External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G23900 347 / 2e-122
Nucleoside diphosphate kinase family protein (.1)
AT4G11010 338 / 5e-119
NDPK3
nucleoside diphosphate kinase 3 (.1)
AT4G23895 347 / 7e-119
Pleckstrin homology (PH) domain-containing protein (.1), Pleckstrin homology (PH) domain-containing protein (.2), Pleckstrin homology (PH) domain-containing protein (.3)
AT4G09320 186 / 8e-60
NDPK1
Nucleoside diphosphate kinase family protein (.1)
AT5G63310 183 / 5e-58
NDPK1A, NDPKIAIA, NDPK2, ATNDPK2
NDP KINASE 1A, NUCLEOSIDE DIPHOSPHATE KINASE IA, ARABIDOPSIS NUCLEOSIDE DIPHOSPHATE KINASE 2, nucleoside diphosphate kinase 2 (.1)
AT1G17410 89 / 6e-22
Nucleoside diphosphate kinase family protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10003878
399 / 1e-142
AT4G23900 362 / 6e-128
Nucleoside diphosphate kinase family protein (.1)
Lus10032392
325 / 4e-114
AT4G11010 328 / 1e-115
nucleoside diphosphate kinase 3 (.1)
Lus10023076
220 / 3e-71
AT4G23900 189 / 4e-59
Nucleoside diphosphate kinase family protein (.1)
Lus10024286
190 / 1e-61
AT4G09320 272 / 3e-95
Nucleoside diphosphate kinase family protein (.1)
Lus10003844
186 / 5e-60
AT4G09320 264 / 7e-92
Nucleoside diphosphate kinase family protein (.1)
Lus10014787
186 / 8e-59
AT5G63310 301 / 2e-104
NDP KINASE 1A, NUCLEOSIDE DIPHOSPHATE KINASE IA, ARABIDOPSIS NUCLEOSIDE DIPHOSPHATE KINASE 2, nucleoside diphosphate kinase 2 (.1)
Lus10017931
186 / 9e-59
AT5G63310 301 / 3e-104
NDP KINASE 1A, NUCLEOSIDE DIPHOSPHATE KINASE IA, ARABIDOPSIS NUCLEOSIDE DIPHOSPHATE KINASE 2, nucleoside diphosphate kinase 2 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G091001
361 / 6e-128
AT4G23900 335 / 2e-117
Nucleoside diphosphate kinase family protein (.1)
Potri.003G140600
360 / 1e-127
AT4G23900 332 / 2e-116
Nucleoside diphosphate kinase family protein (.1)
Potri.014G049900
193 / 4e-63
AT4G09320 242 / 2e-83
Nucleoside diphosphate kinase family protein (.1)
Potri.012G093800
193 / 1e-61
AT5G63310 281 / 1e-96
NDP KINASE 1A, NUCLEOSIDE DIPHOSPHATE KINASE IA, ARABIDOPSIS NUCLEOSIDE DIPHOSPHATE KINASE 2, nucleoside diphosphate kinase 2 (.1)
Potri.002G138800
189 / 3e-61
AT4G09320 271 / 5e-95
Nucleoside diphosphate kinase family protein (.1)
Potri.003G068100
76 / 4e-17
AT1G17410 229 / 1e-77
Nucleoside diphosphate kinase family protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00334
NDK
Nucleoside diphosphate kinase
Representative CDS sequence
>Lus10001824 pacid=23154504 polypeptide=Lus10001824 locus=Lus10001824.g ID=Lus10001824.BGIv1.0 annot-version=v1.0
ATGAGCTCTCACATTTTCAGATCTGCTTCCAGAGCCGCCAGGTCCCTCCTCTCCTCTGCTTCCAGGGGCTCTCGCTTTTACTCCGGGGGACGAGCGGCTG
CTGTTCCGTTTAGAGGAAAGCTGCCACTCTTTGCTTCGGCTTATGAAAACACAGGTTCTGCAAATGTTCCGCAATGGATCTCTGGAGCCCTTGCGATTCC
CATGGCAACTTACATGCTCCAAGAACAGGAGGCCTGTGCTGGAGAGCTGGAGCGCACTTTCATTGCTATCAAGCCCGACGGTGTGCAGAGAGGATTGGTG
TCAGAAATCATTTCCCGTTTCGAGCGCAAAGGTTTCAAGCTTGTGGCGATCAAGATAGTGGTTCCATCAAAGGACTTCGCTCAAAAGCACTATCATGACC
TGAAAGAGAGACCATTTTTCAACGGTCTGTGCAACTTCCTTAGCTCTGGTCCTGTTGTGGCCATGGTCTGGGAAGGAGAAGGGGCGATCAAATATGGCAG
GAAGCTGATTGGAGCTACCGATCCCCAGAAATCTGAACCTGGAACAATTAGAGGTGATCTGGCCGTGGTTGTTGGAAGGAACATCATTCATGGAAGTGAT
GGCCCAGAAACCGCCAAGGACGAGATCAAGTTGTGGTTCAAGCCAGAAGAACTGGTTAGCTACACAAGCAATGCTGAGAAATGGATATACGGAAACAACT
GA
AA sequence
>Lus10001824 pacid=23154504 polypeptide=Lus10001824 locus=Lus10001824.g ID=Lus10001824.BGIv1.0 annot-version=v1.0
MSSHIFRSASRAARSLLSSASRGSRFYSGGRAAAVPFRGKLPLFASAYENTGSANVPQWISGALAIPMATYMLQEQEACAGELERTFIAIKPDGVQRGLV
SEIISRFERKGFKLVAIKIVVPSKDFAQKHYHDLKERPFFNGLCNFLSSGPVVAMVWEGEGAIKYGRKLIGATDPQKSEPGTIRGDLAVVVGRNIIHGSD
GPETAKDEIKLWFKPEELVSYTSNAEKWIYGNN
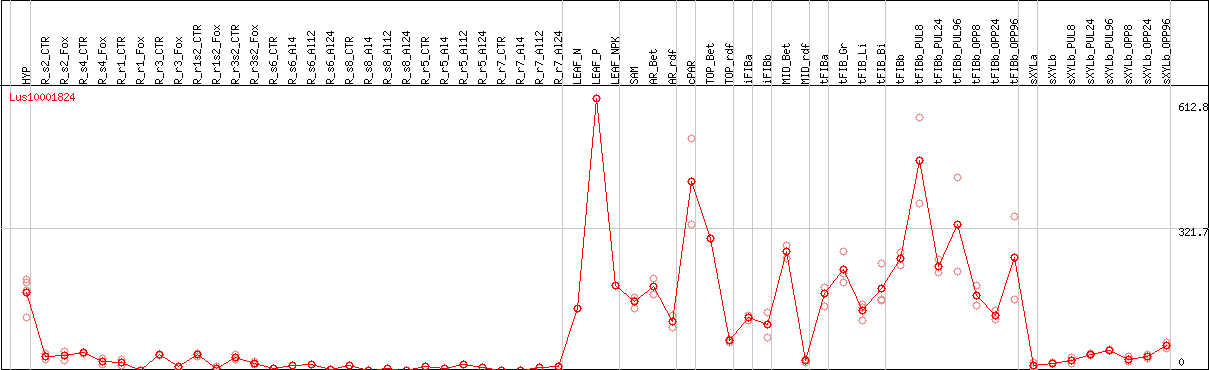
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10001824 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.