Lus10003130 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10003130 pacid=23169990 polypeptide=Lus10003130 locus=Lus10003130.g ID=Lus10003130.BGIv1.0 annot-version=v1.0
ATGCCTAGAAATGTTATCTCCGACGAAGAAGACGAGATGGAGTTCGATGACGAGGACAGGGAGCCGGAGCAAGGAGGTGCTGGCGATGATGACGAGGATG
AAGATGAAGAAGGGCAGGACGAGTATGAAAAGGATGATTTTATAGTGGACGATGTTGAAGAAGAGCCTGACGAAGAAGAAGAGAGGGCGGACAGTGATGA
GGAAAGGCAACGGAAGAAGAAGAAGAAGAAGAAACGAGAGGCTGAGGTTCTCGATGATGAAGACTACGAGCTTCTATTGGATAATAATGCCATACAACGA
AGACCACCAAAGGATAGCAAAAAGTTTAAAAGGCTTAAAAAGGCTCAACGGGATACAGATGGTGACGGGAATGAGTTCTCAGACGAGGAGTTTGATGGAA
GTGGTAAGGGTGGACGGACTGCTGAGGAAAAAATTAAGCGCAGCTTATTCGGTGATTATGAAGGAAACAATCTAGAGGATATTGCTGAAGAAGAGGAACA
AATAGAAGAGGAAGAAGATGTTGAAGTCGGGGAAGAGGATGAGATGGCTGATTTCATCGTTGATGAAGAAGAAATCGACGAGAATGGGCTTCCCGTTAGG
CGTAAGAAGATAAAGAGAAAGAAATTAAGGCAAGCATTTGGTGCTTCCTCTTCTGCTTTGCAGGAAGCTCATGAGATATTTGGAGATGTGGATGAGCTCT
TAGAGCTGCGTAAGCAACAAGGGAAAGATTCTGCTGAACAGCAGGAAAGGCGATTGGAAGATGAATTCGAGCCGACGGTACTTGCTGAGAAATACATGAC
TGAGAAAGATGATGTGATTCGACAAACCGACATTCCCGAAAGGATGCAGATATTAGAGGAGACTACTGGCCCGCCAACAGTTAATGCAGACAGTTTAGAC
GGTGAAAGCACGTGGATTTATAATCAGCTAGCAAGTGGTGTTGCGTCTATACAAGGTCTTGCTGCGTCTCTACAGGATCTGGCCAGCAATAAGGATGCGG
TCGAGCGGTTTAAGGATGAGATTAGCCGTTTTTTGGATTTTCACCACGTGCAAAAGTTGGATATCCCATTCATTGCCACATACCGGAAAGATGAGTGCTC
TAGCCTTTTCAAAGATCCTGAGCAACTCAAGTCCGATTACGACATTCACGATGACAATGACCGATCGCCAAAATTGAAGTGGCATAAGGTGGTTTCGGCA
ATTCAGGACTTGGATAGGAAGTGGCTACTTCTCCAAAAGAGGAAGACTGCTCTCCGCTTGTACTACCGCAAGCGATATGACGAAGAATCTATGCGGATTT
ATGACGAGACGAGACTTAATTTGAACCGGCACCTTTTCAAGTCCATTATTAAGTCGCTTGAGGCTGCCGAATCAGAACGAGAAGTTGATGATGTTGATGC
AAAGTTTAACTTGCACTTCCCGCCTGGTGAGGTGGGTGCCAATGAAGGACAATATAAGAGACCAAAAAGGAAATCTACATATAGTACCTGCAACAAGGCT
