External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G53160 306 / 2e-107
RCAR3, PYL8
PYR1-like 8, regulatory components of ABA receptor 3 (.1.2)
AT1G01360 290 / 3e-101
PYL9, RCAR1
PYRABACTIN RESISTANCE 1-LIKE 9, regulatory component of ABA receptor 1 (.1)
AT4G01026 278 / 3e-96
RCAR2, PYL7
regulatory components of ABA receptor 2, PYR1-like 7 (.1)
AT4G27920 273 / 1e-94
RCAR4, PYL10
regulatory components of ABA receptor 4, PYR1-like 10 (.1)
AT5G05440 176 / 6e-56
RCAR8, PYL5
regulatory component of ABA receptor 8, PYRABACTIN RESISTANCE 1-LIKE 5, Polyketide cyclase/dehydrase and lipid transport superfamily protein (.1)
AT2G38310 173 / 6e-55
RCAR10, PYL4
regulatory components of ABA receptor 10, PYR1-like 4 (.1)
AT2G40330 171 / 1e-53
RCAR9, PYL6
regulatory components of ABA receptor 9, PYR1-like 6 (.1)
AT2G26040 166 / 4e-52
RCAR14, PYL2
regulatory components of ABA receptor 14, PYR1-like 2 (.1)
AT4G17870 157 / 6e-49
RCAR11, PYR1
regulatory component of ABA receptor 11, PYRABACTIN RESISTANCE 1, Polyketide cyclase/dehydrase and lipid transport superfamily protein (.1)
AT1G73000 149 / 2e-45
RCAR13, PYL3
regulatory components of ABA receptor 13, PYR1-like 3 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10039335
390 / 2e-140
AT5G53160 295 / 4e-103
PYR1-like 8, regulatory components of ABA receptor 3 (.1.2)
Lus10038818
320 / 1e-112
AT5G53160 287 / 1e-99
PYR1-like 8, regulatory components of ABA receptor 3 (.1.2)
Lus10014929
315 / 8e-111
AT5G53160 289 / 2e-100
PYR1-like 8, regulatory components of ABA receptor 3 (.1.2)
Lus10040916
266 / 6e-92
AT1G01360 265 / 1e-91
PYRABACTIN RESISTANCE 1-LIKE 9, regulatory component of ABA receptor 1 (.1)
Lus10014239
187 / 4e-60
AT2G38310 250 / 5e-85
regulatory components of ABA receptor 10, PYR1-like 4 (.1)
Lus10022675
186 / 6e-59
AT2G38310 249 / 1e-83
regulatory components of ABA receptor 10, PYR1-like 4 (.1)
Lus10012231
175 / 2e-55
AT2G38310 234 / 1e-78
regulatory components of ABA receptor 10, PYR1-like 4 (.1)
Lus10007530
161 / 5e-51
AT2G38310 198 / 1e-65
regulatory components of ABA receptor 10, PYR1-like 4 (.1)
Lus10024991
154 / 2e-47
AT2G26040 264 / 7e-91
regulatory components of ABA receptor 14, PYR1-like 2 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G000800
326 / 2e-115
AT5G53160 303 / 1e-106
PYR1-like 8, regulatory components of ABA receptor 3 (.1.2)
Potri.015G020500
326 / 2e-115
