External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G13680 286 / 8e-89
GLS2, ATGSL02, CALS5
ARABIDOPSIS THALIANA GLUCAN SYNTHASE-LIKE 2, callose synthase 5 (.1)
AT5G13000 181 / 3e-52
CALS3, ATGSL12
callose synthase 3, glucan synthase-like 12 (.1.2)
AT2G31960 180 / 7e-52
ATGSL3, ATGSL03
glucan synthase-like 3 (.1.2)
AT1G05570 178 / 2e-51
ATGSL6, ATGSL06, GSL6, CALS1
GLUCAN SYNTHASE-LIKE 6, callose synthase 1 (.1.2)
AT3G14570 118 / 2e-30
ATGSL4, ATGSL04
glucan synthase-like 4 (.1.2)
AT5G36870 117 / 7e-30
ATGSL9, ATGSL09
glucan synthase-like 9 (.1)
AT3G59100 108 / 4e-27
ATGSL11
glucan synthase-like 11 (.1)
AT1G06490 105 / 7e-26
CalS7, ATGSL7, ATGSL07
callose synthase 7, Arabidopsis thaliana glucan synthase-like 7, glucan synthase-like 7 (.1)
AT4G03550 81 / 2e-17
ATGSL5, PMR4, GSL5, ATGSL05
POWDERY MILDEW RESISTANT 4, glucan synthase-like 5 (.1)
AT3G07160 77 / 3e-16
CALS9, ATGSL10
glucan synthase-like 10 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002097
342 / 5e-110
AT2G13680 1916 / 0.0
ARABIDOPSIS THALIANA GLUCAN SYNTHASE-LIKE 2, callose synthase 5 (.1)
Lus10033482
328 / 2e-106
AT2G13680 1556 / 0.0
ARABIDOPSIS THALIANA GLUCAN SYNTHASE-LIKE 2, callose synthase 5 (.1)
Lus10020893
239 / 1e-72
AT2G13680 3037 / 0.0
ARABIDOPSIS THALIANA GLUCAN SYNTHASE-LIKE 2, callose synthase 5 (.1)
Lus10039199
176 / 3e-50
AT5G13000 3524 / 0.0
callose synthase 3, glucan synthase-like 12 (.1.2)
Lus10013744
176 / 3e-50
AT1G05570 2981 / 0.0
GLUCAN SYNTHASE-LIKE 6, callose synthase 1 (.1.2)
Lus10003920
169 / 6e-48
AT1G05570 3231 / 0.0
GLUCAN SYNTHASE-LIKE 6, callose synthase 1 (.1.2)
Lus10037469
168 / 7e-48
AT1G05570 3143 / 0.0
GLUCAN SYNTHASE-LIKE 6, callose synthase 1 (.1.2)
Lus10042478
124 / 3e-32
AT3G14570 2889 / 0.0
glucan synthase-like 4 (.1.2)
Lus10020751
111 / 5e-28
AT3G59100 934 / 0.0
glucan synthase-like 11 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G058300
295 / 3e-92
AT2G13680 3190 / 0.0
ARABIDOPSIS THALIANA GLUCAN SYNTHASE-LIKE 2, callose synthase 5 (.1)
Potri.001G012200
186 / 3e-54
AT5G13000 3492 / 0.0
callose synthase 3, glucan synthase-like 12 (.1.2)
Potri.003G214200
178 / 3e-51
AT5G13000 3363 / 0.0
