External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G44920 269 / 3e-92
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT1G12250 63 / 4e-12
Pentapeptide repeat-containing protein (.1.2)
AT5G55000 59 / 2e-10
FIP2
potassium channel tetramerisation domain-containing protein / pentapeptide repeat-containing protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10000520
329 / 3e-116
AT2G44920 286 / 6e-99
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10024602
58 / 5e-10
AT1G12250 354 / 1e-123
Pentapeptide repeat-containing protein (.1.2)
Lus10032239
58 / 5e-10
AT1G12250 357 / 9e-125
Pentapeptide repeat-containing protein (.1.2)
Lus10020270
52 / 1e-07
AT5G55000 430 / 2e-149
potassium channel tetramerisation domain-containing protein / pentapeptide repeat-containing protein (.1.2)
Lus10002622
49 / 5e-07
AT5G55000 462 / 1e-165
potassium channel tetramerisation domain-containing protein / pentapeptide repeat-containing protein (.1.2)
Lus10007788
42 / 0.0001
AT5G53490 301 / 2e-104
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2), Tetratricopeptide repeat (TPR)-like superfamily protein (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G200000
286 / 4e-99
AT2G44920 256 / 4e-87
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.008G058800
284 / 3e-98
AT2G44920 253 / 1e-85
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.003G112900
64 / 4e-12
AT1G12250 363 / 1e-128
Pentapeptide repeat-containing protein (.1.2)
Potri.013G062500
50 / 3e-07
AT5G55000 413 / 2e-146
potassium channel tetramerisation domain-containing protein / pentapeptide repeat-containing protein (.1.2)
Potri.019G038400
49 / 8e-07
AT5G55000 414 / 1e-146
potassium channel tetramerisation domain-containing protein / pentapeptide repeat-containing protein (.1.2)
Potri.012G018100
42 / 0.0002
AT5G53490 318 / 1e-110
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2), Tetratricopeptide repeat (TPR)-like superfamily protein (.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0505
Pentapeptide
PF00805
Pentapeptide
Pentapeptide repeats (8 copies)
Representative CDS sequence
>Lus10002712 pacid=23180557 polypeptide=Lus10002712 locus=Lus10002712.g ID=Lus10002712.BGIv1.0 annot-version=v1.0
ATGAATCTTCTTCTGAATGTCTCGCTCTGTACGGCTAAAACCCCACCAAAACCCTCACTTCCACTCCTTCCAAGCCCTCCTCAACCCCCACTTTCTCTTT
CTTTGCCCCGTTCCCAGGTGTTACGAGACTTGGCTAAAACAAGCTTGCTTGCAGTGCTATCCGCCTCTGTCTTCCTCTCTGACCCTGCTCTCGCATACAA
GGGAGGAGGTCCATACGGTTCTGAGGTGACCAGGGGCCAGGATCTGACTGGCAAGGACTTCAGTGGCAAGACTTTGATCAAGCAGGACTTTAAGACGTCA
ATTCTAAGGCAAGCCAATTTCAAAGGTGCAAAGTTGCTCGGGGCTAGCTTCTTTGATGCTGATTTGACTGGAGCCGATCTTTCGAATGCAGACCTTAGAG
GAGTAGATTTCTCCTTGGCCAATGTGACAAAGGTGAATTTGAGCAATGCGAACCTAGAAGGCGCTCTAACCACAGGCAATACTTCATTCAAGGGCTCAAA
CATAGCAGGAGCAGATTTCACTGATGTGCCTCTACGGGAGGATCAGCGTGAATACCTTTGCAAAATTGCAGATGGGGTGAACCCAACTACTGGAAACGCG
ACCCGTGAGACACTACTTTGTAACTAA
AA sequence
>Lus10002712 pacid=23180557 polypeptide=Lus10002712 locus=Lus10002712.g ID=Lus10002712.BGIv1.0 annot-version=v1.0
MNLLLNVSLCTAKTPPKPSLPLLPSPPQPPLSLSLPRSQVLRDLAKTSLLAVLSASVFLSDPALAYKGGGPYGSEVTRGQDLTGKDFSGKTLIKQDFKTS
ILRQANFKGAKLLGASFFDADLTGADLSNADLRGVDFSLANVTKVNLSNANLEGALTTGNTSFKGSNIAGADFTDVPLREDQREYLCKIADGVNPTTGNA
TRETLLCN
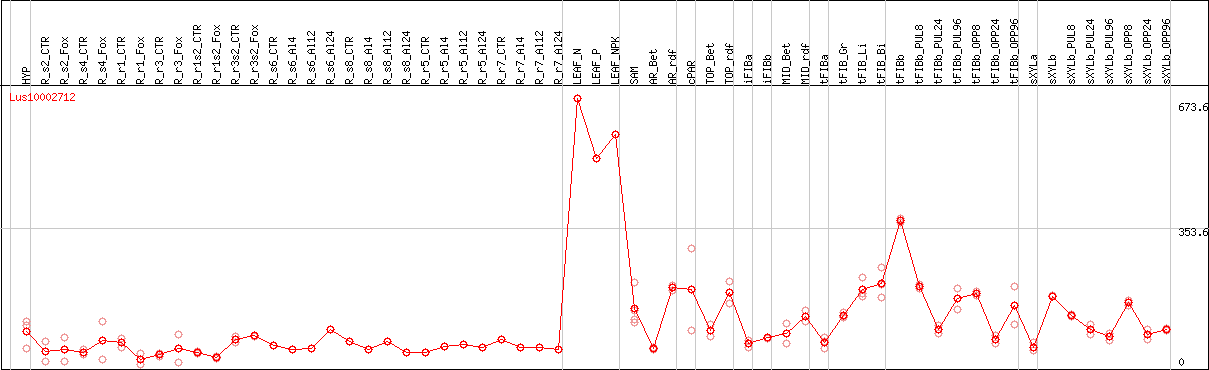
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10002712 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.