Lus10003482 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10003482 pacid=23182491 polypeptide=Lus10003482 locus=Lus10003482.g ID=Lus10003482.BGIv1.0 annot-version=v1.0
ATGGCGCTGTTCAGACGCTTCTTCTATCGGAAGCCGCCCGATCGGCTTCTGGAGATCTCCGAGAGAGTCTACGTTTTCGACTGTTGCTTCTCCACGGATG
TCATGGAAGAAGATCAGTACAAGCTCTACTTAGGTGGCATTGTAGCACAGCTGCAGGATTACTTTCCAGATTCTTCCTTTATGGTGTTTAACTTTAGAGA
AGGAGATAGACGGAGCCAAATTTCTGACATCTTGACGCAGTATGACATGACAGTTATGGATTACCCTCGTCAACACGAAGGATGTCCATTGCTGCCTCTG
GAGATGGTTCATCATTTCCTACGATCCAGTGAAAGCTGGTTGTCATTGGAGGGGCAGCAAAATGTGCTGTTAATGCACTGCGAAAGAGGTGGATGGCCTG
TGCTTGCTTTCATGCTTGCTGGTCTTTTGTTATATAGGGGGCAGTATAGTGGAGAGCAGAAGACTCTTGAAACGGTTTATAAGCAAGCCCCTCGAGAGCG
TCAACATTTTTTGTCGGCTTTAAATCAGTACCCTTCCCAGCTTAGATATCTTCAGTATATTTCTAGAAGAAATCTTGGAGTTGATTGGCCTCCTTCCGAC
ACACCATTAATTCTCGACTGCTTGATGCTAAGATCCATTCCACTGTTCGAAGGAGGAAAGGGTTGTAGGCCAGTTGTGCGTGTTTATGGCCAGGATCCTT
CAAAACCCGCCAATAGAACGTCCAAGCTTTTGTATTCAACTACAAAGACTAAAAAACAAATACGGCAGTATGTACAGGAAGAGTGCATGCTTGTGAAAGT
AGATCTTAGGTGCCGTGTTCAAGGTGATGTTGTTATTGAATGTGTCCACGTAGATGAGGACCTCCTGCAAGAAGAGATGATCTTCAGACTGATGTTCCAC
ACAGCATTTGTGCGAGCAAATGTTTTGATGCTAAGTCGTGATGACATCGATATTATGTGGGATGCCAAAGATCAATTTCCACGGGAATTCAGGGCAGAGG
TACTTTTTGCTGATGCCGACACTATTCTACCGCCTCTCACCAGAGTTCTAACCGGTGAGGATCTGGATGATTCAGAAAGTGTCACACCTGAAGAATTTTA
TGAGGTGGAAGAGATATTCACAGTGGATGCACCGGAATCAAAAGGGAATTTTGATTTTAATAAAACTAAGAAGTCATCCCAATTAGAGTTTTCAGAAGTT
GTGAGAGAGGATGTGGATGTGCCCTCTTTCCAAGATTGTAGGGTCGATGATCTGAACCACAAGAAAAACAGGAAGCTTAATTCTAATGTGGATGCCGTGA
AGGATATTGCTGTAGATGATGTGAAGTACAGAATGGATGAGAAGGAGATTTCTGATCTTGAAGCAGTGAAAGACATTACCGTTGATAATGGGAATATGAA
GTTTGGTCCCAGAAATACAATTTCTGATGTTCAGATAAATGAGGAAACAGATGTAATCACTTATGAAGCAGCTGAAGAATTCAAAATGACGGCAGATAAA
GCTGATGGAGCAAGCAGTTCAAATTTGAAGTTAGAACCGAGTGTCTTATTACAGAAGCTGACTACTGAATTTGATAGAGAGAAACCGGAGAAACTGGAGC
AGCCTTCTCCCTTATGGCATTTTGTCGCTAACACAAAGCCTCCAGAATCGATGCTTGCTAGGCAAAAGAAACAGGGTTTTATTTTGGGCGAGAATTTAAG
ACAGATAAAGCCAGACACTCCACCACTGTGGGTGCTACATAACAAAGGTCCAGTAAGCAGTCTAATGAACGTATCACTTCCACCATCCAGGTATAATAGT
