External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G31100 85 / 3e-20
alpha/beta-Hydrolases superfamily protein (.1)
AT1G06250 79 / 4e-18
alpha/beta-Hydrolases superfamily protein (.1)
AT2G42690 66 / 2e-13
alpha/beta-Hydrolases superfamily protein (.1)
AT1G30370 45 / 4e-06
DLAH
DAD1-like acylhydrolase, alpha/beta-Hydrolases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013935
213 / 1e-69
AT4G18550 337 / 2e-113
Arabidopsis thaliana DAD1-like seeding establishment-related lipase, alpha/beta-Hydrolases superfamily protein (.1)
Lus10013936
138 / 1e-40
AT4G18550 345 / 2e-116
Arabidopsis thaliana DAD1-like seeding establishment-related lipase, alpha/beta-Hydrolases superfamily protein (.1)
Lus10000985
84 / 3e-20
AT4G18550 375 / 1e-129
Arabidopsis thaliana DAD1-like seeding establishment-related lipase, alpha/beta-Hydrolases superfamily protein (.1)
Lus10013932
84 / 5e-20
AT4G18550 427 / 5e-149
Arabidopsis thaliana DAD1-like seeding establishment-related lipase, alpha/beta-Hydrolases superfamily protein (.1)
Lus10017668
71 / 2e-15
AT4G18550 420 / 1e-145
Arabidopsis thaliana DAD1-like seeding establishment-related lipase, alpha/beta-Hydrolases superfamily protein (.1)
Lus10017975
65 / 3e-13
AT2G42690 524 / 0.0
alpha/beta-Hydrolases superfamily protein (.1)
Lus10041967
64 / 9e-13
AT2G42690 490 / 3e-173
alpha/beta-Hydrolases superfamily protein (.1)
Lus10015521
49 / 2e-07
AT1G30370 524 / 0.0
DAD1-like acylhydrolase, alpha/beta-Hydrolases superfamily protein (.1)
Lus10038526
47 / 6e-07
AT1G30370 539 / 0.0
DAD1-like acylhydrolase, alpha/beta-Hydrolases superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G026500
91 / 1e-22
AT4G18550 326 / 5e-109
Arabidopsis thaliana DAD1-like seeding establishment-related lipase, alpha/beta-Hydrolases superfamily protein (.1)
Potri.015G026400
91 / 2e-22
AT4G18550 330 / 5e-110
Arabidopsis thaliana DAD1-like seeding establishment-related lipase, alpha/beta-Hydrolases superfamily protein (.1)
Potri.011G064400
87 / 3e-21
AT4G18550 501 / 3e-177
Arabidopsis thaliana DAD1-like seeding establishment-related lipase, alpha/beta-Hydrolases superfamily protein (.1)
Potri.004G054800
82 / 2e-19
AT4G18550 516 / 0.0
Arabidopsis thaliana DAD1-like seeding establishment-related lipase, alpha/beta-Hydrolases superfamily protein (.1)
Potri.004G054600
81 / 7e-19
AT4G18550 384 / 1e-131
Arabidopsis thaliana DAD1-like seeding establishment-related lipase, alpha/beta-Hydrolases superfamily protein (.1)
Potri.003G101800
60 / 1e-11
AT2G42690 531 / 0.0
alpha/beta-Hydrolases superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10000983 pacid=23162254 polypeptide=Lus10000983 locus=Lus10000983.g ID=Lus10000983.BGIv1.0 annot-version=v1.0
ATGAGGATAAGGAACTTTCAAGATGTGGTTGTGTCAAACTTTCCGTCGAAGATTTTTGGATTTTTTGAGGTTGGGAACGAGTTGTTGATCCATTCGTTCT
CGTCCTACCTAAAGCCGTTGGTGAACCCTCATAGCTTGGAGGTGTACATGCATGGCTTAGCCGGGGTGGATGGCTTCGGGAAGTTCAAGCTGGTGGTTCA
TAGGGATGTTTCGTTACTCAACAAGTGGCTCGATGCGCTTAAGGACGAATATAAGGTTCCAAAAGAGTGGTGGATCGCGAAGAACTGGTCCATGGTCGAC
AGTGGTGTGTGGATGTTGCAGGAGGTTATGCCGCCACCTCCGGTGGCTCTTGTTTGA
AA sequence
>Lus10000983 pacid=23162254 polypeptide=Lus10000983 locus=Lus10000983.g ID=Lus10000983.BGIv1.0 annot-version=v1.0
MRIRNFQDVVVSNFPSKIFGFFEVGNELLIHSFSSYLKPLVNPHSLEVYMHGLAGVDGFGKFKLVVHRDVSLLNKWLDALKDEYKVPKEWWIAKNWSMVD
SGVWMLQEVMPPPPVALV
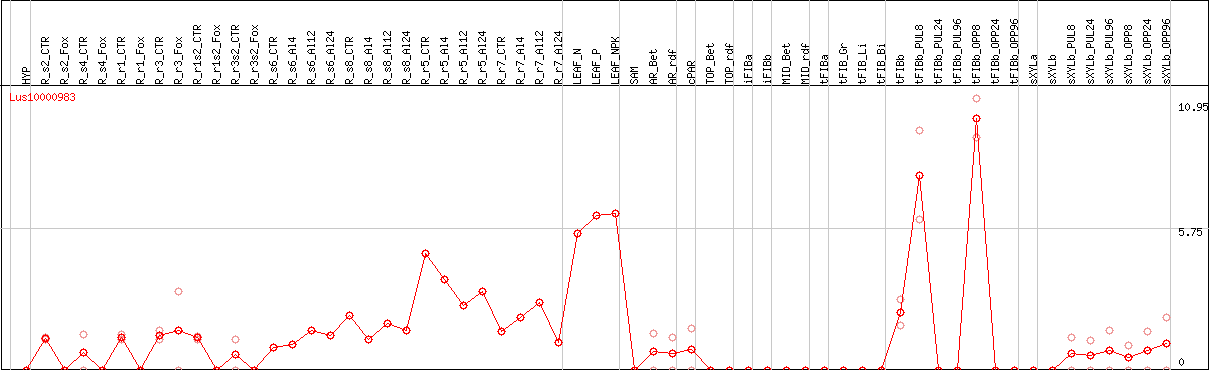
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10000983 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.