External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G47490 382 / 6e-134
ATNDT1
NAD+ transporter 1, ARABIDOPSIS THALIANA NAD+ TRANSPORTER 1, NAD+ transporter 1 (.1)
AT1G25380 316 / 8e-107
ATNDT2
NAD+ transporter 2, ARABIDOPSIS THALIANA NAD+ TRANSPORTER 2, NAD+ transporter 2 (.1)
AT5G66380 155 / 7e-45
ATFOLT1
folate transporter 1 (.1)
AT5G46800 123 / 8e-33
BOU
A BOUT DE SOUFFLE, Mitochondrial substrate carrier family protein (.1)
AT4G26180 113 / 7e-29
Mitochondrial substrate carrier family protein (.1)
AT5G48970 110 / 7e-28
Mitochondrial substrate carrier family protein (.1)
AT5G56450 108 / 3e-27
PM-ANT
Mitochondrial substrate carrier family protein (.1)
AT4G01100 108 / 9e-27
ADNT1
adenine nucleotide transporter 1 (.1.2)
AT3G51870 107 / 2e-26
Mitochondrial substrate carrier family protein (.1)
AT4G32400 106 / 7e-26
EMB42, EMB104, ATBT1, SHS1
SODIUM HYPERSENSITIVE 1, EMBRYO DEFECTIVE 42, EMBRYO DEFECTIVE 104, ARABIDOPSIS THALIANA BRITTLE 1, Mitochondrial substrate carrier family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10034104
337 / 7e-116
AT1G25380 414 / 3e-145
NAD+ transporter 2, ARABIDOPSIS THALIANA NAD+ TRANSPORTER 2, NAD+ transporter 2 (.1)
Lus10003041
242 / 5e-80
AT1G25380 299 / 6e-102
NAD+ transporter 2, ARABIDOPSIS THALIANA NAD+ TRANSPORTER 2, NAD+ transporter 2 (.1)
Lus10000906
117 / 1e-30
AT5G46800 482 / 1e-173
A BOUT DE SOUFFLE, Mitochondrial substrate carrier family protein (.1)
Lus10028339
116 / 1e-30
AT5G66380 384 / 2e-135
folate transporter 1 (.1)
Lus10032522
119 / 3e-30
AT5G51050 563 / 0.0
ATP/phosphate carrier 2, Mitochondrial substrate carrier family protein (.1)
Lus10043022
119 / 6e-30
AT5G51050 758 / 0.0
ATP/phosphate carrier 2, Mitochondrial substrate carrier family protein (.1)
Lus10008572
115 / 2e-29
AT4G26180 494 / 4e-177
Mitochondrial substrate carrier family protein (.1)
Lus10032690
115 / 2e-29
AT4G26180 493 / 4e-177
Mitochondrial substrate carrier family protein (.1)
Lus10002902
115 / 5e-29
AT4G32400 522 / 0.0
SODIUM HYPERSENSITIVE 1, EMBRYO DEFECTIVE 42, EMBRYO DEFECTIVE 104, ARABIDOPSIS THALIANA BRITTLE 1, Mitochondrial substrate carrier family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G200900
431 / 5e-153
AT2G47490 471 / 1e-168
NAD+ transporter 1, ARABIDOPSIS THALIANA NAD+ TRANSPORTER 1, NAD+ transporter 1 (.1)
Potri.014G125600
410 / 1e-144
AT2G47490 448 / 1e-159
NAD+ transporter 1, ARABIDOPSIS THALIANA NAD+ TRANSPORTER 1, NAD+ transporter 1 (.1)
Potri.010G121500
355 / 4e-122
AT1G25380 395 / 2e-137
NAD+ transporter 2, ARABIDOPSIS THALIANA NAD+ TRANSPORTER 2, NAD+ transporter 2 (.1)
Potri.008G123700
331 / 7e-113
