External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G04550 307 / 1e-105
DSPTP1E, IBR5
DUAL SPECIFICITY PROTEIN PHOSPHATASE 1E, indole-3-butyric acid response 5 (.1.3)
AT3G23610 95 / 4e-23
DSPTP1
dual specificity protein phosphatase 1 (.1.2.3)
AT3G06110 77 / 4e-17
DSPTP1B, MKP2, ATMKP2
DUAL-SPECIFICITY PROTEIN PHOSPHATASE 1B, ARABIDOPSIS MAPK PHOSPHATASE 2, MAPK phosphatase 2 (.1.2.3)
AT3G55270 72 / 5e-14
MKP1, ATMKP1
ARABIDOPSIS MITOGEN-ACTIVATED PROTEIN KINASE PHOSPHATASE 1, mitogen-activated protein kinase phosphatase 1 (.1)
AT5G23720 53 / 1e-07
PHS1
PROPYZAMIDE-HYPERSENSITIVE 1, dual specificity protein phosphatase family protein (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002263
499 / 0
AT2G04550 327 / 1e-113
DUAL SPECIFICITY PROTEIN PHOSPHATASE 1E, indole-3-butyric acid response 5 (.1.3)
Lus10021940
90 / 9e-22
AT3G23610 248 / 1e-84
dual specificity protein phosphatase 1 (.1.2.3)
Lus10041227
80 / 1e-17
AT3G23610 237 / 4e-80
dual specificity protein phosphatase 1 (.1.2.3)
Lus10003291
72 / 8e-14
AT3G55270 800 / 0.0
ARABIDOPSIS MITOGEN-ACTIVATED PROTEIN KINASE PHOSPHATASE 1, mitogen-activated protein kinase phosphatase 1 (.1)
Lus10006229
70 / 3e-13
AT3G23610 512 / 1e-171
dual specificity protein phosphatase 1 (.1.2.3)
Lus10034033
66 / 3e-13
AT3G23610 168 / 5e-54
dual specificity protein phosphatase 1 (.1.2.3)
Lus10030324
69 / 4e-13
AT3G55270 806 / 0.0
ARABIDOPSIS MITOGEN-ACTIVATED PROTEIN KINASE PHOSPHATASE 1, mitogen-activated protein kinase phosphatase 1 (.1)
Lus10036877
69 / 6e-13
AT3G23610 511 / 3e-171
dual specificity protein phosphatase 1 (.1.2.3)
Lus10032545
61 / 2e-10
AT5G23720 1120 / 0.0
PROPYZAMIDE-HYPERSENSITIVE 1, dual specificity protein phosphatase family protein (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G160500
389 / 2e-137
AT2G04550 333 / 6e-116
DUAL SPECIFICITY PROTEIN PHOSPHATASE 1E, indole-3-butyric acid response 5 (.1.3)
Potri.010G033000
86 / 3e-20
AT3G23610 249 / 4e-85
dual specificity protein phosphatase 1 (.1.2.3)
Potri.010G210900
72 / 7e-14
AT3G55270 715 / 0.0
ARABIDOPSIS MITOGEN-ACTIVATED PROTEIN KINASE PHOSPHATASE 1, mitogen-activated protein kinase phosphatase 1 (.1)
Potri.008G049900
71 / 8e-14
AT3G55270 710 / 0.0
ARABIDOPSIS MITOGEN-ACTIVATED PROTEIN KINASE PHOSPHATASE 1, mitogen-activated protein kinase phosphatase 1 (.1)
