External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G61430 340 / 2e-116
NAC
ANAC100, ATNAC5
NAC domain containing protein 100 (.1)
AT5G07680 331 / 4e-113
NAC
ANAC079, ATNAC4, ANAC080
Arabidopsis NAC domain containing protein 79, NAC domain containing protein 80 (.1.2)
AT5G39610 291 / 4e-98
NAC
ORE1, AtNAC2, ANAC092, ATNAC6
ORESARA 1, NAC domain containing protein 2, Arabidopsis NAC domain containing protein 92, NAC domain containing protein 6 (.1)
AT3G29035 283 / 2e-94
NAC
ORS1, AtNAC3, ANAC059
ORE1 SISTER1, Arabidopsis NAC domain containing protein 59, NAC domain containing protein 3 (.1)
AT3G04060 273 / 4e-90
NAC
ANAC046
NAC domain containing protein 46 (.1)
AT5G18270 269 / 1e-88
NAC
ANAC087
Arabidopsis NAC domain containing protein 87 (.1.2)
AT5G53950 263 / 8e-86
NAC
ATCUC2, CUC2, ANAC098
CUP-SHAPED COTYLEDON 2, Arabidopsis NAC domain containing protein 98, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT3G15170 257 / 3e-84
NAC
ATNAC1, CUC1, ANAC054
CUP-SHAPED COTYLEDON1, Arabidopsis NAC domain containing protein 54, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT3G18400 253 / 2e-82
NAC
ANAC058
NAC domain containing protein 58 (.1)
AT2G24430 239 / 4e-77
NAC
ANAC039, ANAC038
Arabidopsis NAC domain containing protein 39, NAC domain containing protein 38 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021659
567 / 0
AT5G61430 361 / 8e-125
NAC domain containing protein 100 (.1)
Lus10005537
265 / 2e-87
AT5G53950 317 / 3e-107
CUP-SHAPED COTYLEDON 2, Arabidopsis NAC domain containing protein 98, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10026879
265 / 1e-86
AT5G18270 325 / 3e-110
Arabidopsis NAC domain containing protein 87 (.1.2)
Lus10003435
255 / 6e-83
AT5G18270 315 / 2e-106
Arabidopsis NAC domain containing protein 87 (.1.2)
Lus10009029
248 / 4e-80
AT3G18400 311 / 3e-105
NAC domain containing protein 58 (.1)
Lus10020165
248 / 1e-79
AT2G24430 306 / 2e-102
Arabidopsis NAC domain containing protein 39, NAC domain containing protein 38 (.1.2)
Lus10009669
246 / 2e-79
AT3G18400 308 / 3e-104
NAC domain containing protein 58 (.1)
Lus10041924
239 / 8e-79
AT5G53950 281 / 9e-95
CUP-SHAPED COTYLEDON 2, Arabidopsis NAC domain containing protein 98, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10026966
243 / 1e-77
AT2G24430 311 / 6e-105
Arabidopsis NAC domain containing protein 39, NAC domain containing protein 38 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G001400
355 / 3e-122
AT5G61430 448 / 1e-158
NAC domain containing protein 100 (.1)
Potri.015G020000
354 / 1e-121
AT5G61430 446 / 1e-157
NAC domain containing protein 100 (.1)
Potri.017G086200
322 / 3e-109
AT5G61430 386 / 2e-134
NAC domain containing protein 100 (.1)
