Lus10001455 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10001455 pacid=23161037 polypeptide=Lus10001455 locus=Lus10001455.g ID=Lus10001455.BGIv1.0 annot-version=v1.0
ATGCAGGAAGCGATCACAAGTTCTTCATCGAAACCGTTCGTCGGCCTTCGCTTTCTTCTCCGTGGCTTCGACTCTATCAGCCAGCAGCAGGTTCGATCAA
AGCTTCTGAGCGGCGGAGGAGTTGAGAGTCAGAACTATACTCATTTAATCGTCGACAACACTGCTTATGATGATCCGGAATGTGTTTCTGCTCGAAATAA
CGGGAAGACAGTGGTCACGGGGTTGTGGGTCGACCACAGTTTTGATATTGGATTGGTTGTTGATCGAACTTCTATAATGTATCAACCTCCAAAAGATATC
AATGGAATACCTGGTGCTGATAAACTGGTTGTGTGCTTGACGGGTTACCAGCGCCAAGATCGAGAAGACATCATGACTATGGTGAGTCTCATGGGTGCAC
GATTTTCCAAGCCCTTGATTGCCAACAAAGTCACTCACCTCATATGCTATAAGTTTGAAGGCGTGGCTTACAAGGTCTTTCATTACAGCAAGGTACTTTT
CAATGATCAATGCCCTTTGAAGATGTGCAATGTAGCTGAGTCTGCATTCCACGGTGAAAAGTACGAACTTGCAAGTAGGATTCAGAAGATAAAGCTTATC
AATCATCGTTGGTTGGAAGATTGTTTGACGAATTGGAAGCTCCTCCCAGAAGATGACTATCATAAGAGTGGTTATGACTTAGAGATGATGGAAACTGAAG
CAAAGGATTCTGAAGAGGATGTGACAGAAACTTCTCTTATGGTATCTACCGGTCATATTCCAAACCTGAGCCTTCGTAATATGGAAAATGGATCTCCAAA
GATTGGCAACTTGTCAAAACCTATTCCTGAAAGTTCTGCAATGGTGGCTGGAGATTTGTATGCCCACAATGCCAATGAGGTGTTAAGAACTCCTGTCAAT
AAGAAAAGGGGGGATATGGGCTGCAGTCCTGATAAGAATCATGCTTCTGATCTAATTGCTGCTCCATCAGAGAGAACTTCAGGTTCGGGGAACGGGGGTG
GCAAGATGAACTCTGTTTCTAAACCTTCACTTTCTTATACCGATTCGCTTGCTATGAGTCATGACAGGAAAACTTCACAAAGGTCTCCGTCACTATTCAT
TGGAGCATCAAAAGCTCCTGGATGCTATCCGAAAGTACAAACAGGGGAACCCATCAATTTCTCTTCTGCTAAAGTTGAAGTTGCAAACCAATGCCTTGGT
TCTGGTTTTGAGGATTTCGAAAGATTAGTTTCTGAAGAGCTATCAGATGGTAAAAAGCAAAGGATAGATGCCTTAGAGTCTAGCTCAAAAAGCCATCAGA
TGTTTCAGGATAGACAAACTTGGATCACAGGGAGTCCATCACTTAGCTATAGAGCTCTAGGATTGAAATCATCTTCTTTGGAAACTGCACCTAGGCTTAA
CAATAGCTCTCCTGATGGTAGTTATGCTCAGTCTCACTCTGGTAAGGGTATATTAAGTTCGAATGGTGCTTCAAATAGCCATAGCATGTTAGCATTGAGA
GAGCAATCACTAGCAGAAGGTGTCCCATTTGGTGAAAATGCAACAATGGAAAAAATAGGAAAGCAAAGTACTGATATGAAACACCAAGCGTCAGTGAGTG
