External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G19810 74 / 2e-16
ChiC
class V chitinase, Glycosyl hydrolase family protein with chitinase insertion domain (.1)
AT4G19800 67 / 3e-14
Glycosyl hydrolase family protein with chitinase insertion domain (.1)
AT4G19770 59 / 2e-11
Glycosyl hydrolase family protein with chitinase insertion domain (.1)
AT4G19720 58 / 5e-11
Glycosyl hydrolase family protein with chitinase insertion domain (.1)
AT4G19730 54 / 2e-09
Glycosyl hydrolase superfamily protein (.1)
AT4G19820 54 / 2e-09
Glycosyl hydrolase family protein with chitinase insertion domain (.1)
AT4G19740 49 / 8e-08
Glycosyl hydrolase superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10019061
204 / 5e-63
AT4G21380 338 / 5e-104
receptor kinase 3 (.1)
Lus10040021
99 / 3e-25
AT4G19810 278 / 5e-89
class V chitinase, Glycosyl hydrolase family protein with chitinase insertion domain (.1)
Lus10003408
76 / 1e-18
AT4G19770 98 / 5e-26
Glycosyl hydrolase family protein with chitinase insertion domain (.1)
Lus10017124
55 / 9e-10
AT4G19810 260 / 9e-84
class V chitinase, Glycosyl hydrolase family protein with chitinase insertion domain (.1)
Lus10018329
54 / 2e-09
AT4G19810 268 / 1e-86
class V chitinase, Glycosyl hydrolase family protein with chitinase insertion domain (.1)
Lus10041639
48 / 2e-07
AT4G11900 338 / 7e-104
S-locus lectin protein kinase family protein (.1)
Lus10024081
47 / 8e-07
AT1G61500 333 / 5e-102
S-locus lectin protein kinase family protein (.1)
Lus10032626
0 / 1
AT4G19810 105 / 3e-27
class V chitinase, Glycosyl hydrolase family protein with chitinase insertion domain (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G111900
107 / 4e-28
AT3G16030 367 / 2e-114
CALLUS EXPRESSION OF RBCS 101, lectin protein kinase family protein (.1)
Potri.018G111800
87 / 4e-21
AT4G23200 362 / 4e-115
cysteine-rich RLK (RECEPTOR-like protein kinase) 12 (.1)
Potri.018G111700
84 / 4e-20
AT4G23200 363 / 1e-115
cysteine-rich RLK (RECEPTOR-like protein kinase) 12 (.1)
Potri.006G188400
62 / 2e-12
AT4G19810 477 / 2e-169
class V chitinase, Glycosyl hydrolase family protein with chitinase insertion domain (.1)
Potri.006G262001
57 / 1e-10
AT4G19810 283 / 2e-92
class V chitinase, Glycosyl hydrolase family protein with chitinase insertion domain (.1)
Potri.006G261800
57 / 1e-10
AT4G19810 286 / 2e-92
class V chitinase, Glycosyl hydrolase family protein with chitinase insertion domain (.1)
Potri.018G112000
54 / 2e-09
AT4G19810 284 / 3e-93
class V chitinase, Glycosyl hydrolase family protein with chitinase insertion domain (.1)
Potri.018G112100
52 / 5e-09
AT4G19810 279 / 3e-91
class V chitinase, Glycosyl hydrolase family protein with chitinase insertion domain (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0058
Glyco_hydro_tim
PF00704
Glyco_hydro_18
Glycosyl hydrolases family 18
Representative CDS sequence
>Lus10000981 pacid=23163396 polypeptide=Lus10000981 locus=Lus10000981.g ID=Lus10000981.BGIv1.0 annot-version=v1.0
ATGACCTCCATTATTTCCTGTCAGAGCACCTTCATCCGATCATCAATCCAAACCACCAGACTCTACGGCTTCCAAAGAATCGATTTGTTGACCCACGGTA
CGGACGTTGGGAAGATAGAAACCCTATTTGAGGAATTCAGATCTGCGATCGATTCAGAAGCCCAAAAATCGAACAATTCGAAGCTCTTACTTACCATGGC
GGTTCCAAGATCTTCGAACCTGAATTCCGCCGGGTATCTAGTTCGATCAATGGCGACGAATCTGGTTTGGGCTAACCCGTTGGCCCACGATTACCACTTG
CCGGCGAAGGAGAACTTCACGGTGAACCACGACGCGTTATTCGGTGATGATGATGGCGGCTAG
AA sequence
>Lus10000981 pacid=23163396 polypeptide=Lus10000981 locus=Lus10000981.g ID=Lus10000981.BGIv1.0 annot-version=v1.0
MTSIISCQSTFIRSSIQTTRLYGFQRIDLLTHGTDVGKIETLFEEFRSAIDSEAQKSNNSKLLLTMAVPRSSNLNSAGYLVRSMATNLVWANPLAHDYHL
PAKENFTVNHDALFGDDDGG
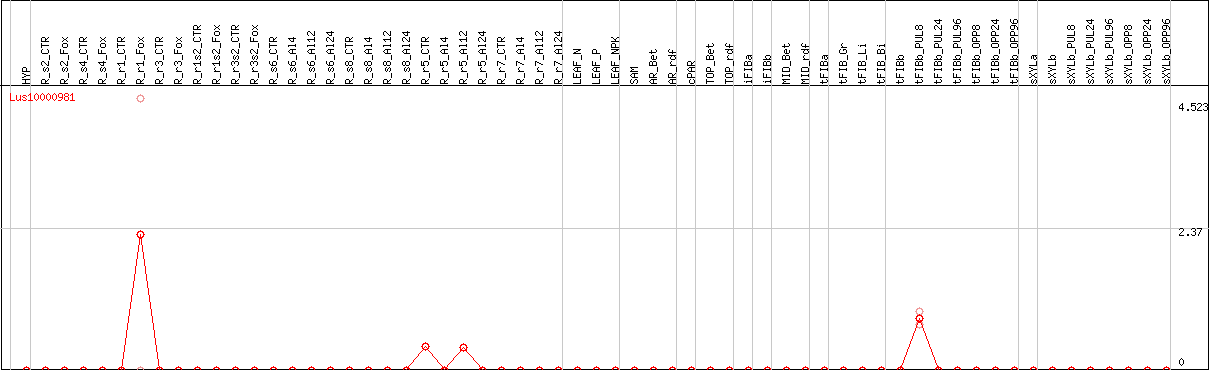
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000981 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.