External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G36690 136 / 4e-37
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G10500 122 / 4e-32
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G10490 121 / 9e-32
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT2G44800 119 / 9e-31
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT5G24530 117 / 2e-30
DMR6
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G22880 110 / 1e-27
TT18, TDS4, ANS, LDOX
TANNIN DEFICIENT SEED 4, ANTHOCYANIDIN SYNTHASE, leucoanthocyanidin dioxygenase (.1.2)
AT3G60290 110 / 2e-27
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT5G59530 107 / 2e-26
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT2G19590 105 / 4e-26
ATACO1, ACO1
ACC oxidase 1 (.1)
AT2G30830 106 / 5e-26
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10018949
430 / 3e-152
AT2G44800 148 / 1e-41
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10018950
427 / 3e-151
AT3G60290 147 / 3e-41
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10011263
183 / 4e-57
AT4G25420 50 / 2e-07
GA REQUIRING 5, ARABIDOPSIS THALIANA GIBBERELLIN 20-OXIDASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10030185
147 / 2e-41
AT4G10490 442 / 3e-156
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10001711
139 / 3e-39
AT2G36690 204 / 3e-64
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10014398
132 / 2e-35
AT2G36690 482 / 4e-171
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10023890
130 / 6e-35
AT2G36690 486 / 5e-173
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10035782
119 / 5e-31
AT5G24530 258 / 4e-84
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10028678
119 / 6e-31
AT1G77330 435 / 1e-154
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G185000
354 / 3e-122
AT2G36690 147 / 5e-41
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.017G048700
200 / 9e-62
AT2G36690 255 / 3e-82
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.017G048900
200 / 2e-61
AT2G36690 250 / 5e-80
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G451700
148 / 1e-41
AT4G10500 451 / 6e-160
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.008G029700
145 / 2e-40
AT2G36690 478 / 1e-169
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.014G106700
144 / 4e-40
AT4G10490 468 / 2e-166
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.011G150100
143 / 5e-40
AT4G10490 501 / 2e-179
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G451600
140 / 7e-39
AT4G10500 406 / 5e-142
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.010G231500
140 / 2e-38
AT2G36690 474 / 3e-168
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G451900
138 / 4e-38
AT4G10490 498 / 3e-178
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0029
Cupin
PF03171
2OG-FeII_Oxy
2OG-Fe(II) oxygenase superfamily
Representative CDS sequence
>Lus10001460 pacid=23161034 polypeptide=Lus10001460 locus=Lus10001460.g ID=Lus10001460.BGIv1.0 annot-version=v1.0
ATGGTTCAAATAGAAGCAGTTGTGATGGATGGACAAGTTCCAACCATAGACTACTTCTCCCTTTTCTCTACTGACCATGCCCAGCGATCCGCAGCCCTTG
CATGCCTCTCAACTACGTGCAGAGATTACGGGTTCTTCAACTTGGTGAACCATGGGATTCCAGACGGAGTGATAGAAAGTGCACTGAGCAGCATTGGAGA
ATTTTACGACAGCGTAGAGCTGCAAGAAAAGAACAAGTATAGTAGAGGAGATCCGGCAGCTAAGGTTCTTTGGGATGGTAGATACCACGGTGGTGCTATC
GCTAGGGAGCATCTCAAGCTACTTGCACGTCCCGAACTCCATTGCCCTCCTAATCATCCTTCTGTTAGGGAGAGCATGGATGATTTCCAGACCAGATTCC
ATCAAGTGAAGCTAGGGTTGGCAAGGGCTATGTCTGTAATATTGGGACAGAGAGAATCCTACATTGAGACAGCTTTGAATCTGGAATCAGGGTTTGACGT
TGTAGCAAACAATAGTTACCCACCAAATTCAAACTCAAACGGCACAACTAGACTCCCTGAACATACCGATCCAGGCTTCATCATCTCCCTTATTCAAGAC
ATGGATGGTGGCCTTCAAATCCTAACTCACCAAGGAAAATGGATCAACATCCGCATCCCTCCCAATGGTATCTTAATTCAACTTGGGGATCATCTCGAAA
TTCTGACTAATGGAAAGTACAAGAGTCACATTCATCGAGCGATGATACATAACTATAAAGTAAGAAGGATCTCTATAGCCACACTCCATGGACCATCTAT
TGGCACATTTGTAGCTCCGGCGCCGGAATTCGTTGATGATTCCCATCCTTCAGCTTACCGTGGAATGACCTACTCGGAATTCCTGAAAGGCAACAATTGT
TACGAGATTGAAGTCCAATCATGCCTCGATCAAATTAGGCTTCTACCCAAAATAGAATAG
AA sequence
>Lus10001460 pacid=23161034 polypeptide=Lus10001460 locus=Lus10001460.g ID=Lus10001460.BGIv1.0 annot-version=v1.0
MVQIEAVVMDGQVPTIDYFSLFSTDHAQRSAALACLSTTCRDYGFFNLVNHGIPDGVIESALSSIGEFYDSVELQEKNKYSRGDPAAKVLWDGRYHGGAI
AREHLKLLARPELHCPPNHPSVRESMDDFQTRFHQVKLGLARAMSVILGQRESYIETALNLESGFDVVANNSYPPNSNSNGTTRLPEHTDPGFIISLIQD
MDGGLQILTHQGKWINIRIPPNGILIQLGDHLEILTNGKYKSHIHRAMIHNYKVRRISIATLHGPSIGTFVAPAPEFVDDSHPSAYRGMTYSEFLKGNNC
YEIEVQSCLDQIRLLPKIE
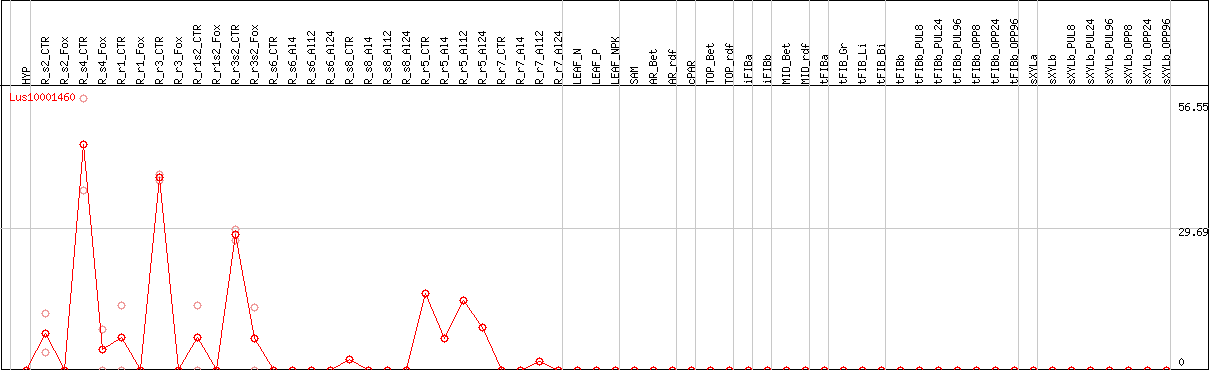
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10001460 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.