External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G03560 130 / 2e-38
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT1G74850 64 / 2e-13
PDE343, PTAC2
PIGMENT DEFECTIVE 343, plastid transcriptionally active 2 (.1)
AT4G38150 62 / 9e-13
Pentatricopeptide repeat (PPR) superfamily protein (.1), Pentatricopeptide repeat (PPR) superfamily protein (.2)
AT3G25210 61 / 2e-12
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G31840 60 / 5e-12
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT5G61990 59 / 1e-11
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G05670 58 / 3e-11
Pentatricopeptide repeat (PPR-like) superfamily protein (.1), Pentatricopeptide repeat (PPR-like) superfamily protein (.2)
AT1G06580 56 / 1e-10
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT4G19440 56 / 1e-10
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT5G14770 56 / 2e-10
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013853
65 / 1e-13
AT4G38150 304 / 2e-102
Pentatricopeptide repeat (PPR) superfamily protein (.1), Pentatricopeptide repeat (PPR) superfamily protein (.2)
Lus10026570
64 / 3e-13
AT4G38150 303 / 5e-102
Pentatricopeptide repeat (PPR) superfamily protein (.1), Pentatricopeptide repeat (PPR) superfamily protein (.2)
Lus10013972
61 / 5e-12
AT5G46100 520 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10015397
61 / 5e-12
AT5G46100 593 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10025013
59 / 2e-11
AT1G05670 161 / 1e-43
Pentatricopeptide repeat (PPR-like) superfamily protein (.1), Pentatricopeptide repeat (PPR-like) superfamily protein (.2)
Lus10013841
58 / 3e-11
AT5G65560 868 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10026558
58 / 3e-11
AT5G65560 881 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10013077
58 / 4e-11
AT5G40400 587 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10038974
58 / 4e-11
AT1G74580 911 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G209500
62 / 8e-13
AT4G38150 314 / 4e-106
Pentatricopeptide repeat (PPR) superfamily protein (.1), Pentatricopeptide repeat (PPR) superfamily protein (.2)
Potri.010G035700
60 / 5e-12
AT1G05670 922 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1), Pentatricopeptide repeat (PPR-like) superfamily protein (.2)
Potri.003G105700
59 / 2e-11
AT4G19440 758 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.005G050180
58 / 3e-11
AT1G12700 488 / 2e-164
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.002G248100
57 / 5e-11
AT3G25210 506 / 2e-180
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G090600
57 / 8e-11
AT5G08310 851 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G046000
57 / 1e-10
AT1G63130 405 / 9e-133
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.001G297300
56 / 1e-10
AT5G02860 1129 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.005G014500
56 / 1e-10
AT1G09820 696 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Potri.009G092100
56 / 2e-10
AT5G02860 1146 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10001125 pacid=23161785 polypeptide=Lus10001125 locus=Lus10001125.g ID=Lus10001125.BGIv1.0 annot-version=v1.0
ATGCCTGATGTAGCAGCTTACACAGCGGTGATAGAAGTCTATGCCAAGGCCGGTGGGCAGTCAAAAGAAGCCCTGAAGGTGTTCATGAGAATGCTCGCCT
CTGGAGTTGCTCCCAATGCATACACTTACAGTGTGTTGGTCAAGGGACTGGCTGCCGATGCCAAAATTGGTGATGGTAAGAAGTATCTGCTGGAAATGAT
GGGGAAAGGGATGAGGCGGAATGCTGGGACTTACTGTGCGGTGTTTGAAGGTTTGGTGAGAGGAGAGAGTTGA
AA sequence
>Lus10001125 pacid=23161785 polypeptide=Lus10001125 locus=Lus10001125.g ID=Lus10001125.BGIv1.0 annot-version=v1.0
MPDVAAYTAVIEVYAKAGGQSKEALKVFMRMLASGVAPNAYTYSVLVKGLAADAKIGDGKKYLLEMMGKGMRRNAGTYCAVFEGLVRGES
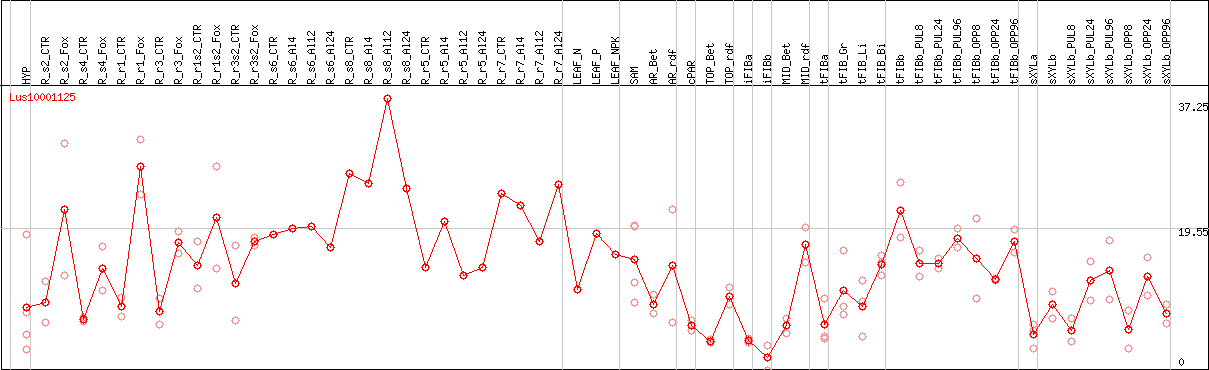
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10001125 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.