External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G46790 314 / 2e-104
CRR2
CHLORORESPIRATORY REDUCTION 2, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G20230 222 / 3e-68
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT4G02750 220 / 2e-67
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G57430 213 / 2e-64
OTP84
ORGANELLE TRANSCRIPT PROCESSING 84, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G14330 210 / 5e-64
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G18750 211 / 1e-63
DOT4
DEFECTIVELY ORGANIZED TRIBUTARIES 4, Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G13770 207 / 1e-63
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G49142 206 / 1e-62
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G12770 201 / 1e-60
MEF22
mitochondrial editing factor 22 (.1)
AT4G14850 200 / 1e-60
MEF11, LOI1
lovastatin insensitive 1, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10004987
357 / 2e-123
AT3G46790 715 / 0.0
CHLORORESPIRATORY REDUCTION 2, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10040504
224 / 6e-70
AT3G13770 904 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10016387
206 / 2e-64
AT4G33990 717 / 0.0
embryo defective 2758, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10040577
201 / 3e-63
AT1G25360 577 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10028837
202 / 4e-63
AT1G11290 650 / 0.0
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10001617
206 / 7e-63
AT2G03880 840 / 0.0
required for efficiency of mitochondrial editing 1, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10017446
206 / 5e-62
AT1G11290 1152 / 0.0
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10029436
203 / 1e-61
AT3G08820 828 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10019735
203 / 4e-61
AT4G33990 1015 / 0.0
embryo defective 2758, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10001557 pacid=23154583 polypeptide=Lus10001557 locus=Lus10001557.g ID=Lus10001557.BGIv1.0 annot-version=v1.0
ATGCGAATCGAACCAGGACCGAAGGTCTGGGGAGCGCTACTCGGTTCGTGCAGGATCCATTGCAACGTCGAGCCGGCAGAGTGGGCAAGCACGAAGCTTT
TTGAGCTCGAGCCTACAAATGCAGGGAACTATGTTCTTCTAGCCAATATTTACGCAGAAGCAGGAATGTGGGACGAGGTGAAGCGAGTCAAAAAGATGAT
GGAATCGAGGGGGATCCGGAAACTCCATGGACGGAGTTGGATCGAAGTGAAGAGGGAGCTTCACTCGTTCGTATCAGTAGACGAGTTCAATCATCTGCTG
TTCGAATTATCGTCTCAGATGAAGGAGAATGGGTATGTGCCGTGGACTAATGTGGTGCTGTACGATCTGGAGGAAGAGGAGAAGGAGAGGATTGTGCTTG
GACATAGTGAGAAGTTGGCTTTGGCGTTTGGACTTGTGAATAGCAGTAAAGGGGAGGTGATTCGGATTACAAAGAGTTTGAGGCTGTGGGAAGACTGTCA
TTCCTTTACGAAGTTTGTTTCGAGGTTTGTGGGGAGGGGGATTCTGGTTAGGGATGTGAATCGGTTTCACCGGTTCCGAGATGGGGTCTGTTCTTGTGGG
GATTTCTGGTGA
AA sequence
>Lus10001557 pacid=23154583 polypeptide=Lus10001557 locus=Lus10001557.g ID=Lus10001557.BGIv1.0 annot-version=v1.0
MRIEPGPKVWGALLGSCRIHCNVEPAEWASTKLFELEPTNAGNYVLLANIYAEAGMWDEVKRVKKMMESRGIRKLHGRSWIEVKRELHSFVSVDEFNHLL
FELSSQMKENGYVPWTNVVLYDLEEEEKERIVLGHSEKLALAFGLVNSSKGEVIRITKSLRLWEDCHSFTKFVSRFVGRGILVRDVNRFHRFRDGVCSCG
DFW
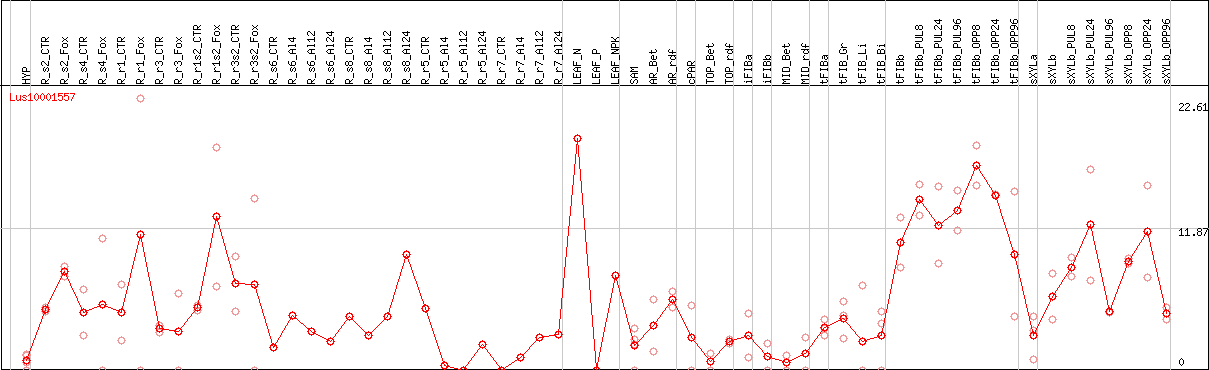
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10001557 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.