External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G05490 201 / 8e-59
CHR31
chromatin remodeling 31 (.1)
AT3G24340 181 / 1e-51
CHR40
chromatin remodeling 40 (.1)
AT5G20420 108 / 2e-26
CHR42
chromatin remodeling 42 (.1)
AT3G42670 108 / 2e-26
CLSY1, CLSY, CHR38
CLASSY 1, CLASSY1, chromatin remodeling 38 (.1)
AT2G16390 105 / 2e-25
DMS1, CHR35, DRD1
DEFECTIVE IN RNA-DIRECTED DNA METHYLATION 1, DEFECTIVE IN MERISTEM SILENCING 1, SNF2 domain-containing protein / helicase domain-containing protein (.1)
AT2G21450 98 / 8e-23
CHR34
chromatin remodeling 34 (.1)
AT3G32330 77 / 6e-16
DNA repair protein-related (.1)
AT1G08600 67 / 1e-12
ATRX, CHR20
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4)
AT3G32280 61 / 3e-10
ATP-dependent helicase family protein (.1)
AT3G31900 55 / 2e-08
ATP-dependent helicase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10040136
491 / 1e-168
AT1G05490 430 / 2e-131
chromatin remodeling 31 (.1)
Lus10003003
344 / 5e-111
AT1G05490 579 / 0.0
chromatin remodeling 31 (.1)
Lus10011033
348 / 7e-111
AT1G05490 590 / 0.0
chromatin remodeling 31 (.1)
Lus10009840
114 / 1e-28
AT5G20420 1283 / 0.0
chromatin remodeling 42 (.1)
Lus10041963
113 / 5e-28
AT2G16390 1018 / 0.0
DEFECTIVE IN RNA-DIRECTED DNA METHYLATION 1, DEFECTIVE IN MERISTEM SILENCING 1, SNF2 domain-containing protein / helicase domain-containing protein (.1)
Lus10017973
112 / 6e-28
AT2G16390 994 / 0.0
DEFECTIVE IN RNA-DIRECTED DNA METHYLATION 1, DEFECTIVE IN MERISTEM SILENCING 1, SNF2 domain-containing protein / helicase domain-containing protein (.1)
Lus10040956
110 / 5e-27
AT5G20420 1316 / 0.0
chromatin remodeling 42 (.1)
Lus10042070
107 / 6e-26
AT2G16390 833 / 0.0
DEFECTIVE IN RNA-DIRECTED DNA METHYLATION 1, DEFECTIVE IN MERISTEM SILENCING 1, SNF2 domain-containing protein / helicase domain-containing protein (.1)
Lus10032785
61 / 3e-10
AT1G08600 1252 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G050200
256 / 3e-78
AT1G05490 592 / 0.0
chromatin remodeling 31 (.1)
Potri.008G073500
117 / 1e-29
AT3G42670 1120 / 0.0
CLASSY 1, CLASSY1, chromatin remodeling 38 (.1)
Potri.009G120700
113 / 3e-28
AT2G16390 938 / 0.0
DEFECTIVE IN RNA-DIRECTED DNA METHYLATION 1, DEFECTIVE IN MERISTEM SILENCING 1, SNF2 domain-containing protein / helicase domain-containing protein (.1)
Potri.010G183832
109 / 2e-27
AT3G42670 656 / 0.0
