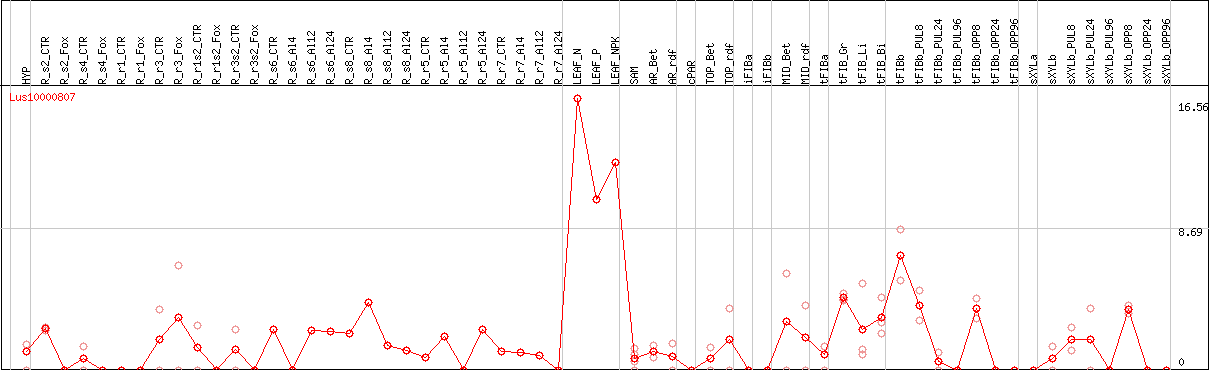
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000807 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.