External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G72180 270 / 2e-84
Leucine-rich receptor-like protein kinase family protein (.1)
AT4G28650 227 / 1e-68
Leucine-rich repeat transmembrane protein kinase family protein (.1)
AT5G49660 215 / 3e-64
XIP1
XYLEM INTERMIXED WITH PHLOEM 1, Leucine-rich repeat transmembrane protein kinase family protein (.1)
AT1G09970 214 / 4e-64
RLK7, LRRXI-23 ,LRR XI-23
receptor-like kinase 7, Leucine-rich receptor-like protein kinase family protein (.1.2)
AT1G08590 207 / 3e-61
Leucine-rich receptor-like protein kinase family protein (.1)
AT5G63930 206 / 8e-61
Leucine-rich repeat protein kinase family protein (.1)
AT3G19700 198 / 3e-58
IKU2
HAIKU2, Leucine-rich repeat protein kinase family protein (.1)
AT5G61480 197 / 5e-58
PXY, TDR
TDIF receptor, PHLOEM INTERCALATED WITH XYLEM, Leucine-rich repeat protein kinase family protein (.1)
AT2G33170 195 / 4e-57
Leucine-rich repeat receptor-like protein kinase family protein (.1)
AT1G17230 195 / 5e-57
Leucine-rich receptor-like protein kinase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10009476
467 / 2e-159
AT1G72180 995 / 0.0
Leucine-rich receptor-like protein kinase family protein (.1)
Lus10027951
224 / 2e-67
AT1G08590 1256 / 0.0
Leucine-rich receptor-like protein kinase family protein (.1)
Lus10028232
223 / 6e-67
AT1G09970 1181 / 0.0
receptor-like kinase 7, Leucine-rich receptor-like protein kinase family protein (.1.2)
Lus10000814
222 / 8e-67
AT1G08590 1263 / 0.0
Leucine-rich receptor-like protein kinase family protein (.1)
Lus10025943
218 / 2e-65
AT4G28650 1284 / 0.0
Leucine-rich repeat transmembrane protein kinase family protein (.1)
Lus10007232
218 / 4e-65
AT1G09970 1186 / 0.0
receptor-like kinase 7, Leucine-rich receptor-like protein kinase family protein (.1.2)
Lus10012462
201 / 3e-63
AT1G09970 391 / 2e-128
receptor-like kinase 7, Leucine-rich receptor-like protein kinase family protein (.1.2)
Lus10028234
209 / 3e-62
AT5G49660 1234 / 0.0
XYLEM INTERMIXED WITH PHLOEM 1, Leucine-rich repeat transmembrane protein kinase family protein (.1)
Lus10007233
209 / 5e-62
AT5G49660 1224 / 0.0
XYLEM INTERMIXED WITH PHLOEM 1, Leucine-rich repeat transmembrane protein kinase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G078400
306 / 8e-98
AT1G72180 1060 / 0.0
Leucine-rich receptor-like protein kinase family protein (.1)
Potri.019G021700
225 / 6e-68
AT1G08590 1286 / 0.0
Leucine-rich receptor-like protein kinase family protein (.1)
Potri.002G256500
