External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G23560 87 / 1e-22
ATMES7
ARABIDOPSIS THALIANA METHYL ESTERASE 7, methyl esterase 7 (.1)
AT2G23610 85 / 8e-22
ATMES3
ARABIDOPSIS THALIANA METHYL ESTERASE 3, methyl esterase 3 (.1)
AT2G23600 85 / 1e-21
ATMES2, ACL, ATME8
ARABIDOPSIS THALIANA METHYL ESTERASE 2, ARABIDOPSIS METHYL ESTERASE 8, acetone-cyanohydrin lyase (.1)
AT2G23580 81 / 9e-21
ATMES4, ABE4
ARABIDOPSIS THALIANA METHYL ESTERASE 4, ALPHA/BETA FOLD HYDROLASE/ESTERASE 4, methyl esterase 4 (.1.2)
AT2G23590 81 / 4e-20
ATMES8
methyl esterase 8 (.1)
AT2G23620 80 / 1e-19
ATMES1
ARABIDOPSIS THALIANA METHYL ESTERASE 1, methyl esterase 1 (.1)
AT4G37140 73 / 4e-18
ATMES20, MEE69
MATERNAL EFFECT EMBRYO ARREST 69, ARABIDOPSIS THALIANA METHYL ESTERASE 20, alpha/beta-Hydrolases superfamily protein (.1)
AT2G23550 75 / 6e-18
ATMES6, ABE1
ALPHA/BETA FOLD HYDROLASE/ESTERASE 1, methyl esterase 6 (.1.2)
AT3G50440 75 / 8e-18
ATMES10
ARABIDOPSIS THALIANA METHYL ESTERASE 10, methyl esterase 10 (.1)
AT5G10300 74 / 1e-17
AtHNL, HNL, ATMES5
HYDROXYNITRILE LYASE, ARABIDOPSIS THALIANA METHYL ESTERASE 5, methyl esterase 5 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10019645
141 / 4e-45
AT2G23600 139 / 7e-42
ARABIDOPSIS THALIANA METHYL ESTERASE 2, ARABIDOPSIS METHYL ESTERASE 8, acetone-cyanohydrin lyase (.1)
Lus10023326
107 / 2e-30
AT3G50440 263 / 3e-88
ARABIDOPSIS THALIANA METHYL ESTERASE 10, methyl esterase 10 (.1)
Lus10012854
92 / 2e-25
AT2G23600 147 / 2e-45
ARABIDOPSIS THALIANA METHYL ESTERASE 2, ARABIDOPSIS METHYL ESTERASE 8, acetone-cyanohydrin lyase (.1)
Lus10022467
86 / 4e-22
AT3G50440 238 / 2e-78
ARABIDOPSIS THALIANA METHYL ESTERASE 10, methyl esterase 10 (.1)
Lus10005400
83 / 5e-21
AT2G23610 206 / 8e-66
ARABIDOPSIS THALIANA METHYL ESTERASE 3, methyl esterase 3 (.1)
Lus10005402
82 / 1e-20
AT2G23580 204 / 3e-65
ARABIDOPSIS THALIANA METHYL ESTERASE 4, ALPHA/BETA FOLD HYDROLASE/ESTERASE 4, methyl esterase 4 (.1.2)
Lus10030707
81 / 1e-19
AT3G29770 494 / 4e-175
ARABIDOPSIS THALIANA METHYL ESTERASE 11, methyl esterase 11 (.1)
Lus10015532
79 / 1e-19
AT2G23600 240 / 2e-79
ARABIDOPSIS THALIANA METHYL ESTERASE 2, ARABIDOPSIS METHYL ESTERASE 8, acetone-cyanohydrin lyase (.1)
Lus10013193
80 / 2e-19
AT3G29770 493 / 8e-175
ARABIDOPSIS THALIANA METHYL ESTERASE 11, methyl esterase 11 (.1)
PFAM info
Representative CDS sequence
>Lus10000840 pacid=23148149 polypeptide=Lus10000840 locus=Lus10000840.g ID=Lus10000840.BGIv1.0 annot-version=v1.0
ATGAAGAAACAACACTTTGTACTAGTTCATGGAACAAACCATGGAGCCTGGTGCTGGTACAAGGTCATGGCCCGGCTAGAATCAGCCGGGCACATCGTTA
CAGCGTTGGACCTCGGAGGAAGTGGAGTCAACTTTGACCGAGTTTCGTCCATCTTCGACTACGTCAACCCGCTGGTTCAGTTCATGGCTGATCTGTCTGA
AGATGAAGAAGAAAGAGTCAATCCTTGTGGGTCATAG
AA sequence
>Lus10000840 pacid=23148149 polypeptide=Lus10000840 locus=Lus10000840.g ID=Lus10000840.BGIv1.0 annot-version=v1.0
MKKQHFVLVHGTNHGAWCWYKVMARLESAGHIVTALDLGGSGVNFDRVSSIFDYVNPLVQFMADLSEDEEERVNPCGS
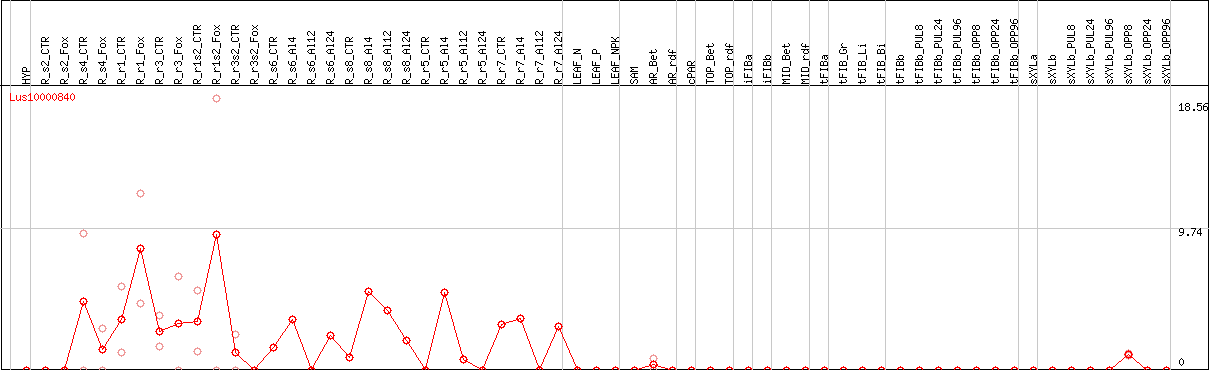
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000840 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.