GATTGAAAGTGTCAAGCTCACCTTCTAAGCAGGACGATGAAGTTCTTTGCTTAGACAAATCTGAAGTTGTGGATCTTAGAAATCAGCAGGTGAGGGCCTC
ACCGGTGGGTAAGATAGATATAACGACTGTGAAGCCTTCTGTGGTTACTGCAAATCAGAGGCAGGATAACAGCTTACAAACCAAACAAGAACGGAAGCAC
ACTGCAACCAAGAAGACATTGGGAACTAGAATGAAACTGAAAAGTACTGTGAAAGAAAAGGGCTCCATTTATTTGAAAGAAAGCACCGCCCAGGAAGATG
TGGCAACTGATCTACGCAGAAGAAATGACAGGACAGATGAAGAAAATCTGTTGATGGCCAAAGAGCTTGAAGACCCCCCTGCCGCTGTTGTTTCAATGGA
AGGTACGGGAAATAAACTTGTCATTGATCTAAGGACTGATGTCCAATATGATATTGACTCTATGAATGATGAAACGGAGCCTCCAGATGATAAAGAGCAG
CAAAGTGATGCTGTGTTGAAAGAAAGGATGGAAACAGGAGGTTTGGCAGGTAGCAGGGTAGAAAAAGAACCAGAAGAAGTCCGTCCTGTGGGGGGAAATT
CTAATGTTTGTGAAGTGGATGGTGTTATGGAAGATCCACAAAAGAAAGATAACTCAATCTCAGAAGGAGATTGTATCTCAGAAGACGATGGTGATAAAAG
GAAGAAAAGTATAAGCAAACGGCTAACTTCAATAAAGAGTGCGAGAACGGTTTCTAGTGGTGATGGTGAATCCAACTCTAGCAAAGCTGTGGAGGCTGAA
GAGACTCATCTTCACACAGTTGACCAAACTGCGAAAAACACATCTGAGCCTAGTGCCGGAAGCTGTCCAAGTAGGAAAAAGCTATCCAGAAGAAGATCAA
GGATCAGCATGGAAGCAGATAAGGAGAATACGGCTATCACCCAAGTACATAGAAGAGGGAAGATGACTGAAAAACATGTCGAATCATGCGAGGCAAGAAT
GGACGATAATCAGGACGTTCTAGAAACCAAACCCCTTTCTGGAAAACTTGAGGATATTTCTACTAACGTGGTGAAGGAACCAGCGTGCTTTATTATGAGC
GGACATAGACTGCAGAGGAAGGAGTTTCAGCAGGTTATAAGACGTTTGAAGGGAAAATCTTGCAGAGATTCTCACCAGTGGTCATACCAGGCAACTCATT
TCATTGCTCCGGACCCACTCCGCAGAACAGAAAAGTTACTTGCTGCTGCTGCCTCTGGAAGGTGGATTCTGAAGACTGACTATCTGTCCGCATGCAGTCA
GGCAGGGAAGTTTCTTCCTGAAGAGCCTTACGAATGGTACAAGAATGGCCTCAGTGAGGATGGAGCTATAAACTTGGAGGCCCCGAGGAAGTGGCGACTA
CTAAGAGAGCGAACAGGTCATGGGGCATTTCATGGTATGCAGATTGTCATCTACGGGGAGTGCATTTCACCACCGTTGGAAACACTTAAACGTGCAGTGA
AGGCTGGAGATGGAACCATACTAGCAACATCCCCTCCTTACTCGCGGTTCCTTGCCTCTGGGATCGACTATGCAATCGTAAGCCCCGGGATGCCACATGT
GGATGTGTGGGTGCAAGAGTTCCTAAGGCTGGAGATACCGTGCATAGTAGCTGATTACTTGGTGGAGTACGTATGCAAGTCGGGGTATTCTTTGGAGAAA
CATGTCCTCTTCGGCACCCAGAAATGGGCAGAGAAGTCGTTTAGCAACCTGATGAGGAAGGCGGAGGAGATAGTTGTGGAAAAGGCTGCAGCTTCAGCTT
