External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G22410 276 / 2e-88
SLO1
SLOW GROWTH 1 (.1)
AT3G05240 184 / 3e-54
MEF19
mitochondrial editing factor 19 (.1)
AT2G29760 177 / 7e-51
OTP81
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G08070 175 / 4e-50
EMB3102, OTP82
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G20540 166 / 6e-48
MEF21
mitochondrial editing factor 21 (.1)
AT2G34400 166 / 3e-47
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
AT5G66520 162 / 7e-46
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G21065 160 / 4e-45
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT2G01510 158 / 1e-44
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G36730 155 / 6e-44
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10018223
184 / 2e-53
AT2G29760 931 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10040680
182 / 1e-52
AT2G29760 926 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10033026
179 / 2e-51
AT1G08070 921 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10020918
172 / 1e-49
AT2G29760 474 / 3e-159
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10004052
167 / 2e-47
AT1G31430 684 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Lus10002024
164 / 3e-47
AT5G56310 344 / 6e-113
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10002288
165 / 6e-47
AT1G31430 689 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Lus10007341
164 / 8e-47
AT2G42920 610 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Lus10014298
160 / 5e-45
AT5G66520 748 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G019500
317 / 3e-104
AT2G22410 791 / 0.0
SLOW GROWTH 1 (.1)
Potri.009G044700
190 / 1e-55
AT2G29760 1013 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.004G217900
180 / 2e-52
AT2G22410 530 / 0.0
SLOW GROWTH 1 (.1)
Potri.018G040100
180 / 4e-52
AT1G08070 974 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.018G071800
177 / 8e-51
AT1G08070 606 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.003G031600
176 / 1e-50
AT3G15930 784 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.017G001000
174 / 4e-50
AT4G21065 793 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.015G034900
167 / 1e-48
AT4G21065 370 / 2e-122
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.017G083900
166 / 4e-48
AT2G36730 552 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.007G073100
166 / 1e-47
AT5G08305 600 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10002552 pacid=23165488 polypeptide=Lus10002552 locus=Lus10002552.g ID=Lus10002552.BGIv1.0 annot-version=v1.0
ATGAAGCAGGTTCAGTCACAAATGATCCTCACGGAGGTAATCAGAGACAGGCTAGCTTCAAGTCGTTTAGTTGCGTTTTGTGCGATTTCAGAATCGAGGG
ACATTGATTACTGTATCTCGATTTTGAATTCGGTTGAGAACCCCATTTTGTTCTCGTGGAATTTGGCCTCCAGAGGTTGTGTAGAGAGTGGTAAGCCTGA
AGGGGCATTGGTTCTGTACAAGAGGATGTTGGGGCGCGGGTTTAGGCCTGATAGTTATACATACCCGTTGTTGTTGAATGCTTGTGCTGGCTTGGGATCT
TGTGATTTGGGTCTTCAGATTCATGGGCATGTTTTGAAGTTTGGGTTTGATGAGGTTGTGGTCGTGCAAAATGCTACGATCCATATGTTTGCCACGATTG
GGGAATTGGGGCTTGCACGCCAGGTGTTTGATGAAAGTTCTGTCAGAGATAAGGTTTCGTGGAATTCGTTGATTAATGGGTATGCTAAGAAGGGGAAGGC
AAAGGAGGCGATTGGTATTTATGATGAAATGATGGAGGAGGGAGTTAAGCCTGATGAGGTGACAATGGTTGGTGTTATTCTGTCATGTGCACAGTTGGAG
AACTTGAAACTAGGGAGAGGTTTTCATCGCTACATTGAAGAGAACAGGCTGAGATTAACGGTTCCCCTCACGAATGCGCTTTTGGATATGTACATGAAAT
GTGGTGATCTTGAGGCAGCAAGATCTCTGTTTAATATGTCAACAGAGAAAACTGTTGTATCATGGACTACAATGATTATTGGGTATTCGAAGTGA
AA sequence
>Lus10002552 pacid=23165488 polypeptide=Lus10002552 locus=Lus10002552.g ID=Lus10002552.BGIv1.0 annot-version=v1.0
MKQVQSQMILTEVIRDRLASSRLVAFCAISESRDIDYCISILNSVENPILFSWNLASRGCVESGKPEGALVLYKRMLGRGFRPDSYTYPLLLNACAGLGS
CDLGLQIHGHVLKFGFDEVVVVQNATIHMFATIGELGLARQVFDESSVRDKVSWNSLINGYAKKGKAKEAIGIYDEMMEEGVKPDEVTMVGVILSCAQLE
NLKLGRGFHRYIEENRLRLTVPLTNALLDMYMKCGDLEAARSLFNMSTEKTVVSWTTMIIGYSK
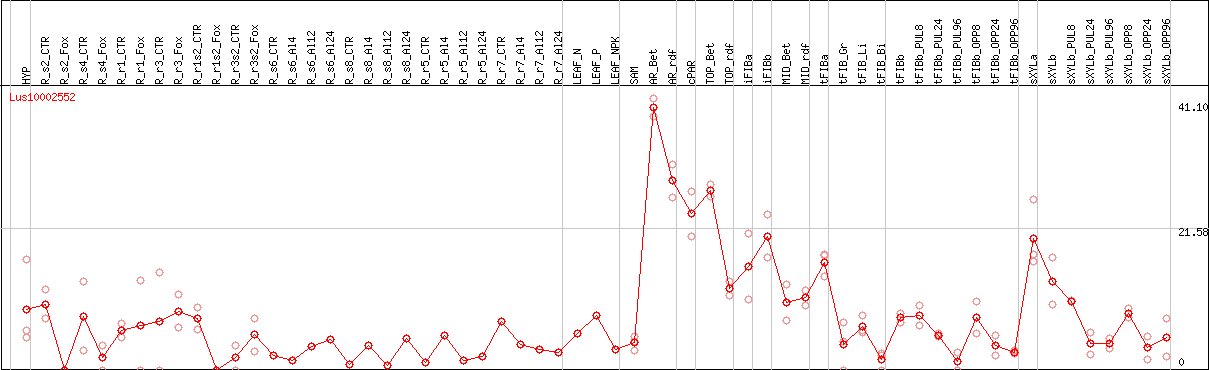
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10002552 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.