Lus10001424 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10001424 pacid=23145931 polypeptide=Lus10001424 locus=Lus10001424.g ID=Lus10001424.BGIv1.0 annot-version=v1.0
ATGGCAAGCACCAGCGGAACCAAGAGCGACCTCCCGAGGTCCTTCTCCCGCGCTTCTACCATGATGGTAGACTTGCGGGATGGAGACGCTTCGGTGATTG
ACAGTGAGGTTGTCCCGTCTTCTTTGGCTTCAATTGCTCCAATTCTCCGTGTTGCTAACGAGATTCAGGATGATAATCCCAGAGTCGCCTATCTATGTCG
GTTCCATGCATTTGAGAAGGCTCATAAGATGGACCCAACATCAAGTGGACGTGGGGTCCGTCAATTCAAGACGTATCTGCTGCACAGATTAGAGAGGGAA
GAAAGTGAAACAGAACCCCGTCTTGCAAGAACTGATCCTAGAGAAATTCAATTATATTATCAGAAGTTTTATGAGGAAAATATAAGAGATGGACAATATA
CAAAAAGACCGGAGGAGATGGCCAAGATTCTTCAGATTGCATCAGTACTATATGACGTGCTGAAAACAGTGGTACCTGCTTCCAGAGTCGATGAAGAGAC
ACATAAATATGCTAAGGATGTTGAGAAGAAAAGGGAGCAATATGAGCACTATAACATTCTTCCATTGTATTCCGTTGGCTTAAAACCGGCAATAATGGAA
CTTCCCGAGATCAAAGCTGCATTTAATGCTGTACGCAATGTGGACAATCTTCCAATGCCTAGAATCAGTTCAATTTATAGTGCCATTCATGATGTGCCAA
CTGGAAAGATCAAACCTGTCAATGATATACTTGAATGGCTTTCTTCACTTTTTGGTTTCCAGAAAGGAAATGTGGCAAATCAGAGAGAGCACCTAATATT
GCTGTTAGCCAATATGGACACGAGGAAAAGGAGCCTTGACGATTACACAGAGTTAAATACTAGCACAATACAACAACTGGTGAACAAAACTTTCAAAAAT
TATCTCTCCTGGTGTCACTATTTGCGAGTTGAAAGCAATCTTAGGTTCCCAGGAGGATTTGATAAACCACAACTGGAGCTTATCTACATTGGACTGTATC
TACTAATATGGGGCGAAGCTTCAAACGTACGCTTCATGCCGGAATGTATTTGCTATATCTTCCACAATATGGCATATGAAGTTTTTGGTGTATTGTTCCG
CAATTTACATCCTGTTAGTGGAGAAACCTATGAATCAGCAGCTCCTGATGACGAAGCCTTTCTGCGGAACGTAGTTACTCCGATATACCAGGTCGTGCGT
AAGGAAGCTAGGAGAAACAAAAGTGGGAAAGCGAGTCATTCAAGGTGGAGAAATTATGATGATCTAAATGAGTATTTCTGGTCTAACAGATGTCTAAGGA
TAAAGTGGCCAATGGATCTCAAAACTGACTTTTTTGTTCATTCAGATGAGGTTTTGCCAGCAAATGAGAGATCTACTCAGAGAGGCAGCGGGAGGAGGAA
GCCCAAGACTAATTTTGTTGAAGTCCGTACTTTCTGGCACCTTTATAGAACCTTCGACAGAATGTGGATATTTTTCATATTAGCTTTTCAGGCTATGTTT
