Lus10001926 [FLAX]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||
|
Representative CDS sequence |
>Lus10001926 pacid=23175274 polypeptide=Lus10001926 locus=Lus10001926.g ID=Lus10001926.BGIv1.0 annot-version=v1.0
ATGAGCTCCTCCAACAGCAAATCGCGAAACTTTCGCCGTCGGCTGGACGACGAAGACTCTAGCTCAACTGACCCTTCCGCCGCCATCCCTTCACCGCCCA
CCGTCCGCAGACCTCAACCACCTTCTTCCTCAGCTTCCAAGCCTAAAAAGCTCCTTAGTTTCGCCGACGACGAAGGCGATGAAGGTGCTGTAACTCCTCT
CAATCATTCTTCTCTCTCTAAGTCTTCTTCTTCTTCTAGCCGGCGAAAGCCTCCTCCTCCTTCTTCTTCCGGTTCTTCTTCTAGACTCGGCAGGCCCAGT
TCTGCTCACAAGCTCACCGCATCCAGAGACCGAGCTCCTCTTAGCTCCTCCGCTACGACTTCCAATGTTCTTCCTCAGGCCGGAACTTACACCAAAGAAG
CCATTCTCGAGCTTCAGAAGAACACCCGCACTTTAGCCAAACCTTCCCCCAAGCCTTCTCCTTCCTCTGAGCCTAGGATCATCCTTAGAGGCCTTCTCAA
GCCTGCTGCTCCATCCCCGAATCAAACCCAAACTCACCCCAACTCTGTCATAGATGATGATGATGATAATGATGACGGAGATGTGGCTGACGCCAATACT
CGATTGGCATCAATTGGGTTGATGAAATCCGCATCCCATGATGACTATCCTGATCCTGAAACCATTAGGAAGATCAGAGCCGAGAGAGAAAAGAAGAGAC
AGTCTAGAGCTTCTGCTGCTGACTTCATTCCCCTTAATGGACCTTCTCACACTGAAGCTGAAGGGTCCAGTGACGATGACGAGCTTGAGCTTAGAACTCG
TGTTGCTATGGTTGGCAAAAAAGAAAGCAAAACTCATGGCGTCTTTGAGGATGTTGACTCTGGTACTGCTGCTTCTGCCAGATATATGAATGCTCAGACC
TCGGTGAATGAGATTGCTGATGAAGTTGATGATGAGGATGAAGAGGACAAGATTTGGGAAGAAGAGCAGTTTAGGAAGGCCTTGGGGAAGAGAATGGATG
ATGGTTCTGCTACTACTCGAGTCGTTTCTGGTGGTGTAGGTGGTACTAATGCTACTGCCACGATGCAGCAGCAGCCTCCATTGCAGCATAGGGCCAGCAG
TGCATATTCAAACATTGGAGGGGGTCTTGGGGCATCTCAACGGCTGGACACCTTGACCATTCAGCAACAATCTGAGATTGCTTTGAAGGCATTGCAGGAT
AATTCCAGGAAGCTTCAGGAATCTCATGATAGAACTGTTTCAACACTAGCAAAAACTGACGAGAACTTATCTGCCTCGCTGGATAATATTACTTCTCTTG
AAGCCTCGTTGTCTGCTGCTGGTGAGAAGTTCATCTTTATGCAAAGTCTCCGTGACTTTGTTTCTGTTTTATGTGAATTTTTGCAGCATAAAGCTCCATT
CATAGAGGAGCTTGAAGAGCGAATGCAGAAGGTTCAAGAGAAACGGGCAGCAGCTGTTTTGGAAAGAAGAAGTGCTGATAATGATGATGAAATGGTGGGG
GTAGAGGCCGCTGTTAAAGCAGCAATGTCAGTTTTCAGGGAACGTGGAAATACTGCAATGATGATCGCTACTGCTGCAAGTGCAGCCCAGACTGCTTCAG
CTTCTGTGAAAGAGCAAGCTACTCAACCAGAATTAGATGAACTGGGTAGGGATATGAGTAGACAAAAGAATATGGAGATGAAACGGAGGGCTGAAGCACG
ACAACGCAGAAGATCCAAGTTTGATTCAAAGAGATTTTCATGCATGGAAGTTGACAACTCTGATCAGAAAATAGAAGGAGAATCAAGCACTGACGAGAGC
GAGAGTGAAAGTGAAGCTTACCGGTCTAGTCGCGATATGTGCCTACAAACTGCTGGTCAAATTTTAAGTGATGCTTCCGAAGAATTTTCCCAGTTGCAAT
