External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G28050 237 / 5e-81
Cytidine/deoxycytidylate deaminase family protein (.1.2)
AT3G05300 142 / 1e-44
Cytidine/deoxycytidylate deaminase family protein (.1)
AT1G68720 74 / 1e-15
ATTADA, TADA
ARABIDOPSIS THALIANA TRNA ADENOSINE DEAMINASE A, tRNA arginine adenosine deaminase (.1)
AT1G48175 40 / 0.0002
TAD1, EMB2191
tRNA adenosine deaminase 1, embryo defective 2191, Cytidine/deoxycytidylate deaminase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015942
279 / 9e-98
AT5G28050 288 / 3e-101
Cytidine/deoxycytidylate deaminase family protein (.1.2)
Lus10019153
73 / 2e-15
AT1G68720 301 / 2e-84
ARABIDOPSIS THALIANA TRNA ADENOSINE DEAMINASE A, tRNA arginine adenosine deaminase (.1)
Lus10034402
73 / 2e-15
AT1G68720 436 / 2e-131
ARABIDOPSIS THALIANA TRNA ADENOSINE DEAMINASE A, tRNA arginine adenosine deaminase (.1)
Lus10028482
44 / 1e-05
AT1G48175 269 / 5e-93
tRNA adenosine deaminase 1, embryo defective 2191, Cytidine/deoxycytidylate deaminase family protein (.1)
Lus10009163
41 / 0.0001
AT1G48175 265 / 1e-91
tRNA adenosine deaminase 1, embryo defective 2191, Cytidine/deoxycytidylate deaminase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G049300
250 / 5e-86
AT5G28050 281 / 4e-98
Cytidine/deoxycytidylate deaminase family protein (.1.2)
Potri.013G036000
245 / 7e-84
AT5G28050 275 / 1e-95
Cytidine/deoxycytidylate deaminase family protein (.1.2)
Potri.010G003500
241 / 1e-82
AT5G28050 267 / 2e-92
Cytidine/deoxycytidylate deaminase family protein (.1.2)
Potri.008G111100
79 / 2e-17
AT1G68720 522 / 3e-162
ARABIDOPSIS THALIANA TRNA ADENOSINE DEAMINASE A, tRNA arginine adenosine deaminase (.1)
Potri.010G132600
72 / 4e-15
AT1G68720 555 / 2e-174
ARABIDOPSIS THALIANA TRNA ADENOSINE DEAMINASE A, tRNA arginine adenosine deaminase (.1)
Potri.006G075300
45 / 4e-06
AT1G48175 227 / 1e-76
tRNA adenosine deaminase 1, embryo defective 2191, Cytidine/deoxycytidylate deaminase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0109
CDA
PF00383
dCMP_cyt_deam_1
Cytidine and deoxycytidylate deaminase zinc-binding region
Representative CDS sequence
>Lus10001791 pacid=23169676 polypeptide=Lus10001791 locus=Lus10001791.g ID=Lus10001791.BGIv1.0 annot-version=v1.0
ATGGCAGATGGCAACGCTGTGAAAGACAGAGACCACAAGTTCTTGACTAAAGCTGTTGAAGAAGCGTACAAAGGTGTTGAATGTGGACATGGAGGTCCTT
TTGGCGCTGTCGTTGTTTGCAATGACGAAGTTGTGGTTAGCTGTCACAACATGGTCATGGAACACACTGACCCAACTGCTCACGCGGAGGTCACAGCTGT
TAGAGAGGCATGTAAAAAGCTCAACAGAATTGAGCTGTCGGATTGCGAGATTTATGCATCCTGTGAACCATGCCCAATGTGCTTTGGGGCCATCCATCTG
TCTAGAATCAAGAGGCTGGTCTATGGAGCCAAAGCTGAAGCAGCAATTGCGATTGGTTTCGATGATTTCATTGCTGATGCGTTGAGAGGAACAGGGTTCT
ACCAGAAGGCTAATTTGGAGATCAAGCAGGCTGATGGCAGCGGTGCTCTTATTGCAGAACAAGTGTTCGAGAACACCAAGGCCAGGTTTACCATGTACTG
A
AA sequence
>Lus10001791 pacid=23169676 polypeptide=Lus10001791 locus=Lus10001791.g ID=Lus10001791.BGIv1.0 annot-version=v1.0
MADGNAVKDRDHKFLTKAVEEAYKGVECGHGGPFGAVVVCNDEVVVSCHNMVMEHTDPTAHAEVTAVREACKKLNRIELSDCEIYASCEPCPMCFGAIHL
SRIKRLVYGAKAEAAIAIGFDDFIADALRGTGFYQKANLEIKQADGSGALIAEQVFENTKARFTMY
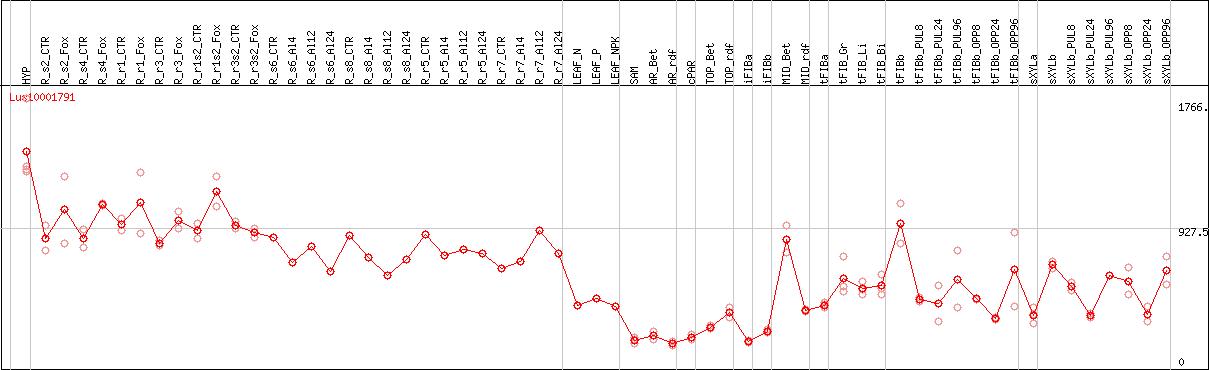
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10001791 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.