ATCATTGCTTGGAATGCTGGATCTCTTGCTGGCATTTTTTACCCAGATGTCTTCAGAAGTGTTTTAAGCATCTTTATCACTGCTGCTTTTCTCAATCTTC
TGCGAGCTGTTCTTGATATAATCCTCAATCTTTACGCATGGAGGAGCTTAAAGTTTACACAAATAGTGCGGTATTTGCTCAAATTTGCAGTAGCAGCAGC
CTGGGCTGTTGCTATGCCGATTGCTTATGCCAAATCTGTTCAGAACCCAACAGGACTTATCAAATTTTTCAGCAGCTGGGCTAGTGACTGGCAGAATCAA
TCTCTTTATAATTATGCTGTTGCTATTTATTTGATACCCAACATCCTGTCTACTCTGTTGTTTGTGTTTCCGCCATTTCAAAGGACAATGGAAAAATCAA
ACTGGCGGATATTCACTCTTCTTCTATGGTGGGCTCAGGCAAGAATATTTAGCACTTTAAATTCTCGGTATACTCTCTTTTGGATCTTACTTCTAATAAG
CAAGTTAGCATTTAGCTATTATATTGAGATACTGCCTCTTGTTGAACCAACAAAGCTGATCATGGATATGAGTATCGATAATTATCAGTGGCATGAGTTC
TTCCCACATGCAAAGCACAACCTTGGCGCTGTTATTGCAATATGGGCACCCATTGTCCTGGTATATTTTATGGATACACAAATATGGTACGCAATATTTT
CAACTATATTCGGTGGGATTCGTGGAGCCTTCAGCCATCTGGGTGAGATAAGGACTCTTGGTATGCTACGGTCCAGATTTGAGGACGTGCCCTTGGCTTT
CAGTAACTGCCTTGTGCCATCATCAAAGAAAAGAGGCAAAAGGAAACATTTGGATGAGTCAGCTGAAAGGCAAAGCATCGCCAATTTTTCTCATGTCTGG
AATGAGATCATTAACTCTATGAGATTGGAGGATCTGATCAGCAATGAGTTGAGACCCTTCTACCGTCATTTCATCTACGAAAGAGATTTACTACTGGTAC
CATCTTCCTCTAATGATGTATCAGTCATCCAATGGCCTCCATTTTTGCTTGCAAGCAAGATTCCTATCGCACTGGACATGGCAAAAGATTTCAAGGGAAC
CAATGGTGCTGATCTATTCAGGAGAATGGATGAGTATATGCGGTTTGCTGTTATTGAATTTTACGAGACCATTAGAGACATAATTTACAGCTTGCTGCAA
GATGATTCAGACAGGATGATTATAAGACAGATTTGTTATGAAGTTGACGTGAGCATACAACAACAGAGGTTTCTTCAAGAATTCCGAACGAGTGGGTTGC
CTCCGCTTAGTGATAAATTGGAGAAGTTTTTGCACACCTTGGTTAGCAATTATGACGATCCTGAAATGTTTAAAGCTCAAATAATAAATATTATCCAAGA
TATTATAGAAATTATTGTTCAAGATGTTATGATTCGGGGCCGTGAAATCCTTGAAAGAGCTCACTATGCCACAGAAGATGACGAGAGTGTCAAGAAAGAG
CAGAGGTTTGGTAAGATAAACATTGACCTCATACGAAACAAAACTTGGAGAGAGAAGGTTGTTAGACTTCATTTGCTCTTGACAACGAAGGAATCTGCAA