AT5G53160 304 / 7e-107
PYR1-like 8, regulatory components of ABA receptor 3 (.1.2)
Potri.003G139200
321 / 2e-113
AT5G53160 297 / 3e-104
PYR1-like 8, regulatory components of ABA receptor 3 (.1.2)
Potri.001G092500
319 / 8e-113
AT5G53160 296 / 7e-104
PYR1-like 8, regulatory components of ABA receptor 3 (.1.2)
Potri.002G169400
312 / 5e-110
AT1G01360 305 / 2e-107
PYRABACTIN RESISTANCE 1-LIKE 9, regulatory component of ABA receptor 1 (.1)
Potri.014G097100
311 / 1e-109
AT1G01360 301 / 6e-106
PYRABACTIN RESISTANCE 1-LIKE 9, regulatory component of ABA receptor 1 (.1)
Potri.016G125400
188 / 1e-60
AT2G38310 256 / 3e-87
regulatory components of ABA receptor 10, PYR1-like 4 (.1)
Potri.006G104100
186 / 7e-60
AT2G38310 249 / 2e-84
regulatory components of ABA receptor 10, PYR1-like 4 (.1)
Potri.018G054400
174 / 1e-55
AT2G26040 273 / 1e-94
regulatory components of ABA receptor 14, PYR1-like 2 (.1)
Potri.010G183900
171 / 6e-54
AT2G40330 243 / 8e-82
regulatory components of ABA receptor 9, PYR1-like 6 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0209
Bet_v_1_like
PF10604
Polyketide_cyc2
Polyketide cyclase / dehydrase and lipid transport
Representative CDS sequence
>Lus10001059 pacid=23141285 polypeptide=Lus10001059 locus=Lus10001059.g ID=Lus10001059.BGIv1.0 annot-version=v1.0
ATGAATTTCGATACTAGCAATGGAGCGGCGACCACGACTGCCAAGGGGCTAGGGAGCGTGGAGACCGAGTCCATCAGGAGACACCATAGGCATGATCCTG
GCGAGAATCAGTGCAGCTCCGTTCTCATTAAGCACATCAAAGCTCCGGTTCATCTGGTTTGGTCCCTGGTTAGGAGATTCGATCAACCGCAGAAGTACAA
GCCATTCATCAGCAGATGCGTTGCTCGTGGGAATCTCGAGATTGGGAGTCTTAGAGAGATTGATGTTAAATCAGGACTTCCAGCTACCACAAGTACTGAA
AGATTAGAGCTTCTTGATGATGAGGAGCATATCCTCAGCATGCGCATTGTTGGCGGAGATCACAGACTAAAGAACTATTCTTCAATAGTTTCCCTACACC
CGGAGATTCTCGATGGAAGACCAGGGACATTAGTGATCGAATCATTTGTGGTGGATGTACCAGATGGGAACACCAAGGACGAGACCTGCTACTTTGTTGA
AGCTCTCATCAAATGCAACCTCAAGTCTCTGGCTGATGTATCTGAGCGGCTTGCGGTACAGGACAGGACCGAACCCATCGATCGTAACTGA
AA sequence
>Lus10001059 pacid=23141285 polypeptide=Lus10001059 locus=Lus10001059.g ID=Lus10001059.BGIv1.0 annot-version=v1.0
MNFDTSNGAATTTAKGLGSVETESIRRHHRHDPGENQCSSVLIKHIKAPVHLVWSLVRRFDQPQKYKPFISRCVARGNLEIGSLREIDVKSGLPATTSTE
RLELLDDEEHILSMRIVGGDHRLKNYSSIVSLHPEILDGRPGTLVIESFVVDVPDGNTKDETCYFVEALIKCNLKSLADVSERLAVQDRTEPIDRN
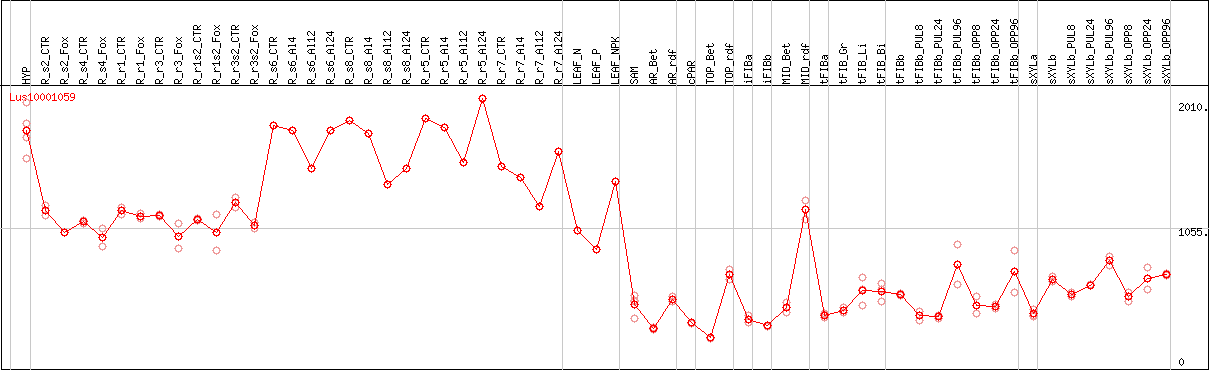
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10001059 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.