GGTCTATGGGAAGTTGCCAATAAGTTTGGTCGTAGTGCTGAGCAGCTCGGACGGCACTTGTCACTGTTGGAGATGGGTGTTGATTTGGAGGATGCCAAGG
AAGCACCTGAAGAGATGGCTTCGAACTTTACATGTGCAATGTTTGAAACTCCAGACGCTGTTCTTAAAGGTGCCAGGCACATGGCTGCCGTCGAGATAAG
TTGCGAGCCATACGTCAGGAAACATGTCCGTGCTGCTTACATGGAGAATGCAGTCGTATCCACCAGTCCAACTTCCGACGGAAACTCAACAATAGATTCA
TTCCACCAGTTTTACGGGGTGAAATGGCTGCGTGAAAAACCAGTCAGCGAGTTCAAGGACGGACAATGGCTTCTCATCCAAAAGGCCGAAGAGGAGAAGC
TTCTTCAAGTCACCTTCAAGCTTCCGGAAAAACGTATGGACGAATTAGTCACCGAATGCAAAGATCATTATCTTAGTGACGGTGTAAGTAAATCCGCCCA
GTTATGGAACGAGCAAAGGCAGCTAATCTTGCAAGACGCTCTCTTCGGGTTTCTTATGCCTTCTATGGAGAAAGAAGCGAGATCGTTGCTTACAAGTAGA
GCCAAACGGTGGCTACTATGGGAATACGGGCAAGTCTTGTGGAATAAAGTATCGGTCGGCCCGTATCAGAGGAAAGAAAATGACCGGTCTTCAGATGATG
AAGCTGCACCGAGGGTCATGGCTTGCTGTTGGGGCCCCGGAAAACCTGCAACGACGTTTGTTATGTTGGATTCGGCTGGGGAGGTGCTGGACGTGCTTTA
TACGGGTTCGCTTACGTTACGTTCTCAAAATGTTGATGACCAGCAAAGGAAGAAAAATGATCAGCAGCGAGTTTTGAAGTTTATGATGGATCATCAACCG
CATGTGGTCGCTCTAGGGGCTGTCAATTTGTCTTGTACAAAACTGAAGGAAGACATCTATGAGATCATTTTTAAGATGGTGGAAGAAAATCCCAGAGATG
TTGGACACGATATGGACGAATTAAGCATTTTGTACGGGGATGAGTCTCTTCCACGCCTTTACGAGAATTCCCGAATTTCGTCTGATCAACTACCTTCACA
ATCAGGGATCGTTAAAAAGGCCGTTGCTCTCGGACGCTATCTTCAAAACCCATTGGCAATGGCGGCAACTCTCTGTGGACCTGAAAGAGAGATCCTATCT
TGGAAGCTAAACCCTTTAGAGAGCTTTCTCACATCTGACGAAAAGTACGCAATGGTCGAACAGGTTATGGTGGACGTCACAAACCAGGTCGGGTTCGACG
TGAACCTGGCAATAGGTCACGAATGGTTATTCGCTCCGCTGCAGTTTATATCCGGTCTCGGACCGAGGAAAGCAGCTTCTCTGCAGAGATCCCTCGTAAG
AGCCGGTGGGATTTACACGAGGAAAGACTTTGTTACGATCCACGACCTAGGCAAAAAGGTGTTCGTCAATGCGGTCGGTTTTTTGCGAGTTCGACGGAGC
GGCCTGGCAGCTAGCAGCAGTCAGTTTATTGATTTGTTGGACGACACGAGGATTCACCCAGAATCGTACGGGCTCGCTCAGGATTTGGCCAAAGATATGT