TAAATGTTCCATCCAACTTAGATGCGCGTCGCCGTATCACATTCTTTGCAAATTCCTTGTTCATGCACATGCCAAGTGCTCCAAAAGTGAGGGACATGCT
CTCTTTTAGTGTTTTGACTCCTTATTACAAAGAAGATGTCCTCTACTCAGAGGAGGAGCTTAATCATGAAAATGAAGATGGCATCACAATTTTGTTTTAT
CTTCAAACGATATATCGCGATGAGTGGAGAAATTTCGAGGAAAGAATTTCAGGTTCCGCTCTAAAGGAAAAGGCAGAGCTTACTCGTCACTGGGTATCCT
ATAGGGCACAGACACTTTCAAGAACAGTTAGGGGGATGATGTACTACAGACGGGCTCTTGAACTCCAGTACTTGCTGGAATTTGCAGGAGAAAATGCTAT
TATCAATGGCCTCCCCAGCATGGAACTCAGTAATGAAGATAGATCTCTGTCAGATCGTGCACAGGCTCTTGCTGATCTGAAGTTTACCTATGTGGTTTCA
TGTCAGATTTACGGTGCTCAGAAAAAGTCCAGCGATGGTCGAGACCGGAGCTGTTACAATAACATTCTGAATCTGATGTTAAAGTACCCATCACTACGTG
TTGCGTACATTGATGAGAGAGAAGAGACAGTGAATGGAAAGCAGCAGAAAGTTTACTATTCTGTGCTTGTTAAAGGGGGTGATAAGTTGGATGAGGAAAT
ATACAGAATAAAGCTTCCTGGGCCACCTACAGAAATAGGTGAAGGTAAACCTGAAAATCAGAACCACGCAATTATTTTCACCCGTGGAGAGGCTTTGCAA
ACCATTGATATGAATCAGGATAACTACTTCGAAGAAGCTTTCAAAATGAGAAATGTGTTGGAGGAGTTCTTAACACCTCGCCATGGGCCACGCAAGCCAA
CAATCCTGGGCCTCAGGGAGCATATATTTACTGGCAGTGTTTCATCGCTTGCCTGGTTTATGTCCAACCAAGAAACAAGCTTCGTAACGATTGGCCAACG
CATTTTGGCCAACCCTTTAAGGGTACGTTTCCACTATGGTCATCCTGACATATTTGACCGGCTCTTCCATATAACGAGGGGTGGGATCAGCAAAGCTTCA
AAAATCATAAACCTGAGCGAAGATATATTTTCAGGATTCAATTCTACATTACGTGGTGGATATATAACACATCACGAGTACATACAAGTGGGCAAAGGAC
GTGATGTTGGGATGAATCAGATTTCTCTTTTTGAAGCAAAGGTTGCAAACGGTAATGGGGAGCAGACTCTTAGCCGTGATGTATATCGTCTTGGGCGCCG
ATTCGACTTCTACAGAATGCTGTCTTTCTATTTTACAACAGTTGGTTTTTATTTCAGTAGCATGATAACCGTGATTACGGTGTACATATTTTTGTATGGA
CGTATGTACATGGTAATGAGTGGAGTAGAGGCGGAGATTCTGACGAATCCAACCATTCGTCAAAGCAATGCACTTGAACAAGCTTTGGCCACTCAGTCCA
TATTTCAGTTAGGTTTGCTACTAGTCTTGCCAATGGTCATGGAAATCGGCCTGGAGAAAGGATTTCGCTCTGCTTTAGGTGATTTCATCATTATGCAGCT
GCAGCTAGCCTCTGTCTTCTTCACATTTCAGCTTGGAACTAAAGCTCATTACTACGGCAAGACAATTCTGCATGGTGGTTCCAAATATAGGGCCACTGGA
