External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G04730 199 / 3e-63
AUX_IAA
IAA16
indoleacetic acid-induced protein 16 (.1)
AT3G23050 194 / 4e-61
AUX_IAA
AXR2, IAA7
AUXIN RESISTANT 2, indole-3-acetic acid 7 (.1.2)
AT1G04250 189 / 2e-59
AUX_IAA
IAA17, AXR3
indole-3-acetic acid inducible 17, AUXIN RESISTANT 3, AUX/IAA transcriptional regulator family protein (.1)
AT4G14550 189 / 3e-59
AUX_IAA
SLR, IAA14
SOLITARY ROOT, indole-3-acetic acid inducible 14 (.1)
AT4G29080 153 / 1e-44
AUX_IAA
IAA27, PAP2
indole-3-acetic acid inducible 27, phytochrome-associated protein 2 (.1)
AT5G43700 145 / 7e-43
AUX_IAA
IAA4, ATAUX2-11
indole-3-acetic acid inducible 4, AUXIN INDUCIBLE 2-11, AUX/IAA transcriptional regulator family protein (.1)
AT1G04240 145 / 8e-43
AUX_IAA
IAA3, SHY2
SHORT HYPOCOTYL 2, indole-3-acetic acid inducible 3, AUX/IAA transcriptional regulator family protein (.1)
AT5G65670 147 / 8e-42
AUX_IAA
IAA9
indole-3-acetic acid inducible 9 (.1.2)
AT2G22670 145 / 2e-41
AUX_IAA
IAA8
indoleacetic acid-induced protein 8 (.1.2.3.4)
AT3G15540 132 / 1e-37
AUX_IAA
MSG2, IAA19
MASSUGU 2, indole-3-acetic acid inducible 19 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014731
425 / 2e-151
AT3G04730 241 / 4e-80
indoleacetic acid-induced protein 16 (.1)
Lus10015907
190 / 3e-59
AT3G04730 309 / 5e-107
indoleacetic acid-induced protein 16 (.1)
Lus10024854
182 / 3e-56
AT3G04730 221 / 3e-72
indoleacetic acid-induced protein 16 (.1)
Lus10039414
177 / 3e-54
AT4G14550 305 / 1e-105
SOLITARY ROOT, indole-3-acetic acid inducible 14 (.1)
Lus10028222
165 / 3e-49
AT5G65670 322 / 3e-110
indole-3-acetic acid inducible 9 (.1.2)
Lus10042929
160 / 3e-46
AT5G65670 356 / 3e-122
indole-3-acetic acid inducible 9 (.1.2)
Lus10019241
155 / 5e-45
AT5G65670 356 / 4e-123
indole-3-acetic acid inducible 9 (.1.2)
Lus10039488
149 / 5e-44
AT5G43700 247 / 3e-84
indole-3-acetic acid inducible 4, AUXIN INDUCIBLE 2-11, AUX/IAA transcriptional regulator family protein (.1)
Lus10014729
149 / 6e-44
AT1G04240 230 / 2e-77
SHORT HYPOCOTYL 2, indole-3-acetic acid inducible 3, AUX/IAA transcriptional regulator family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G044900
213 / 2e-68
AT4G14550 286 / 1e-98
SOLITARY ROOT, indole-3-acetic acid inducible 14 (.1)
Potri.005G053900
210 / 3e-67
AT3G04730 317 / 3e-110
indoleacetic acid-induced protein 16 (.1)
Potri.013G041400
205 / 2e-65
AT3G04730 306 / 6e-106
indoleacetic acid-induced protein 16 (.1)
Potri.008G161200
191 / 5e-60
AT4G14550 343 / 1e-120