GCACCACCTGATTTTCCTCGTCTTCTTGAAGAGGCCACTGCTGGTGCCAATGAAAAGTCGTCCTCTACTGCTCCCACTATCCAAGTTTTGGATTCGAGTA
AGTTGATTCCTTCTTCAACTGATGATCAAGGACAGACAGCATCCATCCTTGCTTCTTCTGCAGCCTCAGGCCCGCTGCTACCACCACCACCTCCCTTGTC
TTCATTTTCCAGTACAGTTTCGACGACACCCTCATTCCGGAGTCCCATTCCACCTCCACCGCCACCTCCGGCTTTCATATCATCCAGCCAGGTGAAAGTT
GAAGTAGCATCTGAATGTGCAGTTCCAAAGAGCAGGTCACCACCACCACCACCACCTCCCCCTCTGCATTTCAATAATGGGCATAACAGTGGGATGACAT
TGCCCACTCCTCCCTCTCTATCGTGGAAGTCTGTTCCTGTAAGTACTTATATCCCCTCTACTACTAGTCCTACACCTCCTACACATCCTCTACCACTCCA
TTCTGGCAATGGGAGCACATCCAAAACATTAGAGCCTAGTATTGTAACCCCTACTACACCTTCTCTACCATCAGTACGTGGTACTCCTCTTATACCACCT
CCACCGCCACCACCATATCTTCCTTCAAGGAGTGCCACAGTGCCGCCCCCGCTCCCACCACCACCACCTCCTTATGTTTCTTCAACCTTGCCTCCACCAC
CTCCTTGTATTCATGAAAATGTTCATCCCCCACCTCCTCCTCCAATGCCAGGAGCACCTCCTCCACCGCCTCCTCCGAGAGGTGGGTTTGCACCTCCTCC
ACCGCCTCCTCCGGGAGGTGGGTTGGCACCTCCTCCGCCGCCTCCTCCGGGAGGTGGGTTTGCACCTCCTCCGCCGCCTCCTCCAGGAGGTGGGCGCGGA
CCTCCTCCACCGCCTCCACCAGGAGGTGGGCGCGGACCTCCTCCACCCCCTCCCCCCGGAGGACGTGTCCCAGGTCCCCCAGGTGGTGGACCGCCTCCAC
CACCACCTTTAGGCGCCAAAGGAGCTGATGCAAGAGGCGGACCAGGTAGAGGCCGAGGGCGTGGGGGTGCTGGTATGGCACCTCGAAGATCTTCTTTGAA
GCCTTTGCATTGGAGCAAGGTAACTAGGGCAATACAAGGAAGCTTGTGGGAAGAACTGCAGAGGCATGGGGAGACACAAATATTTCTCGCTAACATTCAT
GTATCGTCCTCTTTGATCGGTGACCTTGCATCACTTTGTAACAGTGCACCAGCGGAATTTGACGTGTCAGAGATAGAGACCCTTTTTTCTGCAATAGTTC
CAAAACCTGCTGACTCGAGTAGAACGAAAAAGAAGGCTGCTGGATCAAAACCTGACAAAGTTCAACTGATTGATCTGAGGAGGGCAAATAATACTGAAAT
TATGCTCACAAAAGTTAAAATGCCTCTTTCAGACATGATGGCTGCAGTACTTGCCATGGATGAGTCGGTTTTAGATGTTGATCAAGTGGACAACCTGATA
AAATTCTGTCCAACGAAAGAAGAGATGGAACAACTCAAGAACTACACCGGCGACAAGGAACTCTTGGGAAGGTGTGAACAGATCACAGAATTTAAGAGGA
GCTTAAACACGGTGAACTCTGCCTGTGATGAGGTTAGAAACTCCGTCAAACTGAAGGAAATCATGAAGAAGATTCTGTATTTGGGGAACACTTTGAATCA
AGGCACTGCAAGGGGTGCTGCAATTGGATTCAAGTTGGACAGTCTTCTGAAGCTTACTGATACACGTGCTGCTAACAGCAAGATGACACTTATGCATTAT
CTTTGTAAGGTACTTGCTGCAAAGTCGCCGCCGCTTTTGGATTTTCACCTCGATCTGGTCAGCATTGAAACTGCCACAAAGATACAATTGAAATCTCTAG
CTGAGGAAATGCAAGCTATCATCAAGGGATTGGAAAAGGTAAAACAGGAGAATGCAGCTTCAGCAAATGACGGCCCTGTATCTGACGTCTTCAGAAAGGC
