External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G61150 57 / 7e-11
HD
HD-GL2-1, HDG1
HOMEODOMAIN-GLABRA2 1, homeodomain GLABROUS 1 (.1)
AT4G00730 57 / 1e-10
HD
AHDP, ANL2
ANTHOCYANINLESS 2, ARABIDOPSIS THALIANA HOMEODOMAIN PROTEIN, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
AT5G52170 55 / 5e-10
HD
HDG7
homeodomain GLABROUS 7 (.1)
AT4G04890 49 / 8e-08
HD
PDF2
protodermal factor 2 (.1)
AT4G21750 48 / 1e-07
HD
ATML1
MERISTEM LAYER 1, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
AT1G05230 47 / 2e-07
HD
HDG2
homeodomain GLABROUS 2 (.1.2.3.4)
AT1G73360 47 / 3e-07
HD
ATHDG11, HDG11, EDT1
ENHANCED DROUGHT TOLERANCE 1, homeodomain GLABROUS 11 (.1)
AT1G17920 44 / 4e-06
HD
HDG12
homeodomain GLABROUS 12 (.1)
AT2G32370 38 / 0.0005
HD
HDG3
homeodomain GLABROUS 3 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005759
105 / 7e-28
AT3G61150 706 / 0.0
HOMEODOMAIN-GLABRA2 1, homeodomain GLABROUS 1 (.1)
Lus10027437
105 / 2e-27
AT3G61150 715 / 0.0
HOMEODOMAIN-GLABRA2 1, homeodomain GLABROUS 1 (.1)
Lus10005757
74 / 3e-17
AT4G00730 155 / 1e-43
ANTHOCYANINLESS 2, ARABIDOPSIS THALIANA HOMEODOMAIN PROTEIN, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Lus10005758
67 / 3e-14
AT3G61150 419 / 3e-137
HOMEODOMAIN-GLABRA2 1, homeodomain GLABROUS 1 (.1)
Lus10011633
66 / 6e-14
AT3G61150 160 / 2e-42
HOMEODOMAIN-GLABRA2 1, homeodomain GLABROUS 1 (.1)
Lus10036567
60 / 9e-12
AT4G00730 979 / 0.0
ANTHOCYANINLESS 2, ARABIDOPSIS THALIANA HOMEODOMAIN PROTEIN, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Lus10038877
56 / 2e-10
AT4G00730 261 / 6e-83
ANTHOCYANINLESS 2, ARABIDOPSIS THALIANA HOMEODOMAIN PROTEIN, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Lus10041362
56 / 2e-10
AT4G00730 379 / 2e-122
ANTHOCYANINLESS 2, ARABIDOPSIS THALIANA HOMEODOMAIN PROTEIN, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Lus10006765
52 / 5e-09
AT4G21750 1152 / 0.0
MERISTEM LAYER 1, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G141800
72 / 6e-16
AT4G00730 829 / 0.0
ANTHOCYANINLESS 2, ARABIDOPSIS THALIANA HOMEODOMAIN PROTEIN, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Potri.012G139300
67 / 4e-14
AT4G00730 838 / 0.0
ANTHOCYANINLESS 2, ARABIDOPSIS THALIANA HOMEODOMAIN PROTEIN, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Potri.014G075200
64 / 4e-13
AT4G00730 1045 / 0.0
ANTHOCYANINLESS 2, ARABIDOPSIS THALIANA HOMEODOMAIN PROTEIN, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Potri.002G154700
59 / 2e-11
AT4G00730 1057 / 0.0
ANTHOCYANINLESS 2, ARABIDOPSIS THALIANA HOMEODOMAIN PROTEIN, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Potri.011G025000
50 / 4e-08
AT4G21750 1164 / 0.0
MERISTEM LAYER 1, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Potri.004G020400
48 / 1e-07
AT4G04890 1185 / 0.0
protodermal factor 2 (.1)
Potri.014G152000
46 / 7e-07
AT1G05230 1179 / 0.0
homeodomain GLABROUS 2 (.1.2.3.4)
Potri.002G230200
46 / 7e-07
AT1G05230 1159 / 0.0
homeodomain GLABROUS 2 (.1.2.3.4)
Potri.012G038500
45 / 1e-06
AT1G73360 920 / 0.0
ENHANCED DROUGHT TOLERANCE 1, homeodomain GLABROUS 11 (.1)
Potri.001G184100
43 / 7e-06
AT1G79840 946 / 0.0
GLABRA 2, HD-ZIP IV family of homeobox-leucine zipper protein with lipid-binding START domain (.1.2)
PFAM info
Representative CDS sequence
>Lus10003300 pacid=23169871 polypeptide=Lus10003300 locus=Lus10003300.g ID=Lus10003300.BGIv1.0 annot-version=v1.0
ATGATGAGGCAGCGTTTAAACGATCCACTCTGTAACAATTGCGGTGGCAATGTGATATCTGCAATCGTCAAGCTTGAGCAACAACAACTACGAATCGAGA
ATGCTCGTCTCAAGAACAAAATCGATCGCGTATTGTCCTTTTCAACCAGCAAGTTGCTCGGCAAACCTGATATTTTGTCATCAGTCTCCTCCCCAACATA
TGCTTCCAATGGTTATGGACCAATCACAAATGTGATTGCACCACTAGTGGGGATTGATTTGGCTGATCCTTGCATGGATACCGGTGTCCGACCTATGATC
ATGAGACCAGCATAA
AA sequence
>Lus10003300 pacid=23169871 polypeptide=Lus10003300 locus=Lus10003300.g ID=Lus10003300.BGIv1.0 annot-version=v1.0
MMRQRLNDPLCNNCGGNVISAIVKLEQQQLRIENARLKNKIDRVLSFSTSKLLGKPDILSSVSSPTYASNGYGPITNVIAPLVGIDLADPCMDTGVRPMI
MRPA
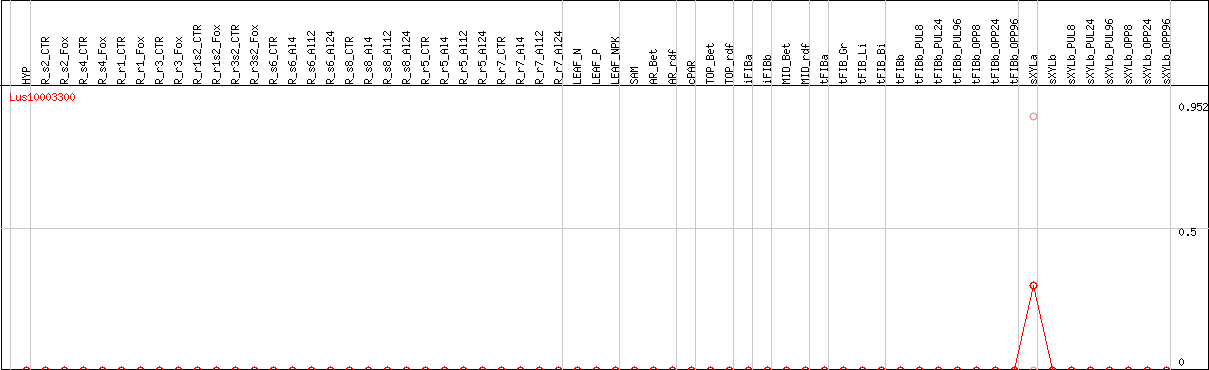
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10003300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.