CLASSY 1, CLASSY1, chromatin remodeling 38 (.1)
Potri.004G159000
94 / 1e-21
AT2G16390 811 / 0.0
DEFECTIVE IN RNA-DIRECTED DNA METHYLATION 1, DEFECTIVE IN MERISTEM SILENCING 1, SNF2 domain-containing protein / helicase domain-containing protein (.1)
Potri.013G048500
62 / 7e-11
AT1G08600 1603 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4)
Potri.019G021500
61 / 2e-10
AT1G08600 1612 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4)
Potri.004G141500
52 / 2e-07
AT3G19210 1259 / 0.0
homolog of RAD54 (.1.2)
Potri.003G110100
50 / 1e-06
AT5G44800 1988 / 0.0
PICKLE RELATED 1, chromatin remodeling 4 (.1)
Potri.013G068400
50 / 1e-06
AT3G54280 2717 / 0.0
ROOT GROWTH DEFECTIVE 3, DNA binding;ATP binding;nucleic acid binding;binding;helicases;ATP binding;DNA binding;helicases (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00176
SNF2_N
SNF2 family N-terminal domain
Representative CDS sequence
>Lus10001090 pacid=23148932 polypeptide=Lus10001090 locus=Lus10001090.g ID=Lus10001090.BGIv1.0 annot-version=v1.0
ATGAAGCTTTTTCCTCACTGCAGACCAGTCATAATCGTCCCTTGCAGTATGCTACTATCTTGGGAAGCCGAGTTCGAGGCCTTGGGAGTCGACATTCCTC
GTCACTATCTGAATGATCCAAAGCTATCTGGCAAAAAGAGTGTCGCAGCTACAAAGTTGTTGGACGATCACAGGAGTTTACAAGCTGTCAGGTGGGTGAA
GCTATATTCCTGGCAAAAGGAGTATGGTATTTTGGTATTAAGCTACAAGCTTTTTGAAGAACTTGTTGGCAAAGGGATGAGAAAGAAGAGGAAAACCGAA
AGCAGAAAACAAAAGGGAAGGTCTGTAGATGAAAGAATAAAGGACGTCCTGCTTGATGTTCCTGGCTTGTTGGTATTTGATGAAGGCCATACAGCTCGCA
ATAACAGTAGCCTCATCTGGAGAGCTTTGTCCAAAGTAAAAACAGAGAAGCGGGTTGTTCTTTCGGGGACTCCATTTCAAAACAATTTTGTAGAGTTCTT
CAATACTCTCTGGATGGCAAGACCTAACTTTGTAGACAACATATCATCCAGCACATTTGAAACATCTCATAGAAAACGTGGTCGCAGGGCGGTGAATGAA
GCCACGGAGAAATGGCAGTCTTTGACGAGTTATCTCGGCAAAGATACTGATGATTGGCTAAAAGTCATGAAACTGCAAGAGCTTAGAGATGGGATCAGCC
CCTTTGTGCATGTCTACAAGGGCAATATACTCCAAGAAAAGCTCCCAGGTTTAAGAGACTCTGTGGTTATTTTACAGCCAGGCCCTTTCCCGGGATGA
AA sequence
>Lus10001090 pacid=23148932 polypeptide=Lus10001090 locus=Lus10001090.g ID=Lus10001090.BGIv1.0 annot-version=v1.0
MKLFPHCRPVIIVPCSMLLSWEAEFEALGVDIPRHYLNDPKLSGKKSVAATKLLDDHRSLQAVRWVKLYSWQKEYGILVLSYKLFEELVGKGMRKKRKTE
SRKQKGRSVDERIKDVLLDVPGLLVFDEGHTARNNSSLIWRALSKVKTEKRVVLSGTPFQNNFVEFFNTLWMARPNFVDNISSSTFETSHRKRGRRAVNE
ATEKWQSLTSYLGKDTDDWLKVMKLQELRDGISPFVHVYKGNILQEKLPGLRDSVVILQPGPFPG
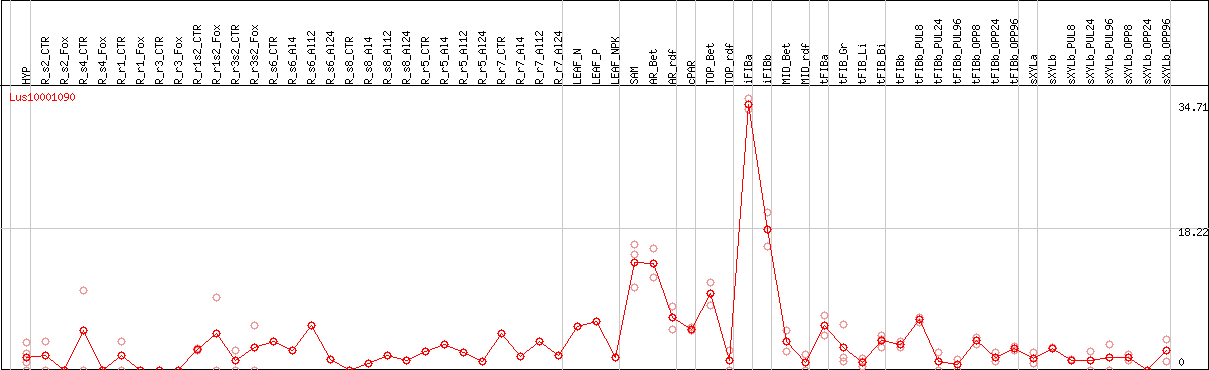
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10001090 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.