GTTGAAAGAATTCATTTCCGGTGCTGAGGCAGAGGTGGCATCTGTGACCAACTTATACTCTGTGGTGGGTAGAAATGCAGATGCACTTGCGTTATACTTT
GGTGAGGATCCTGCTCGGTATCCTTTTGAGCAAGTTACTGCAACCCTCTTGAACTTTGTGAAGTTGTTCCGGAAAGCACATGACGAGAATTTGAAGCAGG
CTGAAGCGGATAAGAAGAAAGCCGAGAAGGACGCAGAAATGGAGAAGGCAAAGGCATCGAAGAAAAGCGATACGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10003482 pacid=23182491 polypeptide=Lus10003482 locus=Lus10003482.g ID=Lus10003482.BGIv1.0 annot-version=v1.0
MALFRRFFYRKPPDRLLEISERVYVFDCCFSTDVMEEDQYKLYLGGIVAQLQDYFPDSSFMVFNFREGDRRSQISDILTQYDMTVMDYPRQHEGCPLLPL
EMVHHFLRSSESWLSLEGQQNVLLMHCERGGWPVLAFMLAGLLLYRGQYSGEQKTLETVYKQAPRERQHFLSALNQYPSQLRYLQYISRRNLGVDWPPSD
TPLILDCLMLRSIPLFEGGKGCRPVVRVYGQDPSKPANRTSKLLYSTTKTKKQIRQYVQEECMLVKVDLRCRVQGDVVIECVHVDEDLLQEEMIFRLMFH
TAFVRANVLMLSRDDIDIMWDAKDQFPREFRAEVLFADADTILPPLTRVLTGEDLDDSESVTPEEFYEVEEIFTVDAPESKGNFDFNKTKKSSQLEFSEV
VREDVDVPSFQDCRVDDLNHKKNRKLNSNVDAVKDIAVDDVKYRMDEKEISDLEAVKDITVDNGNMKFGPRNTISDVQINEETDVITYEAAEEFKMTADK
ADGASSSNLKLEPSVLLQKLTTEFDREKPEKLEQPSPLWHFVANTKPPESMLARQKKQGFILGENLRQIKPDTPPLWVLHNKGPVSSLMNVSLPPSRYNS
APPDFPRLLEEATAGANEKSSSTAPTIQVLDSSKLIPSSTDDQGQTASILASSAASGPLLPPPPPLSSFSSTVSTTPSFRSPIPPPPPPPAFISSSQVKV
EVASECAVPKSRSPPPPPPPPLHFNNGHNSGMTLPTPPSLSWKSVPVSTYIPSTTSPTPPTHPLPLHSGNGSTSKTLEPSIVTPTTPSLPSVRGTPLIPP
PPPPPYLPSRSATVPPPLPPPPPPYVSSTLPPPPPCIHENVHPPPPPPMPGAPPPPPPPRGGFAPPPPPPPGGGLAPPPPPPPGGGFAPPPPPPPGGGRG
PPPPPPPGGGRGPPPPPPPGGRVPGPPGGGPPPPPPLGAKGADARGGPGRGRGRGGAGMAPRRSSLKPLHWSKVTRAIQGSLWEELQRHGETQIFLANIH
VSSSLIGDLASLCNSAPAEFDVSEIETLFSAIVPKPADSSRTKKKAAGSKPDKVQLIDLRRANNTEIMLTKVKMPLSDMMAAVLAMDESVLDVDQVDNLI
KFCPTKEEMEQLKNYTGDKELLGRCEQITEFKRSLNTVNSACDEVRNSVKLKEIMKKILYLGNTLNQGTARGAAIGFKLDSLLKLTDTRAANSKMTLMHY
LCKVLAAKSPPLLDFHLDLVSIETATKIQLKSLAEEMQAIIKGLEKVKQENAASANDGPVSDVFRKALKEFISGAEAEVASVTNLYSVVGRNADALALYF
GEDPARYPFEQVTATLLNFVKLFRKAHDENLKQAEADKKKAEKDAEMEKAKASKKSDT
|
DESeq2's median of ratios [FLAX]
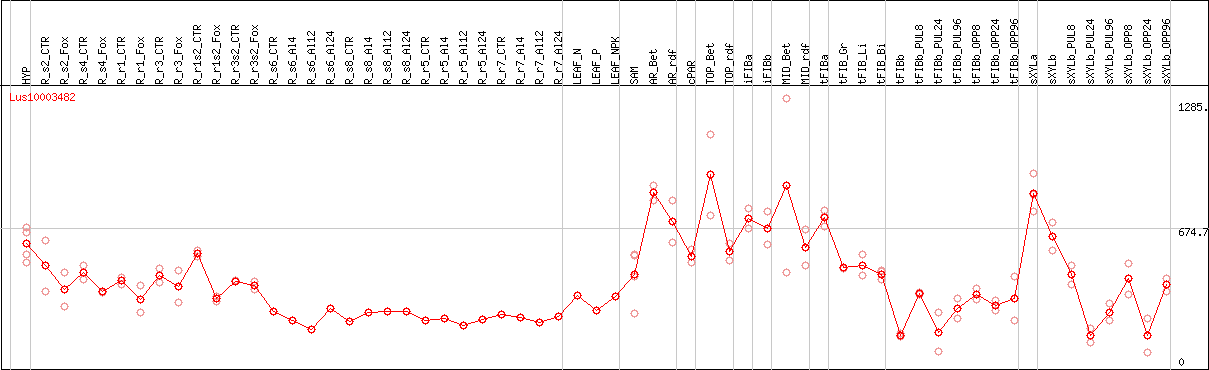
Coexpressed genes
Lus10003482 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
