External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G25300 197 / 5e-61
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
AT1G17020 197 / 1e-60
ATSRG1, SRG1
senescence-related gene 1 (.1)
AT1G17010 194 / 9e-60
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G78550 186 / 1e-56
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G25310 183 / 2e-55
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT3G21420 129 / 1e-34
LBO1
LATERAL BRANCHING OXIDOREDUCTASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G49390 111 / 3e-28
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT5G20400 109 / 2e-27
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT2G38240 107 / 8e-27
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT3G11180 104 / 3e-25
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10022292
285 / 7e-95
AT1G17020 449 / 5e-159
senescence-related gene 1 (.1)
Lus10032574
227 / 5e-72
AT4G25300 419 / 2e-146
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Lus10026173
187 / 8e-57
AT1G17020 443 / 5e-156
senescence-related gene 1 (.1)
Lus10011985
176 / 2e-52
AT1G17020 367 / 4e-126
senescence-related gene 1 (.1)
Lus10011980
173 / 1e-51
AT1G17020 382 / 1e-132
senescence-related gene 1 (.1)
Lus10011979
173 / 2e-51
AT1G17020 374 / 4e-129
senescence-related gene 1 (.1)
Lus10011981
172 / 3e-51
AT1G17020 375 / 2e-129
senescence-related gene 1 (.1)
Lus10015645
160 / 2e-48
AT5G20630 98 / 4e-33
GERMIN-LIKE PROTEIN 3, ARABIDOPSIS THALIANA GERMIN 3, germin 3 (.1)
Lus10040112
163 / 2e-47
AT1G17020 332 / 1e-112
senescence-related gene 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G381700
226 / 6e-72
AT1G17020 436 / 2e-153
senescence-related gene 1 (.1)
Potri.001G382400
215 / 7e-68
AT1G17020 446 / 1e-157
senescence-related gene 1 (.1)
Potri.001G355100
188 / 3e-57
AT1G17020 439 / 1e-154
senescence-related gene 1 (.1)
Potri.009G025900
181 / 2e-54
AT4G25300 410 / 3e-143
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Potri.009G022800
144 / 1e-40
AT1G17020 302 / 6e-101
senescence-related gene 1 (.1)
Potri.001G355200
137 / 7e-38
AT1G17020 329 / 3e-111
senescence-related gene 1 (.1)
Potri.010G023600
129 / 1e-34
AT3G21420 511 / 0.0
LATERAL BRANCHING OXIDOREDUCTASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.010G200900
124 / 5e-33
AT3G21420 288 / 4e-95
LATERAL BRANCHING OXIDOREDUCTASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.010G201000
117 / 2e-30
AT3G21420 290 / 5e-96
LATERAL BRANCHING OXIDOREDUCTASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.006G062500
108 / 6e-27
AT5G20400 432 / 2e-152
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10003652 pacid=23171428 polypeptide=Lus10003652 locus=Lus10003652.g ID=Lus10003652.BGIv1.0 annot-version=v1.0
ATGCTGCTACTCCGCACCCGCCGCGGATTTCTGCGTAGCCGATCTATCTCTCCAAACCGCATCAGGCTACACGTGCAAGGACCCCACCAAAGCCACCGCC
GCCGACTTCCTCTTTACCGGAATATCCCAACCGGGCAACACCACCAACCTCTTCAGGGAAGTTGCTGCCCTGGCATCCGTCCGGCAGTTCCCGGCCCTGA
ACGGGCTGGGAATCTCCATGGCCCGCATCGTTTCGCTCTCGGCGGGGTGATCAAGCTGCACACCCACCCGGTGGCTACCGAGATAGCCTACCTGATTGAG
GGGTCTGTCACGGTGGGATTCGTCACGGCCGGGACGAACAAGGTGTACCTGAACAAGATCAATAAAAAGCTGCTCGATGAATCCTCGATGGATTCCGAGA
TTCAGAAGCTCCACTCTGCCTGTAAAGATTGGGGCTTTTTCCAGTTGGTGAATCGCGGAGTGAGCTCTGCTCTGGTAGAGGACATGAAAACACAGGTTCA
GGAATTCTTCAACCTTCCAATGGAAGAAAAGAAGAAGTTATGGCAATATCCAGGCGAAGTGGAAGGGTTCGGTCAATCATTCGTTGTGTCCGAGGAGCAG
AAACTTGATTGGGGAGACATGCTTTCCTTCTCACTCACCCTCTCCACCACAGAAAGCCTCACTTGCTCCCAAACCTTCCACTTCCTTTCAGGCAAGGTTC
TCGACCAAATGTCCAAAGCATTAAACATGAAGCCAGAGGAAATGAGGGACATGTTCTCAGGGGACATAAGACAGACAACGAGGATGAACTACTATCCTCC
GTCCCCTCAGCCGAACAAAGTCATCGGCTTGACCCCTCATTCCGACGGAACAGGGATAACAATCCTGTTACAAGTCAACGAAGTCGGAGGCTTGTAG
AA sequence
>Lus10003652 pacid=23171428 polypeptide=Lus10003652 locus=Lus10003652.g ID=Lus10003652.BGIv1.0 annot-version=v1.0
MLLLRTRRGFLRSRSISPNRIRLHVQGPHQSHRRRLPLYRNIPTGQHHQPLQGSCCPGIRPAVPGPERAGNLHGPHRFALGGVIKLHTHPVATEIAYLIE
GSVTVGFVTAGTNKVYLNKINKKLLDESSMDSEIQKLHSACKDWGFFQLVNRGVSSALVEDMKTQVQEFFNLPMEEKKKLWQYPGEVEGFGQSFVVSEEQ
KLDWGDMLSFSLTLSTTESLTCSQTFHFLSGKVLDQMSKALNMKPEEMRDMFSGDIRQTTRMNYYPPSPQPNKVIGLTPHSDGTGITILLQVNEVGGL
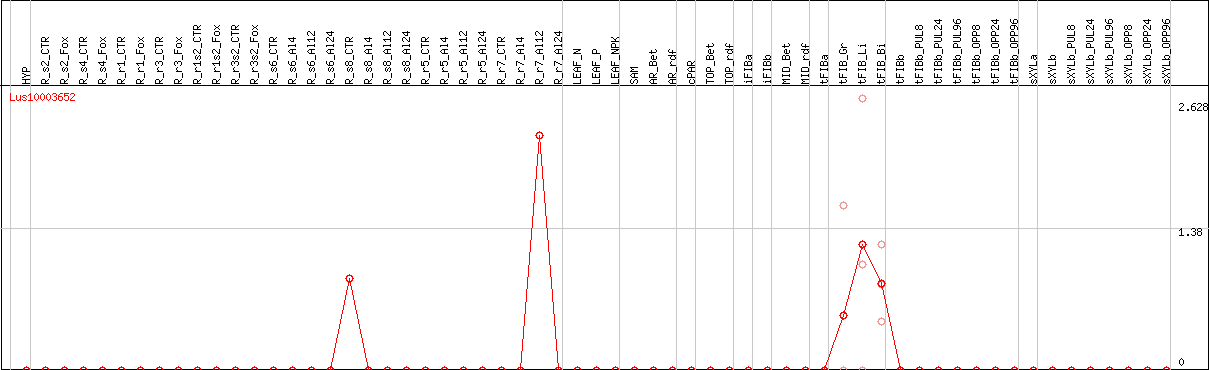
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10003652 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.