External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G05750 285 / 7e-94
PDE247, CLB19
pigment defective 247, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G20540 176 / 2e-51
MEF21
mitochondrial editing factor 21 (.1)
AT2G33760 177 / 3e-51
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT2G02980 176 / 5e-51
OTP85
ORGANELLE TRANSCRIPT PROCESSING 85, Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G77170 174 / 5e-51
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G59720 176 / 7e-51
CRR28
CHLORORESPIRATORY REDUCTION28, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G15930 176 / 1e-50
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G28640 173 / 2e-50
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G33680 176 / 4e-50
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G11290 175 / 1e-49
CRR22
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013341
385 / 5e-133
AT1G05750 525 / 0.0
pigment defective 247, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10031028
189 / 2e-55
AT4G19191 727 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10026886
178 / 7e-52
AT2G44880 376 / 3e-124
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Lus10030125
177 / 7e-52
AT2G20540 618 / 0.0
mitochondrial editing factor 21 (.1)
Lus10033858
178 / 2e-51
AT1G08070 422 / 4e-139
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10010385
178 / 2e-51
AT4G37380 758 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10007341
176 / 3e-51
AT2G42920 610 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Lus10033026
179 / 4e-51
AT1G08070 921 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10030053
178 / 4e-51
AT1G08070 884 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G217600
319 / 1e-106
AT1G05750 548 / 0.0
pigment defective 247, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.018G071800
193 / 1e-56
AT1G08070 606 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.007G050200
188 / 3e-55
AT4G37380 817 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.012G135600
187 / 7e-55
AT4G38010 672 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Potri.011G052300
184 / 4e-54
AT2G33760 747 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.018G040100
186 / 5e-54
AT1G08070 974 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.004G125500
184 / 1e-53
AT3G29230 829 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.004G043400
183 / 1e-53
AT2G33760 775 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.013G103600
184 / 2e-53
AT1G08070 477 / 7e-160
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.003G191000
183 / 2e-52
AT1G11290 1178 / 0.0
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10001852 pacid=23158795 polypeptide=Lus10001852 locus=Lus10001852.g ID=Lus10001852.BGIv1.0 annot-version=v1.0
ATGCGCCTCTCCGCCGTGGAACCAAATCACGTCACCTTCGTAACTTTACTCTCCGCATGCGCCGATCACCCATCAGAAGGCAGATCCTTCGGCCGCGTGA
TTCACTCCTACCTCCGAAAATTGGGCCTGGATAGGAACCAAGTGGTGGTCTGGACGGCACTGGTCAAAATGTACGCCAAGTACGGCGAGTTGGAGCTCGC
TAGGGTCTGCTTCGACGAGATCAAGGGTCTGTTCGGGGAAGCGATCGAGTCGTTTCGGGAGATGCAGGCTGCCAAAGTGGTCCCCGATTACGTGACTGTA
ACCTCTGTTCTCGCCGCCTGTGCTAATCTTGGCGCACTCGGGTTGGGATTATGGATCCATCGCTATATTCTACACCAAGAGTGGAGAAACAATGTAAGGA
TACAGAACTCGTTGATCGACGTGTATGCCAGATGCGGATGTGTCGAATTAGCCCGCCAAGTGTTTGACAAAATGCCCCAAAGAACTCTGGTCACCTGGAA
TTCGATGATCGTCGGTCTGGCTACAAACGGATTCGCGGAACAAGCTCTGGAATGCTTCAAAAAAATGCGAAAAGAAGGATTCAATCCTGATGGAGTAGGC
TTCACCGGTGCTCTTACAGCCTGTAGTCATGCCGGTTTCATCCAGGAAGGGTTACAGTATTTCGACCTCATGACGAAACAGTACAAGATCCCTCCCCGGA
TCGAACACTATGGCTGCATTGTGGATCTTTACAGCAGAGCAGGGGAATTGGAAGCTGCAATGAACGTAGTGGAAAGTATTGAGTATGAAGCCAAACGACG
TCGTATTAGGGTCACTACTTACAGCTTGCAAGAGTTGTAA
AA sequence
>Lus10001852 pacid=23158795 polypeptide=Lus10001852 locus=Lus10001852.g ID=Lus10001852.BGIv1.0 annot-version=v1.0
MRLSAVEPNHVTFVTLLSACADHPSEGRSFGRVIHSYLRKLGLDRNQVVVWTALVKMYAKYGELELARVCFDEIKGLFGEAIESFREMQAAKVVPDYVTV
TSVLAACANLGALGLGLWIHRYILHQEWRNNVRIQNSLIDVYARCGCVELARQVFDKMPQRTLVTWNSMIVGLATNGFAEQALECFKKMRKEGFNPDGVG
FTGALTACSHAGFIQEGLQYFDLMTKQYKIPPRIEHYGCIVDLYSRAGELEAAMNVVESIEYEAKRRRIRVTTYSLQEL
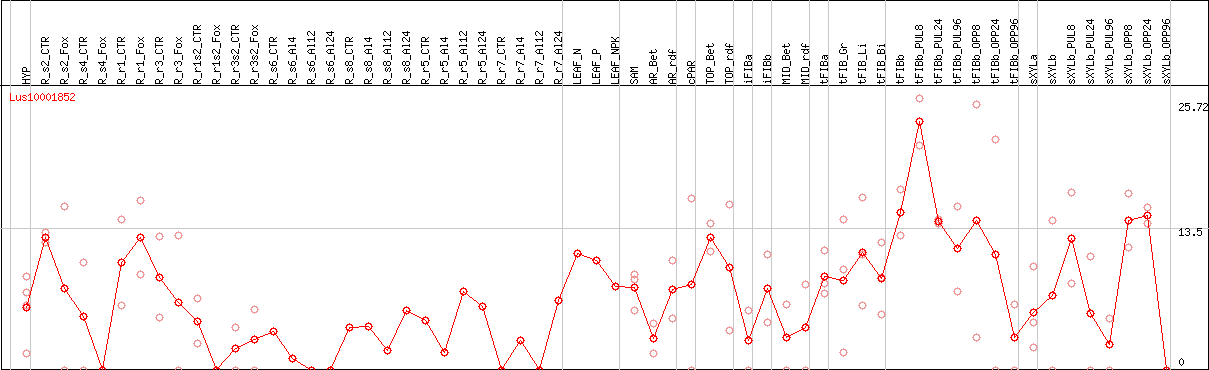
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10001852 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.