Lus10001897 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10001897 pacid=23179011 polypeptide=Lus10001897 locus=Lus10001897.g ID=Lus10001897.BGIv1.0 annot-version=v1.0
ATGCCGCTGGTTAGGTTTCAGGTCAGGAACCAGTACGCTTTGGGCCAGCCGGAGCTCTTCAAGGAGGCCAACATTGATGATCCCAAGGCCGTCCTCGACG
GCGTTGCCGTCGCCGGCCTCGTCGGGATCTTGCGTCAATTAGGCGATCTCGCTGAATTTGCAGCAGAGGTGTTTCACGGATTGCAGGAGCAGCTAACGAC
GACAGCTTCTCGGAGTCAGAAGTTGAAGCTTCGAGTGCAGAACATTGAAGCTGCGTTGCCTCCTATTGAGAAGGCTATATTGGCACAGAATAGCCACATC
CATTTTGCTTACACCGCTGGTTCTGAATGGCATCCTCGACTGCGGAATGAGCAGAATCACTTCATCTACAATGACTTGCCCCGTTTCATTATGGATTCTT
ATGAAGAATGTCGCGATCCGCCACGCTTTCATATACTTGACAAATTTGACACTGGTGGTCCAGGATCGTGTTTGAAGAGGTATTCAGATCCAACCTTCTT
TCGAAGAGCGTCAGGTGGGTTCAAGGAACATCATGCTGAGAAAGTTCAAAGAGATAAGAAGGCTCGTAAAATCAAGAAAAAGAGATCACTTCAAAGGAAA
GGTAGCTCACTGCAGAATGGGTCAATCTCAAGTCACAGCGAGAGATCCCAGTTTTTTCAGCCAACAGTTAATGGACGGAGTTCTTCTTCACATACGGCCT
CTACTATCGACTTGCCATTGAAATCAGAGATTGGTGATCATTCAAATTCTATTGATTCAGGAAATGGGTCAAATCATATTGAATGTGTTTTCCGTCCTAG
TTCTTCTGCACAACTCGAGGAACAGAAGTCTAATGGATTCTCTTCAAGGTCATTCCAGGGTGATGATTCCCGTGATTCGGGCTTTTCAGAGGAACAGTCC
AGGGTTGTTGATGATACTTCTCTGCATTGTTCCTCACCAAACCAGATTCACGATAGGCCCCATGGTGTTACTTGGGATGAAAAGGAAGAACGCGAAGATC
CTGGAGTTGCTGATGATAGTTTTCTGCATTGTTCCTCACCAAAGCAGATTCACCATAGGTCCCCTGGTGTTACTTGGGTTGGAAAGGAAGAACACAAAGA
GCCTGGAGTTGCTGATGATAGTTTTCTGCATGGTTCCTCACCGGAGCTGATTCCCATTAGTTCCTCTTGTCGTACCTGTGAAGAAAAGGCAGAAGGTGCA
CAGCCTGAAACTCATAAAGGTTCTCTACCTAGGTCCTCACTGGAGTGGCTCCCCCCTGGTTTCTCCTGTGTCTCTTGGGATGAAAAAGAAGAATTATCAG
AGGCTGAAATCCAACACTATGATGGAGATGAAGCTCCAGATGTTTTGACTGTAGACTCTGTCTTAGATATGCGAGAATCAGGGACCAATACTGAAATTCA
TCATCCAACAGACCTCAGATTTGAGGATGACAATGTTCGAGGGCCAAGTTTCACTGCCGTCACCCTGCTTGAAGATGTTGAAAGTGAAACTGACAATTTC
ATGGACGCTCTAAATTCAATTGATACAGAATCTGAGAGCGAACTTGATTATCAGACGAAACATGAAGTGAAAGAAGTTGAATGTGCTAACAGGGATAATG