Potri.019G031600
288 / 4e-95
AT5G18270 328 / 6e-111
Arabidopsis NAC domain containing protein 87 (.1.2)
Potri.013G054200
280 / 3e-92
AT5G18270 339 / 4e-115
Arabidopsis NAC domain containing protein 87 (.1.2)
Potri.001G396300
260 / 2e-85
AT5G53950 299 / 5e-100
CUP-SHAPED COTYLEDON 2, Arabidopsis NAC domain containing protein 98, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.011G115400
260 / 9e-85
AT5G53950 312 / 1e-104
CUP-SHAPED COTYLEDON 2, Arabidopsis NAC domain containing protein 98, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.018G003800
251 / 3e-81
AT2G24430 350 / 1e-120
Arabidopsis NAC domain containing protein 39, NAC domain containing protein 38 (.1.2)
Potri.012G056300
248 / 6e-81
AT3G18400 317 / 6e-108
NAC domain containing protein 58 (.1)
Potri.015G046800
249 / 7e-81
AT3G18400 326 / 1e-111
NAC domain containing protein 58 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Lus10001648 pacid=23175237 polypeptide=Lus10001648 locus=Lus10001648.g ID=Lus10001648.BGIv1.0 annot-version=v1.0
ATGGAAAACAACGTTGCGTGTGGAGTTGAGATGGTGATGGAGGAAGAAGATGACCAGATGGATTTGCCACCTGGGTTCCGTTTCCACCCTACTGATGAAG
AGCTCATCTCCCATTATCTCTACAACAAGGTCATCGATGTCAGATTCTCCTGCCGAGCCATCGGCGACGTCGATTTGAACAAGTCTGAGCCCTGGGAACT
TCCCTGGAAAGCGAAGATGGGGGAGAAAGAGTGGTATTTCTTCTGCGTTCGAGATAGGAAGTACCCAACTGGGCTGAGGACGAACAGAGCAACGGAAGCT
GGGTACTGGAAAGCTACAGGGAAAGACAAGGAGATTTACAGAGGGAAATCTCTGGTTGGGATGAAGAAAACCTTGGTCTTTTACAAAGGAAGAGCTCCCA
AGGGGGAGAAATCCAATTGGGTTATGCACGAGTACAGATTAGACGGCAAATTCTCCATCCACAACCTCCCCAAAACCGCCAAGAATGAATGGGTGATATG
CAGGGTATTCCAGAAGACTTGCGGCGGGAAGAAGATTCACATCTCCGGAATCACACGATTCGGAGAGTTGCAGGGTCATCATCATAATTCTTCGACCATG
TTGCCGCCGTTGATGGAATCACCTTCAGCTAATTACGATCAGAAATTTACAATTAAGCCAGCCGTAGCTGCAGAATCTTCTTCAGCTTACGTGCCCTGCT
TCTCCAATCTCATTACCAACATGAACACAACCAACAATTCATCAACATCCTCCTCTCAATTTCTATCTAATCCAATAATTCCTTCATCATCCTCTTCTTC
AATCCCTCTCTTCCCATCTTCTCATTACAATTCCCAGTTGATCCCTTCCGCTAATTTCGGCCAATACAACGACAACTCCATCCTACTCCGAGCTCTGATG
GCGGATAAACAAGGAGGAGGAGGAGGGTCGACGTTGAATCTGATGAACAGTACTGAGATTTCTCCGTCGGTTACTGGACCAGTGGATCTCGACTCTATTT
GGGACTATTAA
AA sequence
>Lus10001648 pacid=23175237 polypeptide=Lus10001648 locus=Lus10001648.g ID=Lus10001648.BGIv1.0 annot-version=v1.0
MENNVACGVEMVMEEEDDQMDLPPGFRFHPTDEELISHYLYNKVIDVRFSCRAIGDVDLNKSEPWELPWKAKMGEKEWYFFCVRDRKYPTGLRTNRATEA
GYWKATGKDKEIYRGKSLVGMKKTLVFYKGRAPKGEKSNWVMHEYRLDGKFSIHNLPKTAKNEWVICRVFQKTCGGKKIHISGITRFGELQGHHHNSSTM
LPPLMESPSANYDQKFTIKPAVAAESSSAYVPCFSNLITNMNTTNNSSTSSSQFLSNPIIPSSSSSSIPLFPSSHYNSQLIPSANFGQYNDNSILLRALM
ADKQGGGGGSTLNLMNSTEISPSVTGPVDLDSIWDY
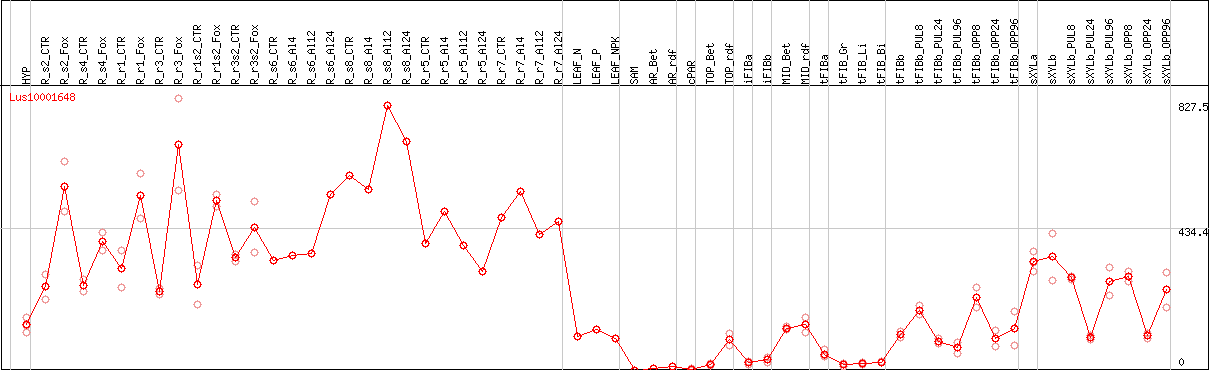
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10001648 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.