External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G22400 121 / 5e-32
ATUGT85A1, UGT85A1
ARABIDOPSIS THALIANA UDP-GLUCOSYL TRANSFERASE 85A1, UDP-Glycosyltransferase superfamily protein (.1)
AT1G78270 118 / 6e-31
ATUGT85A4
UDP-glucosyl transferase 85A4 (.1)
AT1G22340 115 / 8e-30
ATUGT85A7
UDP-glucosyl transferase 85A7 (.1)
AT1G22380 112 / 7e-29
ATUGT85A3
UDP-glucosyl transferase 85A3 (.1)
AT1G22370 111 / 2e-28
ATUGT85A5
UDP-glucosyl transferase 85A5 (.1.2)
AT1G22360 107 / 4e-27
ATUGT85A2, AT2
UDP-glucosyl transferase 85A2 (.1.2)
AT3G02100 65 / 3e-12
UDP-Glycosyltransferase superfamily protein (.1)
AT3G11340 57 / 2e-09
UGT76B1
UDP-dependent glycosyltransferase 76B1, UDP-Glycosyltransferase superfamily protein (.1)
AT5G05860 56 / 6e-09
UGT76C2
UDP-glucosyl transferase 76C2 (.1)
AT3G46660 55 / 1e-08
UGT76E12
UDP-glucosyl transferase 76E12 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013922
180 / 5e-54
AT1G22400 533 / 0.0
ARABIDOPSIS THALIANA UDP-GLUCOSYL TRANSFERASE 85A1, UDP-Glycosyltransferase superfamily protein (.1)
Lus10013923
173 / 3e-51
AT1G22400 537 / 0.0
ARABIDOPSIS THALIANA UDP-GLUCOSYL TRANSFERASE 85A1, UDP-Glycosyltransferase superfamily protein (.1)
Lus10013924
137 / 6e-38
AT1G22400 551 / 0.0
ARABIDOPSIS THALIANA UDP-GLUCOSYL TRANSFERASE 85A1, UDP-Glycosyltransferase superfamily protein (.1)
Lus10013920
133 / 2e-36
AT1G22400 488 / 3e-170
ARABIDOPSIS THALIANA UDP-GLUCOSYL TRANSFERASE 85A1, UDP-Glycosyltransferase superfamily protein (.1)
Lus10039277
130 / 3e-35
AT1G22400 499 / 3e-174
ARABIDOPSIS THALIANA UDP-GLUCOSYL TRANSFERASE 85A1, UDP-Glycosyltransferase superfamily protein (.1)
Lus10000993
122 / 2e-32
AT1G22380 563 / 0.0
UDP-glucosyl transferase 85A3 (.1)
Lus10013925
119 / 3e-31
AT1G22380 560 / 0.0
UDP-glucosyl transferase 85A3 (.1)
Lus10024584
118 / 7e-31
AT1G22400 570 / 0.0
ARABIDOPSIS THALIANA UDP-GLUCOSYL TRANSFERASE 85A1, UDP-Glycosyltransferase superfamily protein (.1)
Lus10031388
118 / 7e-31
AT1G22400 644 / 0.0
ARABIDOPSIS THALIANA UDP-GLUCOSYL TRANSFERASE 85A1, UDP-Glycosyltransferase superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10001880 pacid=23179017 polypeptide=Lus10001880 locus=Lus10001880.g ID=Lus10001880.BGIv1.0 annot-version=v1.0
ATGACTTCAACTGTCGCCGCCGTCGAGAAATCACCGCCGCACGTCATCTGCGTCCCATACCCGACGCAAGGCCACATAAACCCGATGCTCCACGTGGCAA
AACTTCTCCATTCCCGCGGTTTCCATGTCACCTTCGTTAACACGGACTACAACCACAAGCGCCTGCTCAAGTCTTGGGGCACCGGTGGCACCGGCCGCTC
CTCCTCCCCTTCGAGGGACTTGGTGCACAGGCTAAACGACGGGGACAGTGTCGGTTCGCCACGTGTCAGTTGTATTGTGTCCGACGCGGCCATGGCATTC
ACGTTGGATGTGGCTAGAGAGCTCGGGATTCCTGACGCGCTCTTCCTAGCTCTGAGCGCGTGTGCCAACTTGGCCTTGCTCAGCTACCCTGTCCTCGTCG
ACAGAGGCTTGGTGCCTCTTAAAAGTGCGATGAATTGCTCCTATGCTTGTAGAGAATGGGGGATTGAGATGGACATTGGTTCGGAAGTGAAGAGGGGGGA
AGTTGAGAAACTTGTGAGGGAGGTGATGGGTGGGGAGAAAGGAAAGGAGATGAAGTGGAAAGCTATGGAATGGAAAGTTAAGGCGGAAGAAGTTATTCAA
CCTGATAGATTTTCGTTTCAAAATCTGAACAAACTGATCGAAATATTGATGAAGAACACAACTGATTGA
AA sequence
>Lus10001880 pacid=23179017 polypeptide=Lus10001880 locus=Lus10001880.g ID=Lus10001880.BGIv1.0 annot-version=v1.0
MTSTVAAVEKSPPHVICVPYPTQGHINPMLHVAKLLHSRGFHVTFVNTDYNHKRLLKSWGTGGTGRSSSPSRDLVHRLNDGDSVGSPRVSCIVSDAAMAF
TLDVARELGIPDALFLALSACANLALLSYPVLVDRGLVPLKSAMNCSYACREWGIEMDIGSEVKRGEVEKLVREVMGGEKGKEMKWKAMEWKVKAEEVIQ
PDRFSFQNLNKLIEILMKNTTD
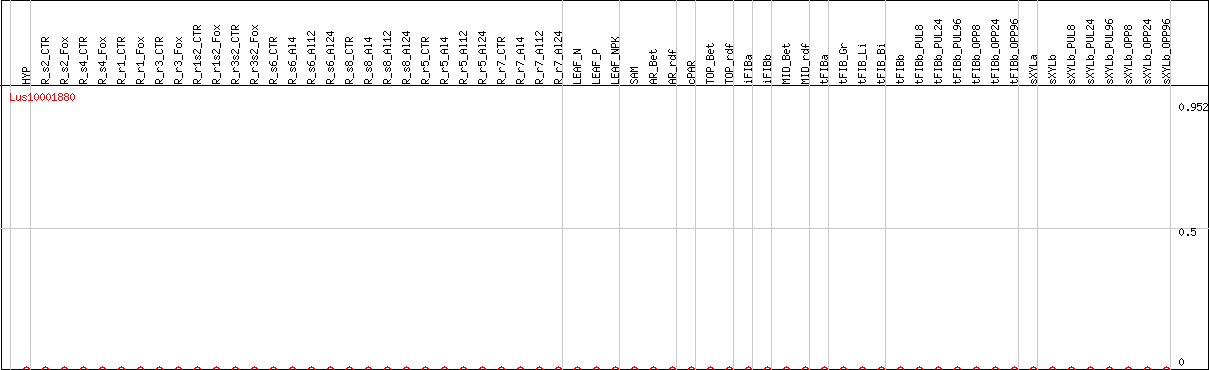
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10001880 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.