AGGGCTTAGCAGTTACTTCAGGTCAGACAAACCAGGAAGTGAAACAAGTTGAATGTGCAATTGATGAACGGAGGACAGAAGATCAGCAGGACGAGGTCTC
TGGAGCTACTTCAGGTCATCATCCACATCATGATAGCTACTCCCCGCCCGATTTTTCTTCAAATCATGGAATGGCTTCTGATGTACTCATCCCAGTTCCT
TTGGTCAGTCCTGATGAGGAACAAACTCATTCTTCTGAATCATCATCTACTGCTTCCGGTTCTGAATCAGATATACTGGCTTCTACAACTGCGGATGCCC
CTGGTGCCTCAAAAGTGGAATCTCCTGTTCTTAACCGGTCATCTGTGATTGTAGTTAACGGAGTAAAGGAACTTTTGGCGGGTGCAGTTCTAAAGGATTC
CGGGAAGCCTCGACAATCTTCTGCCGCTGAGTTATCCCAAGTAGATGGACCATCTGTGGTTGTAGTTAAAGAAGTCGAGGAACCATTGGCTGGTATCGTC
CTAACTGATCCCAGAAAGCCTCGAGAATCTTCTATCACTGAGTTCTCCCATTTGGAACCAATCAGGTTCTGGACTAATGGGGGCCTCTTAGGACTTCAGC
CATCAAAGCCTCCCGATTTCAACGGGTCAAATCCTGGTAGTCAAAGTTCTGCAACAAATACATCTGAGGTGGTAAACACCCCAATACCGAGCAACAACAC
TGACAGTTCTGAAAGACCTGCTGGTTCCAGAATTTCTGAATCGTGCCATGACGTTAATGTTGCCAAAGCTAAAGATGCAATCGATCCTAACTACAGGAGT
AGATTCCATCCGTATGACAGAGGATTGAATGCCAATAGTCCAGTCGCTCCTGGAAATGGATTGCCACTTGGTTCATTCATCAAATTTACTACATCCGGAG
GAGCCAGTCATGAAAGCGATGGATACCCATCTCAAAATATGTTTGGACTTGGGCAAAGATTACTTGCTAACGGCTTTGGCAGAAGGATATCACTTCTGCC
CAGTAGTGAATCTGAGCTGACTAGTTCATTACCCCAACCTGCCGAGTCGGAGAAAAAAGGTAGGCAACCCTTGATTAAATTTCAACCGAGTCAACCAGAC
GATCTCAACATACGGTTTGGACACAAGCCTTCAGATGCAGGCTACCTTACATCTTCACCACCAGTTGAGCACATGAAATTATCTTTCCAACCCACCTGCG
GTTCCGAGGCTTCGAAACTGCAGCTGAAATTTCCTGACGGTGGCTGCTGCAACGAAAGTACTACGGATATGTTTCCATCGTTTCAGTTGGTTCCAGATGC
TGCAACCCGACGAAATGATGCCATCTCCGACTCGGATGATGACACATTCTGCAGATCATCTCCTTACTTGTCAGATGATTGTCTTAGCCATCACTCCGAC
ACGGGTTCTGATCAGTGGGAAGCTGGTCAGTCCCCTGAAAGCGTCCATAACAGTGACCAGGATGTTTTATGTGCATTCGAATCCGTTCCAGCTCCCTTGA
AGCTTGGTGACATGGAAACCAATGGCCTTTCTACAAGTCCTGCAGACGAAACTGAACTCGATCTTCCTAGTTTCGACTCTGTCAAGCTGCCGTTTGAGCA
AGTAAGACAAAAAGAAGACCGGTTATCAAGTCCCTTGCCTCCACCTCTCCCCCCAGCACAATGGCGCCTATCGAAAGAATCAAAACCGGCATCTGATACA
ATAGTAAGCAAACCACAAGCGATACCTGATGCTCCCCAGATCGCATTCGTTCCAAAACTTGTTGCAACTCCTACAACTCAGCAACCTAAACCAGCTCCAG
CCACTGACCAACAGGCTCATCCGGTCATCGTTCCATTCACCCCGAACAGAAAGGATGAACCGAAGTTGAACTGGTTGAAAGATGATAATCCACTCACAAA