callose synthase 3, glucan synthase-like 12 (.1.2)
Potri.001G230000
168 / 9e-48
AT5G13000 3157 / 0.0
callose synthase 3, glucan synthase-like 12 (.1.2)
Potri.005G203500
110 / 1e-27
AT1G06490 2914 / 0.0
callose synthase 7, Arabidopsis thaliana glucan synthase-like 7, glucan synthase-like 7 (.1)
Potri.002G058700
103 / 2e-25
AT1G06490 2961 / 0.0
callose synthase 7, Arabidopsis thaliana glucan synthase-like 7, glucan synthase-like 7 (.1)
Potri.011G095100
93 / 1e-21
AT3G14570 2863 / 0.0
glucan synthase-like 4 (.1.2)
Potri.013G131300
86 / 3e-19
AT4G03550 2752 / 0.0
POWDERY MILDEW RESISTANT 4, glucan synthase-like 5 (.1)
Potri.001G011900
72 / 2e-14
AT3G07160 2981 / 0.0
glucan synthase-like 10 (.1)
Potri.011G052400
72 / 2e-14
AT4G04970 2583 / 0.0
GLUCAN SYNTHASE LIKE-1, GLUCAN SYNTHASE LIKE 1, glucan synthase-like 1 (.1)
PFAM info
Representative CDS sequence
>Lus10000832 pacid=23170690 polypeptide=Lus10000832 locus=Lus10000832.g ID=Lus10000832.BGIv1.0 annot-version=v1.0
ATGATGGTGGATGCGGAACTTCAACTCCGAACAGAATTTCACCAATTTAAGGCCCATGATTCTTGGGCACGAGTCAGCTTCATGGCCTGCCAAGAAAATG
CTTCCAGTATCGCATCCCGGGTTAAGAAGACGGATGCGCGCGAAATCCAGAGCTTCTATTACCAGTACTATAAACAATATGTCGAAGCACTCGAGCAGGG
CGAGCAGGCAGATAGAGCTCAACTTGGCAAGGTCTATGAGACAGCAGGAGTACTTTATGAAGTACTTTGTGCCGTTAATAAGGCTGAGAAAGTTGAAGAG
GTTGCTCCTGAGATTATTGCTGCAGCACGAGACATCCAAGAGAAGAAGGAAATCTATGCTCCATACAACATTCTTCCCTTGGATTCTGCCGGGGCCTCAC
AGTCAATCATGCAACTCGAGGAGGTCAAGGCTGCTGTTGCAGCACTATGGAACACCCGAGGCTTGTCTTGGCCTTCATCTTTTGAGCAGCACAGGCAGAA
AGCTGGAGATCTGGACCTTCTTGATTGGCTAAGGGCCATGTTTGGCTTCCAGAGAGACAGTGTTCGGAACCAAAGGGAGCATTTGATATTGCTGCTTGCC
AACAACCATATAAGGCTCAATCCCAAGCCTGAACCTCTTAACAAGGCACGTGCTGGTCTGCGTGCTTGA
AA sequence
>Lus10000832 pacid=23170690 polypeptide=Lus10000832 locus=Lus10000832.g ID=Lus10000832.BGIv1.0 annot-version=v1.0
MMVDAELQLRTEFHQFKAHDSWARVSFMACQENASSIASRVKKTDAREIQSFYYQYYKQYVEALEQGEQADRAQLGKVYETAGVLYEVLCAVNKAEKVEE
VAPEIIAAARDIQEKKEIYAPYNILPLDSAGASQSIMQLEEVKAAVAALWNTRGLSWPSSFEQHRQKAGDLDLLDWLRAMFGFQRDSVRNQREHLILLLA
NNHIRLNPKPEPLNKARAGLRA
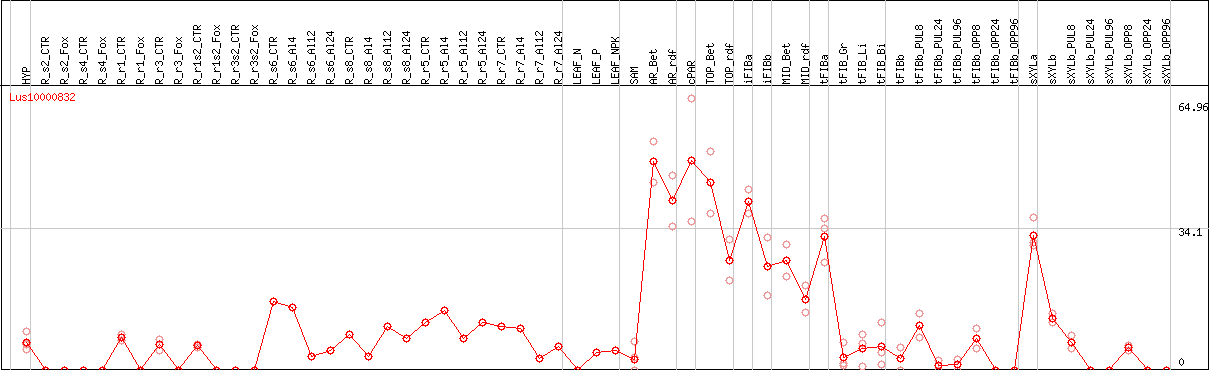
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000832 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.