SOLITARY ROOT, indole-3-acetic acid inducible 14 (.1)
Potri.010G078300
192 / 8e-60
AT4G14550 319 / 1e-110
SOLITARY ROOT, indole-3-acetic acid inducible 14 (.1)
Potri.005G218300
187 / 1e-58
AT4G14550 292 / 7e-101
SOLITARY ROOT, indole-3-acetic acid inducible 14 (.1)
Potri.013G041300
159 / 5e-48
AT5G43700 239 / 5e-81
indole-3-acetic acid inducible 4, AUXIN INDUCIBLE 2-11, AUX/IAA transcriptional regulator family protein (.1)
Potri.001G177400
157 / 4e-47
AT3G04730 211 / 8e-69
indoleacetic acid-induced protein 16 (.1)
Potri.008G161100
154 / 8e-46
AT5G43700 238 / 3e-80
indole-3-acetic acid inducible 4, AUXIN INDUCIBLE 2-11, AUX/IAA transcriptional regulator family protein (.1)
Potri.005G053800
149 / 4e-44
AT5G43700 239 / 7e-81
indole-3-acetic acid inducible 4, AUXIN INDUCIBLE 2-11, AUX/IAA transcriptional regulator family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0072
Ubiquitin
PF02309
AUX_IAA
AUX/IAA family
Representative CDS sequence
>Lus10002724 pacid=23158702 polypeptide=Lus10002724 locus=Lus10002724.g ID=Lus10002724.BGIv1.0 annot-version=v1.0
ATGGAAGTAGCGGTGAAGTCGGCGACGACGACGACGGAGATGATGATGGGTTTTGATGAGACTGAGCTCCGACTTGGCCTTCCGGGCAATAATAGTATTC
TGAGGAAGAGAGGCTTCTCTGAAACGACGTCGGTGGATCTGCAGCTCAATCTTTCTTCTTCTCCGACCACCAAGGATGCGCCAGCTGTCAGTGACTGCAG
CACTACCACTACTACTACTACTGCTGCTCCTAAAAGCCTTCATCTTGCTGATCCAGCCACCAAGCCTCCTCCTCCCAAGGCACAAGTAGTGGGATGGCCG
CCAGTCCGATCTTTCAGGAAGAACATGATACAACCCACACCGCCACGTGCCCAGAAGAGCACCAGCCCCAATATTAAGGAGGAGGCGGTGGCAGAGAGCT
GCGGCGCCGCCGAGCAGAAGGTGGCGGCGGGGAGCTTTGTGAAGGTGAGCATGGACGGAGCTCCTTACCTGAGGAAGGTGGATTTGAGCATGTACGGCAG
CTACAGGGAGCTCTCCGATGCGCTGCTCAAAATGTTCGGCTCCTTCAACGGTGGTGGAGGAGAGCACTGCACCACCATGGATTTCATGAACGGAAGCAAG
CTTGGCGAGAAAAATAATGAAGATATTTTCAACGGCGGCGATGATTGTGTTCCAACCTATGAGGACAAAGACGGCGACTGGATGCTCGTTGGCGATGTCC
CTTGGGGGATGTTTGTGGAATCATGCAAACGGCTCCGCATCATGAAGGGCAAAGAGGCGATGGGTCTCGGCCATAACTACAGTCCCACCAAGAGCCATGG
AGAAACGCAAGAACAGGATGTAGCTAGCCAGCTGGTTGGCGGTGACGGTACCGGCTGGCTGGAGGCGATGCAGAATCTGCTCCGTTGA
AA sequence
>Lus10002724 pacid=23158702 polypeptide=Lus10002724 locus=Lus10002724.g ID=Lus10002724.BGIv1.0 annot-version=v1.0
MEVAVKSATTTTEMMMGFDETELRLGLPGNNSILRKRGFSETTSVDLQLNLSSSPTTKDAPAVSDCSTTTTTTTAAPKSLHLADPATKPPPPKAQVVGWP
PVRSFRKNMIQPTPPRAQKSTSPNIKEEAVAESCGAAEQKVAAGSFVKVSMDGAPYLRKVDLSMYGSYRELSDALLKMFGSFNGGGGEHCTTMDFMNGSK
LGEKNNEDIFNGGDDCVPTYEDKDGDWMLVGDVPWGMFVESCKRLRIMKGKEAMGLGHNYSPTKSHGETQEQDVASQLVGGDGTGWLEAMQNLLR
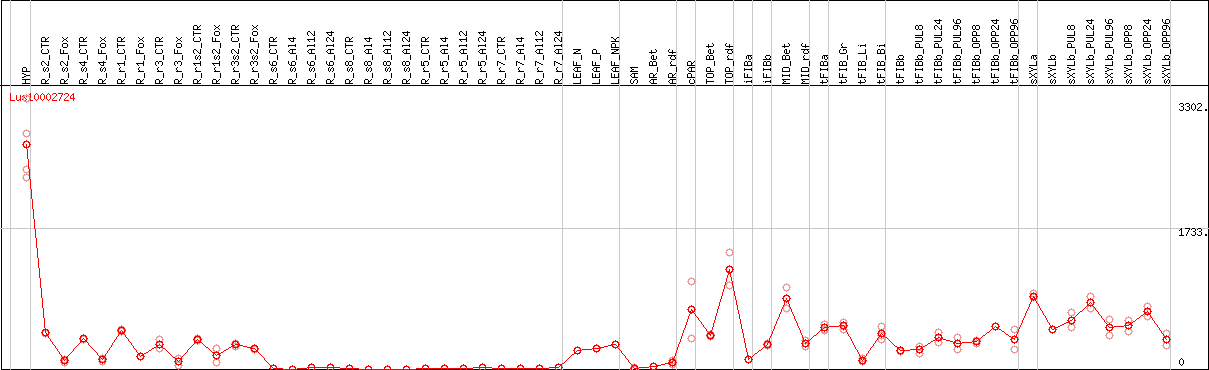
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10002724 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.