External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G13180 270 / 2e-91
NAC
VNDIP2, ANAC083, VNI2
VND-interacting 2, NAC domain containing protein 83 (.1)
AT2G33480 264 / 5e-89
NAC
ANAC041
NAC domain containing protein 41 (.1.2)
AT3G04070 186 / 2e-57
NAC
ANAC047
NAC domain containing protein 47 (.1.2)
AT1G69490 183 / 2e-57
NAC
NAP, ANAC029, ATNAP
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
AT1G61110 182 / 3e-56
NAC
ANAC025
NAC domain containing protein 25 (.1)
AT4G27410 180 / 1e-55
NAC
RD26, ANAC072
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.2), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.3)
AT1G77450 176 / 1e-54
NAC
ANAC032
NAC domain containing protein 32 (.1)
AT3G15510 179 / 2e-54
NAC
ATNAC2, ANAC056, NARS1
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
AT1G65910 181 / 4e-53
NAC
ANAC028
NAC domain containing protein 28 (.1)
AT1G52890 172 / 2e-52
NAC
ANAC019
NAC domain containing protein 19 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002581
469 / 3e-170
AT5G13180 273 / 6e-93
VND-interacting 2, NAC domain containing protein 83 (.1)
Lus10036773
191 / 7e-60
AT1G69490 318 / 1e-109
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Lus10037156
191 / 7e-60
AT1G69490 318 / 2e-109
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Lus10030446
189 / 3e-59
AT1G69490 302 / 2e-103
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Lus10026617
186 / 5e-58
AT1G69490 298 / 1e-101
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Lus10011215
187 / 9e-58
AT1G61110 305 / 1e-102
NAC domain containing protein 25 (.1)
Lus10026496
186 / 3e-57
AT1G61110 244 / 6e-79
NAC domain containing protein 25 (.1)
Lus10023179
186 / 3e-57
AT1G61110 248 / 2e-80
NAC domain containing protein 25 (.1)
Lus10015076
186 / 4e-57
AT1G61110 247 / 5e-80
NAC domain containing protein 25 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Lus10001809 pacid=23140343 polypeptide=Lus10001809 locus=Lus10001809.g ID=Lus10001809.BGIv1.0 annot-version=v1.0
ATGGAGAAGATGAGCTTTGTCAAGAATGGAGTGCTGAGACTGCCACCTGGCTTCAGATTCCACCCAACTGACGAGGAGCTCGTGGTTCAATACCTCAAAC
GGAAAGTCTATTCCTTTCCTTTGCCCGCTTCCATCATTCCCGAAGTCGATGTCTGCAAGTCTGATCCTTGGGATTTACCAGGTGATTTGGAGCAAGAGAG
GTACTTTTTCAGCACCAAGGAGGCCAAATATCCCAATGGGAACCGTTCCAATAGAGCTACTGGGTCAGGCTACTGGAAGGCAACAGGGATAGACAAGCAA
ATTGTGGCTGCTTCTAAAACTGTTGTTGGGATGAAGAAAACTCTGGTTTTTTACAGAGGGAAGCCCCCACATGGCGCTAGAACTGATTGGATCATGCATG
AGTACCGCTTCCTTCCTTCTCCTTCTGATGCTCCCCACCAGCGGATGGAGAATTGGGTTCTGTGCCGAATATTTTTGAAGAAGAGAGGTTCCAAGAATGA
GGAGGATGCAGGATGCTGCATTAAGCAGCTTGGTTCTTCTAACCAGACAGTGACCAAGCATGTCAAACCAGCTTTGACCAAGAAGCCTGTGTTCTACGAC
TTCATGACAAAACCGTGTGGTTCATCCTCAGGAACAAGTGGGATCACAGATGATGTTTCCAATAATGAAACAGATGAGCATGAGGAAAATAGCAGCAGCG
GCAACACTTCTTCTTTCCCTCATTTCAGGAGAAAACCATAG
AA sequence
>Lus10001809 pacid=23140343 polypeptide=Lus10001809 locus=Lus10001809.g ID=Lus10001809.BGIv1.0 annot-version=v1.0
MEKMSFVKNGVLRLPPGFRFHPTDEELVVQYLKRKVYSFPLPASIIPEVDVCKSDPWDLPGDLEQERYFFSTKEAKYPNGNRSNRATGSGYWKATGIDKQ
IVAASKTVVGMKKTLVFYRGKPPHGARTDWIMHEYRFLPSPSDAPHQRMENWVLCRIFLKKRGSKNEEDAGCCIKQLGSSNQTVTKHVKPALTKKPVFYD
FMTKPCGSSSGTSGITDDVSNNETDEHEENSSSGNTSSFPHFRRKP
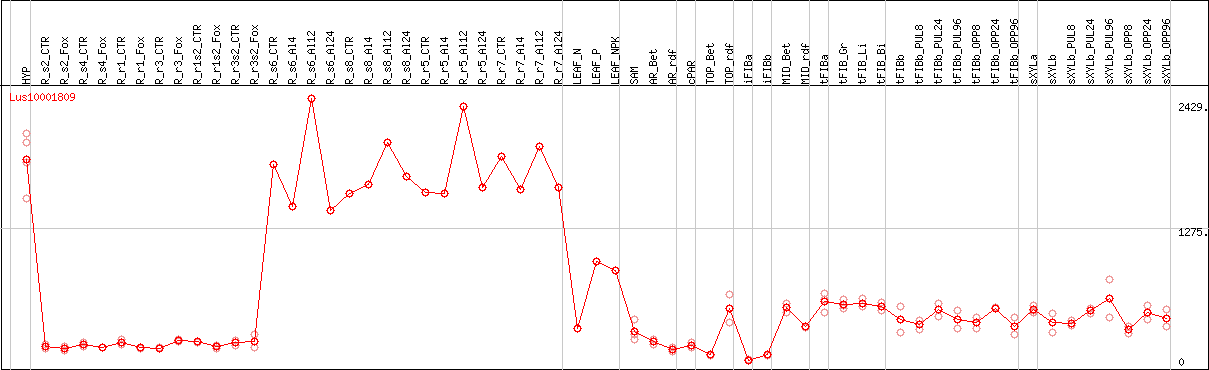
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10001809 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.