External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G25590 311 / 6e-101
Protein of unknown function (DUF630 and DUF632) (.1)
AT1G52320 278 / 3e-91
unknown protein
AT1G02110 113 / 4e-28
Protein of unknown function (DUF630 and DUF632) (.1)
AT4G35240 101 / 4e-24
Protein of unknown function (DUF630 and DUF632) (.1), Protein of unknown function (DUF630 and DUF632) (.2)
AT3G51290 101 / 4e-24
Protein of unknown function (DUF630) ;Protein of unknown function (DUF632) (.1), Protein of unknown function (DUF630) ;Protein of unknown function (DUF632) (.2)
AT4G39790 100 / 1e-23
Protein of unknown function (DUF630 and DUF632) (.1)
AT3G60320 99 / 3e-23
Protein of unknown function (DUF630 and DUF632) (.1)
AT2G17110 91 / 2e-20
Protein of unknown function (DUF630 and DUF632) (.1)
AT2G27090 90 / 4e-20
Protein of unknown function (DUF630 and DUF632) (.1)
AT2G34670 88 / 2e-19
Protein of unknown function (DUF630 and DUF632) (.1), Protein of unknown function (DUF630 and DUF632) (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10038627
447 / 4e-158
AT5G25590 538 / 0.0
Protein of unknown function (DUF630 and DUF632) (.1)
Lus10037901
427 / 4e-150
AT5G25590 627 / 0.0
Protein of unknown function (DUF630 and DUF632) (.1)
Lus10022116
387 / 5e-131
AT5G25590 753 / 0.0
Protein of unknown function (DUF630 and DUF632) (.1)
Lus10000014
289 / 5e-100
AT5G25590 209 / 3e-64
Protein of unknown function (DUF630 and DUF632) (.1)
Lus10025777
294 / 6e-95
AT1G52320 652 / 0.0
unknown protein
Lus10035887
291 / 1e-93
AT1G52320 747 / 0.0
unknown protein
Lus10028431
116 / 3e-29
AT3G51290 728 / 0.0
Protein of unknown function (DUF630) ;Protein of unknown function (DUF632) (.1), Protein of unknown function (DUF630) ;Protein of unknown function (DUF632) (.2)
Lus10041884
115 / 3e-29
AT3G51290 649 / 0.0
Protein of unknown function (DUF630) ;Protein of unknown function (DUF632) (.1), Protein of unknown function (DUF630) ;Protein of unknown function (DUF632) (.2)
Lus10042910
108 / 1e-26
AT3G60320 727 / 0.0
Protein of unknown function (DUF630 and DUF632) (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G036400
343 / 3e-113
AT5G25590 656 / 0.0
Protein of unknown function (DUF630 and DUF632) (.1)
Potri.006G244100
336 / 1e-110
AT5G25590 603 / 0.0
Protein of unknown function (DUF630 and DUF632) (.1)
Potri.003G054800
296 / 9e-96
AT1G52320 626 / 0.0
unknown protein
Potri.001G181100
292 / 2e-94
AT1G52320 615 / 0.0
unknown protein
Potri.005G112300
123 / 1e-31
AT3G51290 576 / 0.0