CGTGGGTTCGTTGTGTTCCATGCTAAGTTTGCAGAAAACTATAGGCTATACTCACGGAGTCACTTCGTGAAGGGATTAGAGTTGACCATGTTATTGGTCA
TATATGAAGTCTATGGGGAATCTTATCGCAGTTCAAGTCTGTACTTCTTCATTACCTTCTCCATGTGGTTCCTTGTTGGTTCCTGGTTGTTTGCACCATT
CGTTTTTAATCCATCTGGCTTTGATTGGCAAAAGACCGTGGATGATTGGACTGATTGGAAACGTTGGATGGGGAATCGTGGTGGGATTGGGATTCCACCG
GAAAAAAGTTGGGAATCTTGGTGGGACGGTGAACAGGAACATCTGAAACACACAAACCTAAGGGGTCGGATACTGGACATAATTCTCGCCTTTCGCTTCT
TTGTCTACCAGTATGGCATCGTCTACCACCTTGACATAGCTCACCGAATCAAAACTCTACTGGCACAGAGTATCCCTTACGTTACGTTACGAGCTTATAT
ATATGTATGTGTGATTCTCCTGATTTCTGGTCCTCAGGTTTATGGACTTTCGTGGGTGGTGATGATCACTGCTCTATTAGTCTTCAAGATGGTGTCAATG
GGGAGAAGGAAATTCGGCACAGATTTCCAGCTAACATTCAGAATCCTCAAAGCACTTCTCTTCATGGGTTTCCTGTCCGTCTTAACTGTGTTATTCGTTG
TCTGCAACCTGACGATAACGGATCTGTTTGCTTCCATCCTCGCCTTCCTTCCCACTGGATGGGCTATCCTTCTCATTGGGCAAGCATGCAGGGGACTGTT
CAAGGCGATTGGGCTGTGGGATTCAATAAAGGAGCTGGGAAGAGGGTACGAATACATAATGGGACTGTTGATCTTCAGCCCCACAGCCATATTGTCGTGG
TTCCCTTTCGTGAACGAGTTCCAAACTCGGCTGCTCTTCAACCAAGCTTTCAGTAGAGGCCTCCAGATTTCAATGATTCTTCAGGGCAGGAAAGATAAAG
ACGATGCAGCTGCCAAGAAAGATAAGCCTCCCCCGCCAAAATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10001424 pacid=23145931 polypeptide=Lus10001424 locus=Lus10001424.g ID=Lus10001424.BGIv1.0 annot-version=v1.0
MASTSGTKSDLPRSFSRASTMMVDLRDGDASVIDSEVVPSSLASIAPILRVANEIQDDNPRVAYLCRFHAFEKAHKMDPTSSGRGVRQFKTYLLHRLERE
ESETEPRLARTDPREIQLYYQKFYEENIRDGQYTKRPEEMAKILQIASVLYDVLKTVVPASRVDEETHKYAKDVEKKREQYEHYNILPLYSVGLKPAIME
LPEIKAAFNAVRNVDNLPMPRISSIYSAIHDVPTGKIKPVNDILEWLSSLFGFQKGNVANQREHLILLLANMDTRKRSLDDYTELNTSTIQQLVNKTFKN
YLSWCHYLRVESNLRFPGGFDKPQLELIYIGLYLLIWGEASNVRFMPECICYIFHNMAYEVFGVLFRNLHPVSGETYESAAPDDEAFLRNVVTPIYQVVR