222 / 7e-67
AT4G28650 1339 / 0.0
Leucine-rich repeat transmembrane protein kinase family protein (.1)
Potri.013G048800
222 / 1e-66
AT1G08590 1306 / 0.0
Leucine-rich receptor-like protein kinase family protein (.1)
Potri.002G106800
219 / 1e-65
AT1G09970 1204 / 0.0
receptor-like kinase 7, Leucine-rich receptor-like protein kinase family protein (.1.2)
Potri.009G081800
214 / 7e-64
AT1G09970 1033 / 0.0
receptor-like kinase 7, Leucine-rich receptor-like protein kinase family protein (.1.2)
Potri.003G107600
213 / 1e-63
AT5G61480 1259 / 0.0
TDIF receptor, PHLOEM INTERCALATED WITH XYLEM, Leucine-rich repeat protein kinase family protein (.1)
Potri.002G111700
212 / 5e-63
AT5G49660 1229 / 0.0
XYLEM INTERMIXED WITH PHLOEM 1, Leucine-rich repeat transmembrane protein kinase family protein (.1)
Potri.001G126100
210 / 2e-62
AT5G61480 1258 / 0.0
TDIF receptor, PHLOEM INTERCALATED WITH XYLEM, Leucine-rich repeat protein kinase family protein (.1)
Potri.004G049100
204 / 2e-60
AT1G28440 1387 / 0.0
HAESA-like 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0016
PKinase
PF00069
Pkinase
Protein kinase domain
Representative CDS sequence
>Lus10003503 pacid=23157266 polypeptide=Lus10003503 locus=Lus10003503.g ID=Lus10003503.BGIv1.0 annot-version=v1.0
ATGCCGAAAGGGAATTTATTCGAAGCGATTCACAGGAAGATCGATAATAAAGGCGGCGAATTGCCTGAATTGGATTGGATTCATAGGTACAGAATTGCAG
TAGGAGCTGCTAAGGGGATTTCGTATCTTCACCACGATTGCTCTCCGCCTGTCATCCATCGAGACATCAAGTCGAGTAATGTGCTGCTCGACGATGATTA
CGAGGCGAAGATTGCTGATTTCGGAGTTGCGAGGAAGCTTGCTGAGGTTTCTGAACAAGGGTGTGATCACAGCTGCTTTGCTGGTACGCATGGATACATT
GCTCCTGAGATGGCGTATTCGATCAAGGTCACAGAGAAGAGCGACGTGTACAGCTTCGGAGTGGTGCTGCTCGAGCTCATAACGGGGAAGAAACCGGTCG
AGGAAGAGTACGGAGAAGGGAAAGACATAGTGTCGTGGGTTTCCAAGCATTTGAGGGACATTGACAGTGTTCTTAACAGGGTTCTGGACGGTTCTATCGC
GAGTGATGAAACTGTGAAGGATGACATGCTGAAGGTATTGAAAATCGGGCTTCTTTGCACCACGAAGCTTCCCAGTCTCCGGCCGACGATGCGAGATGTG
GTGAGGATGCTCGTCGACGCCGAGCCTTCCTGTTCGGCCGAGCCGTTCGACGATTCTTTGAATGTGGATGGTTCTGGGAAAGAAAATCAAAAGCCTTTCC
TGGAATGA
AA sequence
>Lus10003503 pacid=23157266 polypeptide=Lus10003503 locus=Lus10003503.g ID=Lus10003503.BGIv1.0 annot-version=v1.0
MPKGNLFEAIHRKIDNKGGELPELDWIHRYRIAVGAAKGISYLHHDCSPPVIHRDIKSSNVLLDDDYEAKIADFGVARKLAEVSEQGCDHSCFAGTHGYI
APEMAYSIKVTEKSDVYSFGVVLLELITGKKPVEEEYGEGKDIVSWVSKHLRDIDSVLNRVLDGSIASDETVKDDMLKVLKIGLLCTTKLPSLRPTMRDV
VRMLVDAEPSCSAEPFDDSLNVDGSGKENQKPFLE
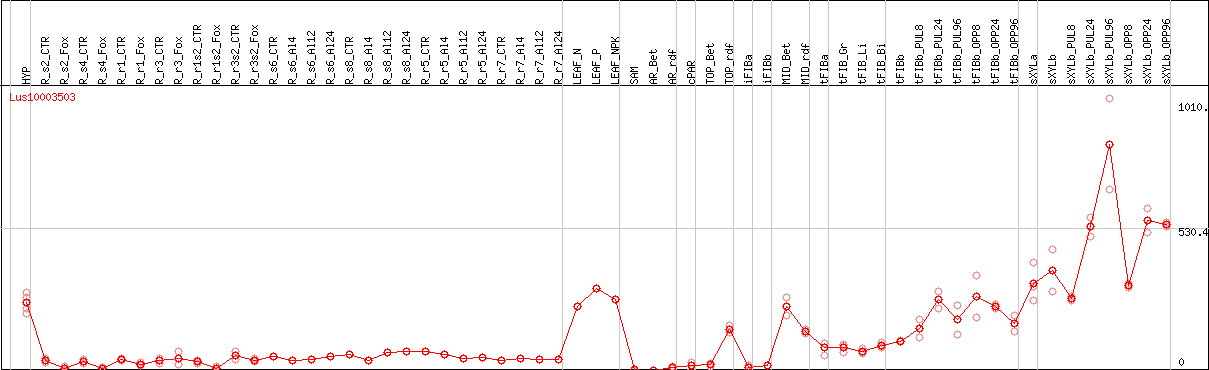
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10003503 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.