Protein of unknown function (DUF630) ;Protein of unknown function (DUF632) (.1), Protein of unknown function (DUF630) ;Protein of unknown function (DUF632) (.2)
Potri.007G056700
117 / 1e-29
AT3G51290 545 / 0.0
Protein of unknown function (DUF630) ;Protein of unknown function (DUF632) (.1), Protein of unknown function (DUF630) ;Protein of unknown function (DUF632) (.2)
Potri.014G047400
110 / 4e-27
AT3G60320 862 / 0.0
Protein of unknown function (DUF630 and DUF632) (.1)
Potri.004G177200
108 / 2e-26
AT1G21740 225 / 2e-62
Protein of unknown function (DUF630 and DUF632) (.1)
Potri.009G137500
107 / 3e-26
AT3G51290 228 / 4e-64
Protein of unknown function (DUF630) ;Protein of unknown function (DUF632) (.1), Protein of unknown function (DUF630) ;Protein of unknown function (DUF632) (.2)
Potri.002G137600
104 / 4e-25
AT3G60320 699 / 0.0
Protein of unknown function (DUF630 and DUF632) (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04782
DUF632
Protein of unknown function (DUF632)
Representative CDS sequence
>Lus10002864 pacid=23181310 polypeptide=Lus10002864 locus=Lus10002864.g ID=Lus10002864.BGIv1.0 annot-version=v1.0
ATGGAGAAGGAGGCAGAGATGACTTGGATTCCGAGGATTACGAAACTCATGCCACTGTTTTGGATAAGTTGTTGGCCTGGGAAAAGAAACTCTTTGATGA
AGTCAAGTCTCACTGAGCAGCAAGGAGAGTTGATGAAGCTGGAGTACAAAAGGAAGGTTGCTTTGCTGAACAAGCAGAAGAAACGTGGTGCTAGTGCTGA
CTCCTTGGAAAAGACTAAAGCGGCCGTGAGTCATTTGCACACTGGATATATTGTTGATATGCAATCTATGGACTCAACTGTTGCTGAAGTTAATCACATA
CGCGATAACCAGTTGTATCCTAAACTTGTTGCTCTTGTCCAAGGGATGTCGAAGATGTGGGGGAACATGTCGATCCACCACGAAAGCCAGCTAAAGATTG
TAATGGACCTTAAAACGCTTGATGTCGCCCATGCCATTAAAGAAACAACGAGGCACCACCACGAACAGACCAGCCAACTATGTGATGTTGTCGAGAAATG
GAATGCACAGTTCGATAAGCTTGTGACGTATCAAAAGCAATACATCAATTGTCTCAATAGCTGGTTAAAGCTAAACCTCATCCCCATAGAAAGCAGCTTG
AAAGAGAAGATATCATCCCCGCAACGAGTCGTGAATCCCCCAATCCAACCCCTTCTTCATGCATGGCACGACCATCTCGAAAGGCTACCAGACGAGCTTG
CCAAAACTGCAATCTCTTCTTTCGAGGCCGTGATCAGAACCATTAAAGTACACCAAGAAGAAGAGATGAAGCTAAAGGAGAGATGTGAAGAAACTAGAAA
GGAGTATGATTGA
AA sequence
>Lus10002864 pacid=23181310 polypeptide=Lus10002864 locus=Lus10002864.g ID=Lus10002864.BGIv1.0 annot-version=v1.0
MEKEAEMTWIPRITKLMPLFWISCWPGKRNSLMKSSLTEQQGELMKLEYKRKVALLNKQKKRGASADSLEKTKAAVSHLHTGYIVDMQSMDSTVAEVNHI
RDNQLYPKLVALVQGMSKMWGNMSIHHESQLKIVMDLKTLDVAHAIKETTRHHHEQTSQLCDVVEKWNAQFDKLVTYQKQYINCLNSWLKLNLIPIESSL
KEKISSPQRVVNPPIQPLLHAWHDHLERLPDELAKTAISSFEAVIRTIKVHQEEEMKLKERCEETRKEYD
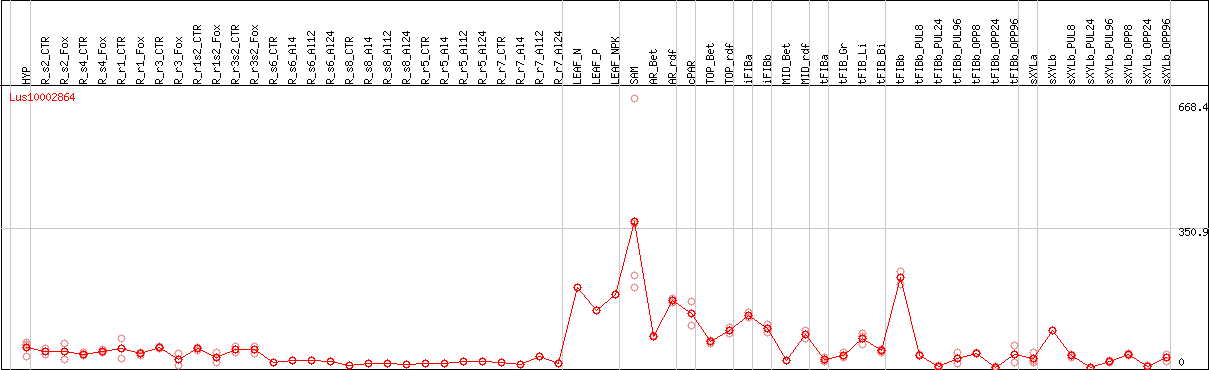
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10002864 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.