ACGAACTCGATAATGTTGATGCAAATGATGATGACGATCTCGAAATGGCAATCGAGCATGTTCGGGACCACCCTTCGAGGTTGAGAAACCTGACTATTGA
TCAGTATCTTCTTGACAAGGGCATGCTGAATAAAACGGAGACGTTGAAGGATATAAGACGGGAATTGATACACGGTTTCCAGGATTGGCGGAAACCGTAT
AAAGAGCCGAGCCAGGAGGAAGAGTTTTACATGATTTCAGGCGAGACGGAAGATACAGTTGCAGAGGGTAGAATCGTCCAAGCCACTGTCCGGAGGGTAG
TGAATGGTTCGAAAGCGATATGTTCTCTTGAATCTGGATTGACGGGGATACTTATGAAGGAAGATTATGGTGATGACTCGAGGGATAATTCAGAGTTGTC
GGATAGGCTCCGTGAAGGTGATATACTCACGTGCAGGATAAAATCTGTTCAGATGAACAGATACCAGGCTTACATATCTTGTAAGGAGAGTGAGATGAGG
AATAATAACCGGTACCAGCCCGTTCGAGATCTCGACCCTTATTACCATGAAGATACAGTCAGCTTGCAGAGTGACCAGGAGAAGGCTCGTAAAGAGAAGG
ACCTTGCGAAGAAGCATTTCAAACCAAGGATGATCGTGCATCCTCAGTTCCAGAATATTACTTCGGATGAAGCTATGGAGTATTTGGCTGATATGGAGCC
TGGGGCAAGCATTATTCGTCCAAGTTCTCGTGGCCCCTCATTTCTGACTCTTACTCTGAAAGTCTACAATGGAGTTTATGCCCACAAAGACATAGTTGAA
GGGGGTAAAGAGCACAAGGACATCACGAGCCTTCTACGTATCGGGAAGACACTGAAAATCGGGGAGGACACTTTCGAGGATTTGGACGAGGTAATCGATA
GGTATGTCGATCCTCTTGTGGTCCACCTGAAAACTATGCTGAACTACCGAAAGTTCCGAAGGGGAACAAAGGCGGAAGTGGACGAGCTCCTGAAGATCGA
GAAAGCAGAGTATCCCATGAGGATAGTCTACTGCTTTGGCATATCTCATGAGCATCCGGGGACTTTCATTCTGACGTATATAAGGAGTACGAATCCACAT
CATGAGTACGTAGGCCTCTACCCGAAGGGATTCAAGTTCCGGAAGAAGATGTTTGAGGACATCGACAGGCTCGTCAATTACTTCCAGAAGCACATCGATG
ATCACCCGCAAGAATCAGGGCCGTCGATTAGATCGATGGCTGCCATGGTGCCGATGAGGAGCTCTACTCCCGGAGGAGGGTCTTCGGGAGATGGTGGTTG
GGGTGGTGGTGGGTCGAACAATGATGGTGGTGGATGGAGAGGACANNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNGGGGGGGGGGGGGCCGGCCCCGGCGGCGGTGGCAGTGGTTGGTGA
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10003130 pacid=23169990 polypeptide=Lus10003130 locus=Lus10003130.g ID=Lus10003130.BGIv1.0 annot-version=v1.0
MPRNVISDEEDEMEFDDEDREPEQGGAGDDDEDEDEEGQDEYEKDDFIVDDVEEEPDEEEERADSDEERQRKKKKKKKREAEVLDDEDYELLLDNNAIQR