AT1G25380 417 / 8e-146
NAD+ transporter 2, ARABIDOPSIS THALIANA NAD+ TRANSPORTER 2, NAD+ transporter 2 (.1)
Potri.007G019000
146 / 2e-41
AT5G66380 463 / 8e-166
folate transporter 1 (.1)
Potri.008G210000
121 / 9e-32
AT5G48970 517 / 0.0
Mitochondrial substrate carrier family protein (.1)
Potri.010G024100
114 / 3e-29
AT5G48970 528 / 0.0
Mitochondrial substrate carrier family protein (.1)
Potri.006G153600
112 / 2e-28
AT4G26180 475 / 4e-170
Mitochondrial substrate carrier family protein (.1)
Potri.012G110700
112 / 1e-27
AT5G51050 714 / 0.0
ATP/phosphate carrier 2, Mitochondrial substrate carrier family protein (.1)
Potri.006G252100
110 / 1e-27
AT4G32400 493 / 4e-175
SODIUM HYPERSENSITIVE 1, EMBRYO DEFECTIVE 42, EMBRYO DEFECTIVE 104, ARABIDOPSIS THALIANA BRITTLE 1, Mitochondrial substrate carrier family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00153
Mito_carr
Mitochondrial carrier protein
Representative CDS sequence
>Lus10001463 pacid=23161044 polypeptide=Lus10001463 locus=Lus10001463.g ID=Lus10001463.BGIv1.0 annot-version=v1.0
ATGGGGAGTAGCAATGAATCTCATGCCCTTACTTTGAAAGGTCTTATCTCCAATGCAGGTGCTGGTGCAGCTGCAGGAATCATTGCTGCGACGTTTGTTT
GTCCGTTAGATGTCATCAAAACACGATTTCAAGTCCACGGTCTCCCACAGCTCAATCATGATGCTAAGTTTAAACGTAATCTGATAGCAGGGAGTTTGGA
ACATATATTCAGAAAGGAAGGTGTACCTGGAATGTATCGTGGGCTTGCAGCTACAGTTCTGGCCTTACTTCCTAATTGGGCCATCTATTTTACAGTGTAC
GAACAACTCAAAGGCCTTCTCAAGTCCCCTGATGAGAATCATCATCTATCACTGGGTGCAAATATGATAGCTGCATCAGGAGCTGGGGTAACAACAACTC
TTTTCACAAATCCTCTCTGGGTTGTTAAGACGAGACTTCAAGCTCAAGGATCAAAGGGAGGTGTTATTCCGTATAAGAACACTTTATCTGCTTTGAAGAG
AATAGCCGGCGAGGAAGGTCTTCGCGGGTTATATAGTGGTCTTGTACCTGCTCTAGCGGGGATAAGTCATGTCGCAATACAATTTCCTACGTACGAGAAG
ATCAAAGTCTATTTAACTGAAAAAGACAATACGACAGTTGACAAGTTGGGGGCACGTGACGTGGCTATTGCTTCTTCAATATCCAAAATATTTGCCTCCA
CCTTGACATATCCACATGAGGTTGTGCGTTCGAGGCTTATGGAACAAGGCCTTCATTCTACACAACGTTATACAGGCGTGATCGACTGTTTACGGAAAAT
GTACGAAAAAGAAGGAATTGTTGGATTTTACAGAGGGTGTGGGATTAATCTTTTGAGGACAACACCCGCAGCTGTTATCACCTTCACAAGCTTCGAAATG
ATACATAGATTTCTTCTTACTGCTTTACCTCCTGGCCCAACAACAACATGA
AA sequence
>Lus10001463 pacid=23161044 polypeptide=Lus10001463 locus=Lus10001463.g ID=Lus10001463.BGIv1.0 annot-version=v1.0
MGSSNESHALTLKGLISNAGAGAAAGIIAATFVCPLDVIKTRFQVHGLPQLNHDAKFKRNLIAGSLEHIFRKEGVPGMYRGLAATVLALLPNWAIYFTVY
EQLKGLLKSPDENHHLSLGANMIAASGAGVTTTLFTNPLWVVKTRLQAQGSKGGVIPYKNTLSALKRIAGEEGLRGLYSGLVPALAGISHVAIQFPTYEK
IKVYLTEKDNTTVDKLGARDVAIASSISKIFASTLTYPHEVVRSRLMEQGLHSTQRYTGVIDCLRKMYEKEGIVGFYRGCGINLLRTTPAAVITFTSFEM
IHRFLLTALPPGPTTT
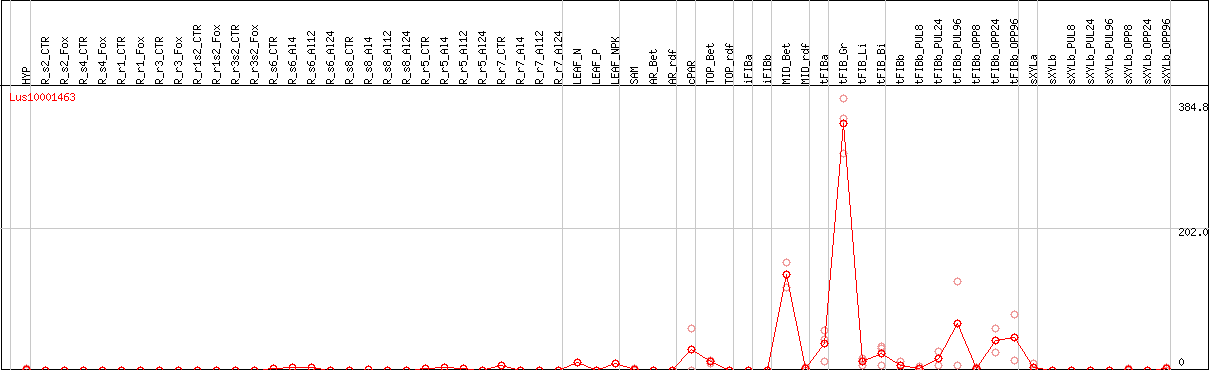
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10001463 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.