TGGTGAAAGAAAGATTGGAGAATTGGAAAATTAAGTATGCAGCTAGCTACAGAGATGCTTACATGTCACTCAGTATACCAGCACTCTTCTCTCCTTATGT
GAGGCTGGAGCTTCTACAGTGGGATCCATTGGACAAGGCTGATTTCTCACACATGAAATGGCATTCTCTGTTATTCGACTATGGTTTACCAGAGGGTGCA
TCTGATTTGAATCCCGACGATGCTGACAGTAATCTCGTACCACAACTGGTAGAAAAAGTAGCTATTCCCAAATTGCATCACCAAATCGTCCACTGTTGGG
ACACACTTAGTACACTTGAAACGGAGAATGCCGTTGCTGCCACAAGCTTGGTCATCGATTACGTTCCAGCCTCAAGTCAGGCTCTAGCAGAGCTATTAGT
TGCAATCCGTACCCGTCTCGCTGATGCTGTAGCTGACATCATGGTTCCAAACTGGGGCTCGCTTGTCCTTAAGGCAGTGCCAAGTGCGGCACAAGTTGCA
GCCTATCGTTTTGGAGTGTCTGTCCGTTTGATAAGGAACATCTGCTTGTGGAAGGATATTCTTGCTTTGCCGGTTCTTGAGGAGCTCGCACTTGATGAGC
TTTTACGGAGGAAGGTGCTACCCCATGTTCAAAGCATTTCATTGGATGTTCATGATGCTGTTGCCAGAACGGAGAGGATCGTTGCTTCTCTGACTGGAGT
GTGGGCAGGTCCAAACGTAACGGGGGATCGCAGTGGCAAGTTGCAGTCAGTGGTGGACTTTGTGGTGTCGCTAGGAAGGGCGTTGGAGAAGAAGCAGGCG
GCGTCAAGAGTGAGCGAGGGGGAGAGTAATGGACTTGCGAGACGGTTGACGAAGCTGCTGGTTGAGCTCAACGACTACGACAATGCACGAGGCGTCGCGA
AGACCTTTCGGTTGAAGGACGCTTTGTAA
|
||||||||||||||||||||
|
AA sequence
|
>Lus10001926 pacid=23175274 polypeptide=Lus10001926 locus=Lus10001926.g ID=Lus10001926.BGIv1.0 annot-version=v1.0
MSSSNSKSRNFRRRLDDEDSSSTDPSAAIPSPPTVRRPQPPSSSASKPKKLLSFADDEGDEGAVTPLNHSSLSKSSSSSSRRKPPPPSSSGSSSRLGRPS
SAHKLTASRDRAPLSSSATTSNVLPQAGTYTKEAILELQKNTRTLAKPSPKPSPSSEPRIILRGLLKPAAPSPNQTQTHPNSVIDDDDDNDDGDVADANT
RLASIGLMKSASHDDYPDPETIRKIRAEREKKRQSRASAADFIPLNGPSHTEAEGSSDDDELELRTRVAMVGKKESKTHGVFEDVDSGTAASARYMNAQT
SVNEIADEVDDEDEEDKIWEEEQFRKALGKRMDDGSATTRVVSGGVGGTNATATMQQQPPLQHRASSAYSNIGGGLGASQRLDTLTIQQQSEIALKALQD
NSRKLQESHDRTVSTLAKTDENLSASLDNITSLEASLSAAGEKFIFMQSLRDFVSVLCEFLQHKAPFIEELEERMQKVQEKRAAAVLERRSADNDDEMVG
VEAAVKAAMSVFRERGNTAMMIATAASAAQTASASVKEQATQPELDELGRDMSRQKNMEMKRRAEARQRRRSKFDSKRFSCMEVDNSDQKIEGESSTDES
ESESEAYRSSRDMCLQTAGQILSDASEEFSQLQLVKERLENWKIKYAASYRDAYMSLSIPALFSPYVRLELLQWDPLDKADFSHMKWHSLLFDYGLPEGA
SDLNPDDADSNLVPQLVEKVAIPKLHHQIVHCWDTLSTLETENAVAATSLVIDYVPASSQALAELLVAIRTRLADAVADIMVPNWGSLVLKAVPSAAQVA
AYRFGVSVRLIRNICLWKDILALPVLEELALDELLRRKVLPHVQSISLDVHDAVARTERIVASLTGVWAGPNVTGDRSGKLQSVVDFVVSLGRALEKKQA
ASRVSEGESNGLARRLTKLLVELNDYDNARGVAKTFRLKDAL
|
DESeq2's median of ratios [FLAX]
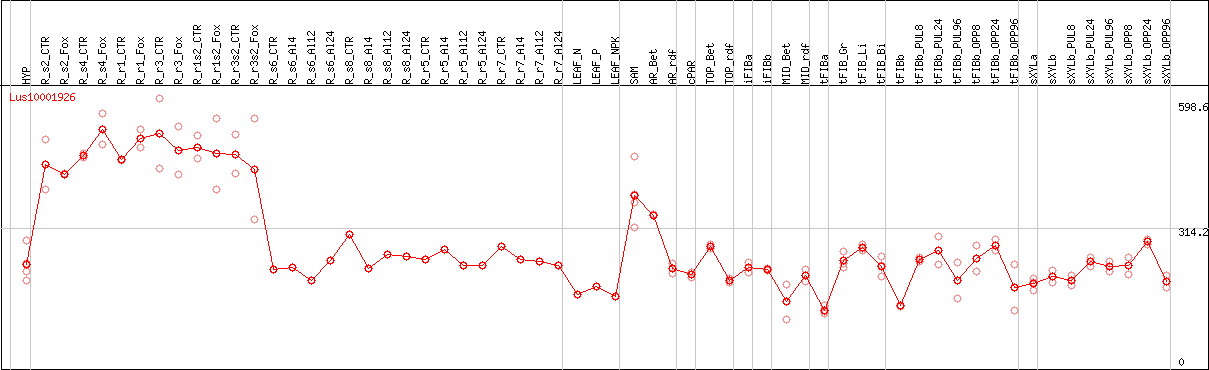
Coexpressed genes
Lus10001926 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