RPPKDSKKFKRLKKAQRDTDGDGNEFSDEEFDGSGKGGRTAEEKIKRSLFGDYEGNNLEDIAEEEEQIEEEEDVEVGEEDEMADFIVDEEEIDENGLPVR
RKKIKRKKLRQAFGASSSALQEAHEIFGDVDELLELRKQQGKDSAEQQERRLEDEFEPTVLAEKYMTEKDDVIRQTDIPERMQILEETTGPPTVNADSLD
GESTWIYNQLASGVASIQGLAASLQDLASNKDAVERFKDEISRFLDFHHVQKLDIPFIATYRKDECSSLFKDPEQLKSDYDIHDDNDRSPKLKWHKVVSA
IQDLDRKWLLLQKRKTALRLYYRKRYDEESMRIYDETRLNLNRHLFKSIIKSLEAAESEREVDDVDAKFNLHFPPGEVGANEGQYKRPKRKSTYSTCNKA
GLWEVANKFGRSAEQLGRHLSLLEMGVDLEDAKEAPEEMASNFTCAMFETPDAVLKGARHMAAVEISCEPYVRKHVRAAYMENAVVSTSPTSDGNSTIDS
FHQFYGVKWLREKPVSEFKDGQWLLIQKAEEEKLLQVTFKLPEKRMDELVTECKDHYLSDGVSKSAQLWNEQRQLILQDALFGFLMPSMEKEARSLLTSR
AKRWLLWEYGQVLWNKVSVGPYQRKENDRSSDDEAAPRVMACCWGPGKPATTFVMLDSAGEVLDVLYTGSLTLRSQNVDDQQRKKNDQQRVLKFMMDHQP
HVVALGAVNLSCTKLKEDIYEIIFKMVEENPRDVGHDMDELSILYGDESLPRLYENSRISSDQLPSQSGIVKKAVALGRYLQNPLAMAATLCGPEREILS
WKLNPLESFLTSDEKYAMVEQVMVDVTNQVGFDVNLAIGHEWLFAPLQFISGLGPRKAASLQRSLVRAGGIYTRKDFVTIHDLGKKVFVNAVGFLRVRRS
GLAASSSQFIDLLDDTRIHPESYGLAQDLAKDMYELDNVDANDDDDLEMAIEHVRDHPSRLRNLTIDQYLLDKGMLNKTETLKDIRRELIHGFQDWRKPY
KEPSQEEEFYMISGETEDTVAEGRIVQATVRRVVNGSKAICSLESGLTGILMKEDYGDDSRDNSELSDRLREGDILTCRIKSVQMNRYQAYISCKESEMR
NNNRYQPVRDLDPYYHEDTVSLQSDQEKARKEKDLAKKHFKPRMIVHPQFQNITSDEAMEYLADMEPGASIIRPSSRGPSFLTLTLKVYNGVYAHKDIVE
GGKEHKDITSLLRIGKTLKIGEDTFEDLDEVIDRYVDPLVVHLKTMLNYRKFRRGTKAEVDELLKIEKAEYPMRIVYCFGISHEHPGTFILTYIRSTNPH
HEYVGLYPKGFKFRKKMFEDIDRLVNYFQKHIDDHPQESGPSIRSMAAMVPMRSSTPGGGSSGDGGWGGGGSNNDGGGWRGXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXGGGGAGPGGGGSGW
|
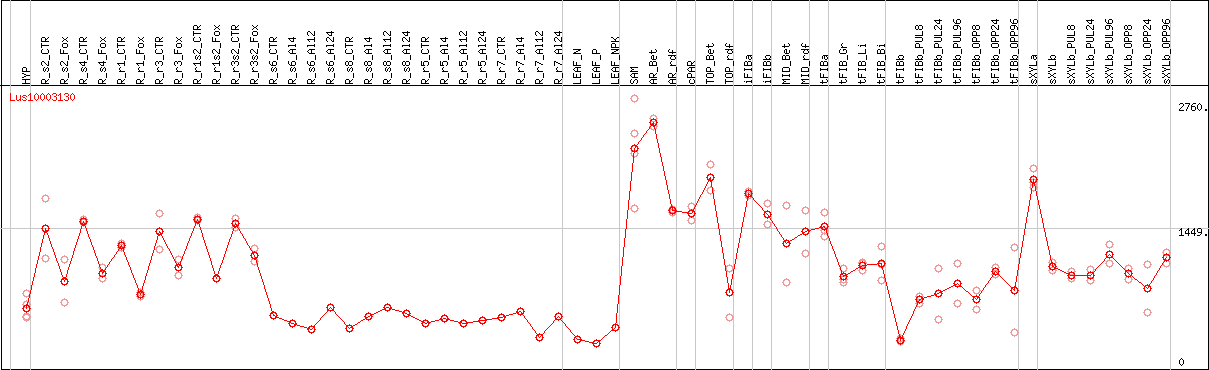
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10003130 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