KEARRNKSGKASHSRWRNYDDLNEYFWSNRCLRIKWPMDLKTDFFVHSDEVLPANERSTQRGSGRRKPKTNFVEVRTFWHLYRTFDRMWIFFILAFQAMF
IIAWNAGSLAGIFYPDVFRSVLSIFITAAFLNLLRAVLDIILNLYAWRSLKFTQIVRYLLKFAVAAAWAVAMPIAYAKSVQNPTGLIKFFSSWASDWQNQ
SLYNYAVAIYLIPNILSTLLFVFPPFQRTMEKSNWRIFTLLLWWAQARIFSTLNSRYTLFWILLLISKLAFSYYIEILPLVEPTKLIMDMSIDNYQWHEF
FPHAKHNLGAVIAIWAPIVLVYFMDTQIWYAIFSTIFGGIRGAFSHLGEIRTLGMLRSRFEDVPLAFSNCLVPSSKKRGKRKHLDESAERQSIANFSHVW
NEIINSMRLEDLISNELRPFYRHFIYERDLLLVPSSSNDVSVIQWPPFLLASKIPIALDMAKDFKGTNGADLFRRMDEYMRFAVIEFYETIRDIIYSLLQ
DDSDRMIIRQICYEVDVSIQQQRFLQEFRTSGLPPLSDKLEKFLHTLVSNYDDPEMFKAQIINIIQDIIEIIVQDVMIRGREILERAHYATEDDESVKKE
QRFGKINIDLIRNKTWREKVVRLHLLLTTKESAINVPSNLDARRRITFFANSLFMHMPSAPKVRDMLSFSVLTPYYKEDVLYSEEELNHENEDGITILFY
LQTIYRDEWRNFEERISGSALKEKAELTRHWVSYRAQTLSRTVRGMMYYRRALELQYLLEFAGENAIINGLPSMELSNEDRSLSDRAQALADLKFTYVVS
CQIYGAQKKSSDGRDRSCYNNILNLMLKYPSLRVAYIDEREETVNGKQQKVYYSVLVKGGDKLDEEIYRIKLPGPPTEIGEGKPENQNHAIIFTRGEALQ
TIDMNQDNYFEEAFKMRNVLEEFLTPRHGPRKPTILGLREHIFTGSVSSLAWFMSNQETSFVTIGQRILANPLRVRFHYGHPDIFDRLFHITRGGISKAS
KIINLSEDIFSGFNSTLRGGYITHHEYIQVGKGRDVGMNQISLFEAKVANGNGEQTLSRDVYRLGRRFDFYRMLSFYFTTVGFYFSSMITVITVYIFLYG
RMYMVMSGVEAEILTNPTIRQSNALEQALATQSIFQLGLLLVLPMVMEIGLEKGFRSALGDFIIMQLQLASVFFTFQLGTKAHYYGKTILHGGSKYRATG
RGFVVFHAKFAENYRLYSRSHFVKGLELTMLLVIYEVYGESYRSSSLYFFITFSMWFLVGSWLFAPFVFNPSGFDWQKTVDDWTDWKRWMGNRGGIGIPP
EKSWESWWDGEQEHLKHTNLRGRILDIILAFRFFVYQYGIVYHLDIAHRIKTLLAQSIPYVTLRAYIYVCVILLISGPQVYGLSWVVMITALLVFKMVSM
GRRKFGTDFQLTFRILKALLFMGFLSVLTVLFVVCNLTITDLFASILAFLPTGWAILLIGQACRGLFKAIGLWDSIKELGRGYEYIMGLLIFSPTAILSW
FPFVNEFQTRLLFNQAFSRGLQISMILQGRKDKDDAAAKKDKPPPPK
|
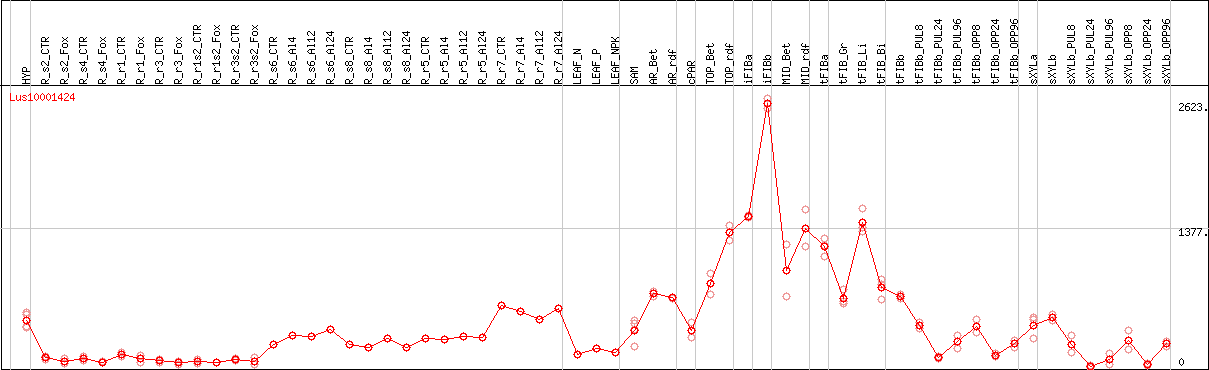
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10001424 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
