External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G17050 62 / 9e-11
disease resistance protein (TIR-NBS-LRR class), putative (.1)
AT3G51560 59 / 1e-09
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT4G16860 56 / 8e-09
RPP5, RPP4
recognition of peronospora parasitica 4, Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT1G69550 56 / 1e-08
disease resistance protein (TIR-NBS-LRR class) (.1)
AT5G46520 55 / 2e-08
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G46510 55 / 2e-08
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G45250 52 / 2e-07
RPS4
RESISTANT TO P. SYRINGAE 4, Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G22690 50 / 6e-07
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G46260 50 / 1e-06
disease resistance protein (TIR-NBS-LRR class) family (.1)
AT3G44480 50 / 1e-06
COG1, RPP10, RPP1
recognition of peronospora parasitica 1, Disease resistance protein (TIR-NBS-LRR class) family (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10042778
244 / 4e-76
AT4G12010 185 / 1e-48
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10024150
204 / 9e-61
AT1G69550 186 / 9e-49
disease resistance protein (TIR-NBS-LRR class) (.1)
Lus10001537
182 / 5e-53
AT5G44510 175 / 1e-45
target of AVRB operation1 (.1)
Lus10015648
183 / 3e-52
AT4G12010 420 / 1e-126
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10010223
178 / 9e-51
AT5G17680 218 / 6e-58
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10017420
167 / 9e-48
AT1G69550 164 / 1e-42
disease resistance protein (TIR-NBS-LRR class) (.1)
Lus10017418
152 / 2e-41
AT5G44510 335 / 1e-96
target of AVRB operation1 (.1)
Lus10015649
137 / 1e-39
AT1G69550 75 / 1e-15
disease resistance protein (TIR-NBS-LRR class) (.1)
Lus10010221
132 / 1e-34
AT5G17680 369 / 4e-109
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G069200
70 / 2e-13
AT5G17680 590 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.005G031101
62 / 7e-11
AT5G17680 557 / 8e-178
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.019G001971
62 / 1e-10
AT5G36930 427 / 3e-130
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G068300
61 / 2e-10
AT5G17680 580 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.019G068200
61 / 2e-10
AT5G17680 629 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.T009323
61 / 3e-10
AT5G36930 449 / 2e-138
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.003G014200
57 / 5e-09
AT5G36930 673 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G070700
57 / 6e-09
AT5G17680 722 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.019G016425
56 / 1e-08
AT5G36930 650 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.T001500
55 / 2e-08
AT5G36930 641 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
PFAM info
Representative CDS sequence
>Lus10001281 pacid=23166378 polypeptide=Lus10001281 locus=Lus10001281.g ID=Lus10001281.BGIv1.0 annot-version=v1.0
ATGTACACCTCTGCATATGCTAAGGTGAAGCATCTAATTGATAGAACCCTCTTAATTTGTGAAGGAAAGAATATCCAAGTGCATGGTCTGCTTAAGGAAC
CGAAGTGGGGAAACGCAGCAGCCATAGAAATGGTGCTTCCGAAAAGGAAAAAAGCCAAAGTGGTAGACATTCCTTGGATTGGCAGTAATCCAAGCTTCAA
ATGCCTGCCATCTAGCATCAGCAATACCAGTTCTCTCAAGTCGCTTAATTTCAAAAAGACTTCGAGTATTCTCGAATTGACGAGCCTTTACTCAATTGAT
GTGAGCTACTGCAAGCGTTTCGAGTCAATGCCCACGGGTATCCATAAGCTTTCCAAGTTACGACGATTGGTAGTGTCCGGCTGTGACCGCATTCGATCTC
TGCCACAACTCCCACCAAATCTGACAGAATTGTATGTAAATGGTTGCAAGTCTTTACAAGCTCTATCAAGCAACACAGGTAAGCTGTTGCATCTGAATAA
ACTCTATTCTGAAGAATGCCCGCAATTGGGACAGACTATACCTGTCGAAATTTTGGCAAACTTCCTCGTTCATGCCAGCTTGTCTCCGGCACCTTGTGTG
ATGTACTGGGAAGTTGTATGTTCAAGAAGCGAGCTTCCGGAATGGTTAACGTACAAAAGCATGAACACGATGAAGGAGGAGGGTTGCAGTGTAGAGGTGC
AGTTGCCTCTTCCCAGCGACCGTGACCAACCAATGATTATGAACGGAATTGCATTCGCCGCTGTGTTTCCTTGCAACTCGATGGTGGAAATGAAATGA
AA sequence
>Lus10001281 pacid=23166378 polypeptide=Lus10001281 locus=Lus10001281.g ID=Lus10001281.BGIv1.0 annot-version=v1.0
MYTSAYAKVKHLIDRTLLICEGKNIQVHGLLKEPKWGNAAAIEMVLPKRKKAKVVDIPWIGSNPSFKCLPSSISNTSSLKSLNFKKTSSILELTSLYSID
VSYCKRFESMPTGIHKLSKLRRLVVSGCDRIRSLPQLPPNLTELYVNGCKSLQALSSNTGKLLHLNKLYSEECPQLGQTIPVEILANFLVHASLSPAPCV
MYWEVVCSRSELPEWLTYKSMNTMKEEGCSVEVQLPLPSDRDQPMIMNGIAFAAVFPCNSMVEMK
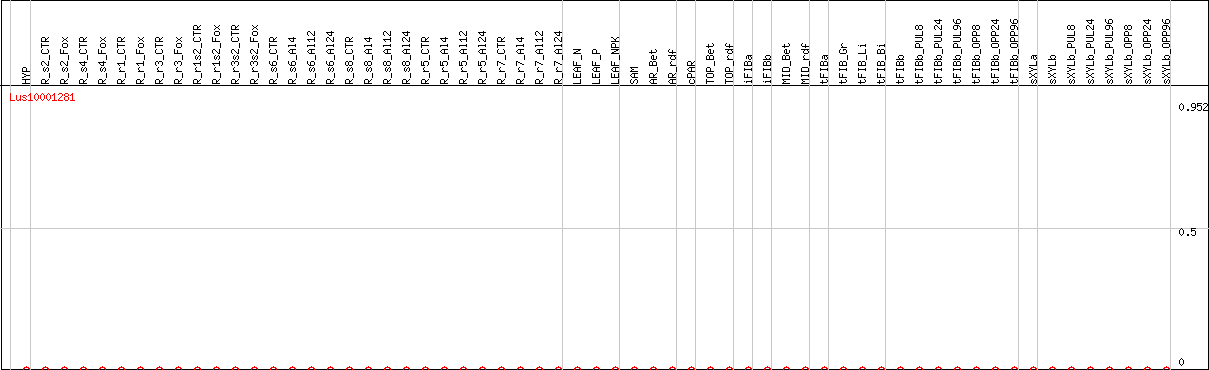
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10001281 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.