Potri.015G105000
60 / 6e-10
AT5G23720 1096 / 0.0
PROPYZAMIDE-HYPERSENSITIVE 1, dual specificity protein phosphatase family protein (.1.2.3)
Potri.012G105800
52 / 2e-07
AT5G23720 1006 / 0.0
PROPYZAMIDE-HYPERSENSITIVE 1, dual specificity protein phosphatase family protein (.1.2.3)
Potri.006G040300
51 / 4e-07
AT4G18593 148 / 5e-44
dual specificity protein phosphatase-related (.1)
Potri.016G035700
50 / 8e-07
AT4G18593 149 / 3e-44
dual specificity protein phosphatase-related (.1)
Potri.016G110700
43 / 0.0002
AT3G09100 914 / 0.0
mRNA capping enzyme family protein (.1.2)
Potri.006G096000
41 / 0.0006
AT3G09100 948 / 0.0
mRNA capping enzyme family protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0031
Phosphatase
PF00782
DSPc
Dual specificity phosphatase, catalytic domain
Representative CDS sequence
>Lus10000921 pacid=23170817 polypeptide=Lus10000921 locus=Lus10000921.g ID=Lus10000921.BGIv1.0 annot-version=v1.0
ATGATGAGGAAGAGGGAAAGGGAGAACCCTTGCGGGATTTGCGGGCACTACCACAAGTACGAGGAAGGAGAAGTTTGCGGGATATGCGGCCACCGCTTGC
CGGATTCTTCCTCCGCCTCCGCCGCCGGCGGCGGCAATCCCTCCGTTCACATCAGCGCTTTCCCTTCCGAGATCGTCCCGGACTTCCTTTACCTAGGTAG
CTACGACAACGCTGCTAGGTCCGAGCTGCTTAAAACCCAGGGGATCTCCCGGATTCTCAATACAGTACCTGCTTGTCAAAACCTTTACAAGAACTCCTTC
ACCTATCATTGTCTCCAAGATGACAAAACTCTACAGTTCGATGAAGCTATACAAATCCTAGAACAATGCGAGAGGGAAAAAGCTCGTATCCTCGTGCACT
GCATGTCCGGAAAGAGTAGGTCTCCGGCTATCGTGATGGCTTATTTGATGAAGTCCAAAGGATGGAGGCTGGCGCAAAGTTACCAGTGGGTTAAAGAACG
AAGGCCTGCTGTTGAATTGACCGAAGGGGTGCACCAGCAGTTGCAGGAGTACGAGCAGAGGCTATTCTCGTCAATTGACAACAAAAATCCTCCGGAAAAT
TTCGTATTCCCGCCACCAGCAGGCATCCCATCTTTCAGTTTCGGCTTCCAAAAGGCAGCCAACGATACCGCACCTTCTCCACCTACATTTCCTTCTGGCG
CTACCTCGATCTTCGGTCAACCACCGGACATCCCTCCAATGGAGTTCAAATTCGGTGCCGGCAGCAGCAGTCCATTTGGCTCAAATCCGAAGAGCTCTAG
CAATGGAGACATCTCAATGGATACATGA
AA sequence
>Lus10000921 pacid=23170817 polypeptide=Lus10000921 locus=Lus10000921.g ID=Lus10000921.BGIv1.0 annot-version=v1.0
MMRKRERENPCGICGHYHKYEEGEVCGICGHRLPDSSSASAAGGGNPSVHISAFPSEIVPDFLYLGSYDNAARSELLKTQGISRILNTVPACQNLYKNSF
TYHCLQDDKTLQFDEAIQILEQCEREKARILVHCMSGKSRSPAIVMAYLMKSKGWRLAQSYQWVKERRPAVELTEGVHQQLQEYEQRLFSSIDNKNPPEN
FVFPPPAGIPSFSFGFQKAANDTAPSPPTFPSGATSIFGQPPDIPPMEFKFGAGSSSPFGSNPKSSSNGDISMDT
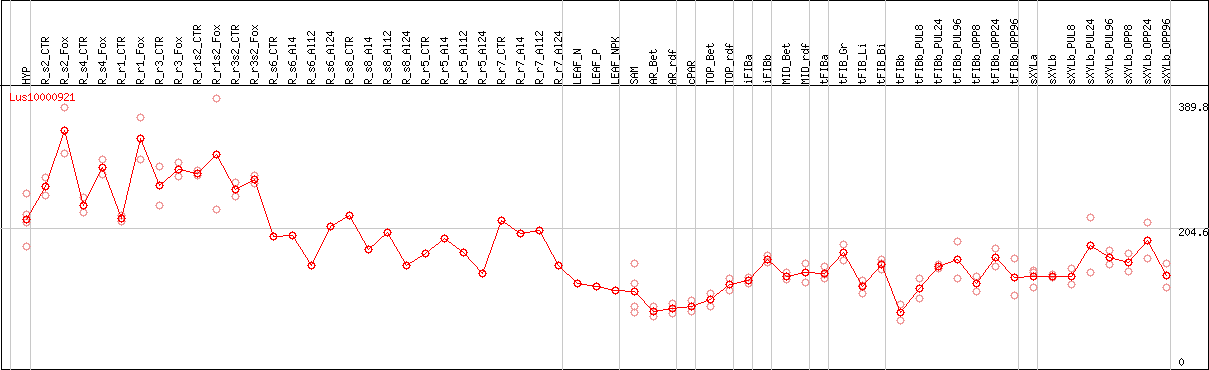
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000921 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.