TGGGAAGGTGATGCACGAACAAGAAGATTTCTTGCAGCAAATTAGATCAAAGTCATTTTCCCTGAGACCCACTGCACCAGTAAAGACAGCGGTGGCCGGT
GCAACTGCAATCGACACAGTGACCGAAATCCTGAAGAAGGCCAGTGCAATTCACCAGGTAGTTGCGAGCGACGTTGATGACGACGATGCATGGAGCGACC
CGTGA
|
||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10001897 pacid=23179011 polypeptide=Lus10001897 locus=Lus10001897.g ID=Lus10001897.BGIv1.0 annot-version=v1.0
MPLVRFQVRNQYALGQPELFKEANIDDPKAVLDGVAVAGLVGILRQLGDLAEFAAEVFHGLQEQLTTTASRSQKLKLRVQNIEAALPPIEKAILAQNSHI
HFAYTAGSEWHPRLRNEQNHFIYNDLPRFIMDSYEECRDPPRFHILDKFDTGGPGSCLKRYSDPTFFRRASGGFKEHHAEKVQRDKKARKIKKKRSLQRK
GSSLQNGSISSHSERSQFFQPTVNGRSSSSHTASTIDLPLKSEIGDHSNSIDSGNGSNHIECVFRPSSSAQLEEQKSNGFSSRSFQGDDSRDSGFSEEQS
RVVDDTSLHCSSPNQIHDRPHGVTWDEKEEREDPGVADDSFLHCSSPKQIHHRSPGVTWVGKEEHKEPGVADDSFLHGSSPELIPISSSCRTCEEKAEGA
QPETHKGSLPRSSLEWLPPGFSCVSWDEKEELSEAEIQHYDGDEAPDVLTVDSVLDMRESGTNTEIHHPTDLRFEDDNVRGPSFTAVTLLEDVESETDNF
MDALNSIDTESESELDYQTKHEVKEVECANRDNEGLAVTSGQTNQEVKQVECAIDERRTEDQQDEVSGATSGHHPHHDSYSPPDFSSNHGMASDVLIPVP
LVSPDEEQTHSSESSSTASGSESDILASTTADAPGASKVESPVLNRSSVIVVNGVKELLAGAVLKDSGKPRQSSAAELSQVDGPSVVVVKEVEEPLAGIV
LTDPRKPRESSITEFSHLEPIRFWTNGGLLGLQPSKPPDFNGSNPGSQSSATNTSEVVNTPIPSNNTDSSERPAGSRISESCHDVNVAKAKDAIDPNYRS
RFHPYDRGLNANSPVAPGNGLPLGSFIKFTTSGGASHESDGYPSQNMFGLGQRLLANGFGRRISLLPSSESELTSSLPQPAESEKKGRQPLIKFQPSQPD
DLNIRFGHKPSDAGYLTSSPPVEHMKLSFQPTCGSEASKLQLKFPDGGCCNESTTDMFPSFQLVPDAATRRNDAISDSDDDTFCRSSPYLSDDCLSHHSD
TGSDQWEAGQSPESVHNSDQDVLCAFESVPAPLKLGDMETNGLSTSPADETELDLPSFDSVKLPFEQVRQKEDRLSSPLPPPLPPAQWRLSKESKPASDT
IVSKPQAIPDAPQIAFVPKLVATPTTQQPKPAPATDQQAHPVIVPFTPNRKDEPKLNWLKDDNPLTNGKVMHEQEDFLQQIRSKSFSLRPTAPVKTAVAG
ATAIDTVTEILKKASAIHQVVASDVDDDDAWSDP
|
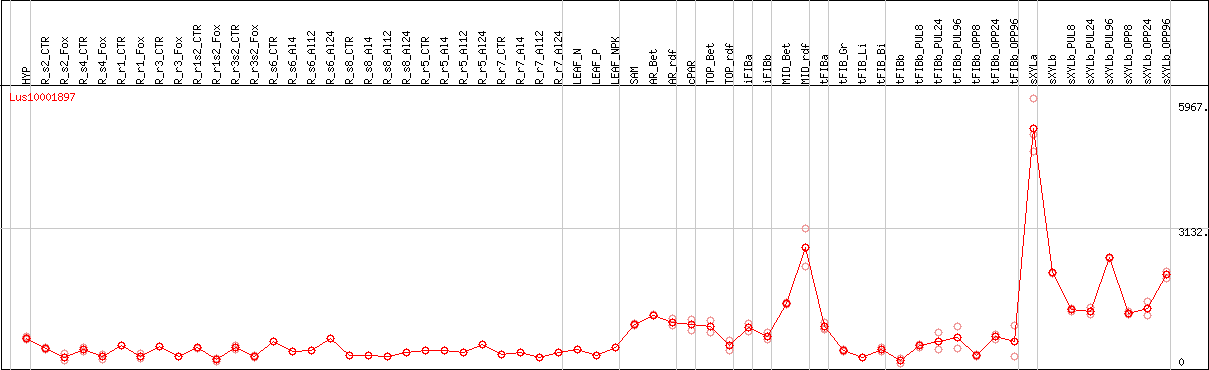
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10001897 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