CAGAAGAAGGTGGTGGTGAGAGCTCCCAGGAAGATGTGGCTTGCCAGGTATGCGGGAGTAGGGATAGAGGTGAAGTAATGCTGCTGTGTGGGGATGAAAG
TGGCAAAGTTGGGTGTGGGGGTGGAAGGCATATAGATTGCTGTGATCCTCCTCTTGACAGTATTCCTGAAGAAGATTGGTTCTGTCCAAACTGTTGCGAT
GACAGTTTAAACCCTTGTTTAAAGACAAAGAAACGGAAGAAGGGGACAAAATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10001455 pacid=23161037 polypeptide=Lus10001455 locus=Lus10001455.g ID=Lus10001455.BGIv1.0 annot-version=v1.0
MQEAITSSSSKPFVGLRFLLRGFDSISQQQVRSKLLSGGGVESQNYTHLIVDNTAYDDPECVSARNNGKTVVTGLWVDHSFDIGLVVDRTSIMYQPPKDI
NGIPGADKLVVCLTGYQRQDREDIMTMVSLMGARFSKPLIANKVTHLICYKFEGVAYKVFHYSKVLFNDQCPLKMCNVAESAFHGEKYELASRIQKIKLI
NHRWLEDCLTNWKLLPEDDYHKSGYDLEMMETEAKDSEEDVTETSLMVSTGHIPNLSLRNMENGSPKIGNLSKPIPESSAMVAGDLYAHNANEVLRTPVN
KKRGDMGCSPDKNHASDLIAAPSERTSGSGNGGGKMNSVSKPSLSYTDSLAMSHDRKTSQRSPSLFIGASKAPGCYPKVQTGEPINFSSAKVEVANQCLG
SGFEDFERLVSEELSDGKKQRIDALESSSKSHQMFQDRQTWITGSPSLSYRALGLKSSSLETAPRLNNSSPDGSYAQSHSGKGILSSNGASNSHSMLALR
EQSLAEGVPFGENATMEKIGKQSTDMKHQASVSGLKVSSSPSKQDDEVLCLDKSEVVDLRNQQVRASPVGKIDITTVKPSVVTANQRQDNSLQTKQERKH
TATKKTLGTRMKLKSTVKEKGSIYLKESTAQEDVATDLRRRNDRTDEENLLMAKELEDPPAAVVSMEGTGNKLVIDLRTDVQYDIDSMNDETEPPDDKEQ
QSDAVLKERMETGGLAGSRVEKEPEEVRPVGGNSNVCEVDGVMEDPQKKDNSISEGDCISEDDGDKRKKSISKRLTSIKSARTVSSGDGESNSSKAVEAE
ETHLHTVDQTAKNTSEPSAGSCPSRKKLSRRRSRISMEADKENTAITQVHRRGKMTEKHVESCEARMDDNQDVLETKPLSGKLEDISTNVVKEPACFIMS
GHRLQRKEFQQVIRRLKGKSCRDSHQWSYQATHFIAPDPLRRTEKLLAAAASGRWILKTDYLSACSQAGKFLPEEPYEWYKNGLSEDGAINLEAPRKWRL
LRERTGHGAFHGMQIVIYGECISPPLETLKRAVKAGDGTILATSPPYSRFLASGIDYAIVSPGMPHVDVWVQEFLRLEIPCIVADYLVEYVCKSGYSLEK
HVLFGTQKWAEKSFSNLMRKAEEIVVEKAAASASEEGGGESSQEDVACQVCGSRDRGEVMLLCGDESGKVGCGGGRHIDCCDPPLDSIPEEDWFCPNCCD
DSLNPCLKTKKRKKGTK
|
DESeq2's median of ratios [FLAX]
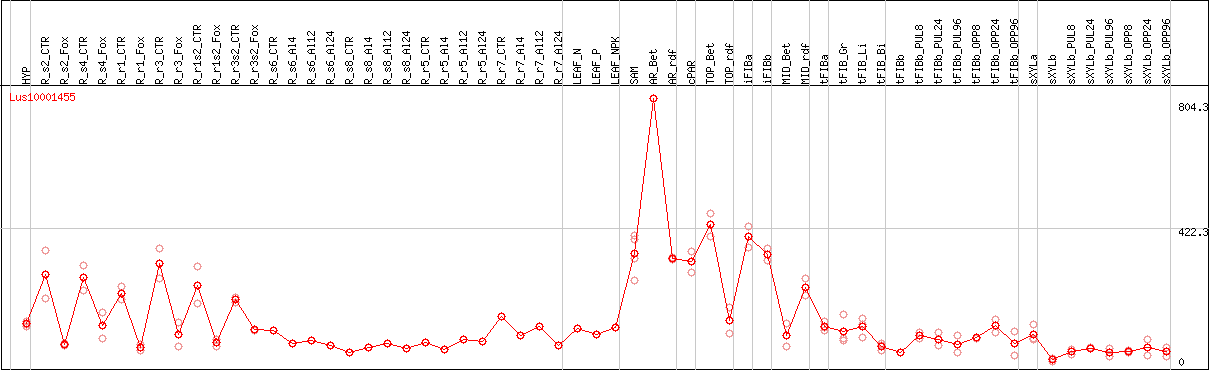
Coexpressed genes
Lus10001455 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
