External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G34520 106 / 2e-28
MBOAT (membrane bound O-acyl transferase) family protein (.1)
AT1G34490 102 / 6e-27
MBOAT (membrane bound O-acyl transferase) family protein (.1)
AT5G55350 102 / 1e-26
MBOAT (membrane bound O-acyl transferase) family protein (.1)
AT1G34500 101 / 2e-26
MBOAT (membrane bound O-acyl transferase) family protein (.1)
AT3G51970 101 / 2e-26
ATASAT1, ASAT1, ATSAT1
ARABIDOPSIS THALIANA STEROL O-ACYLTRANSFERASE 1, acyl-CoA sterol acyl transferase 1 (.1)
AT5G55380 101 / 3e-26
MBOAT (membrane bound O-acyl transferase) family protein (.1)
AT5G55340 100 / 6e-26
MBOAT (membrane bound O-acyl transferase) family protein (.1)
AT5G55370 99 / 2e-25
MBOAT (membrane bound O-acyl transferase) family protein (.1)
AT5G55330 99 / 2e-25
MBOAT (membrane bound O-acyl transferase) family protein (.1)
AT5G55360 99 / 3e-25
MBOAT (membrane bound O-acyl transferase) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10022293
176 / 4e-57
AT5G55340 178 / 2e-55
MBOAT (membrane bound O-acyl transferase) family protein (.1)
Lus10039576
112 / 3e-30
AT3G51970 329 / 7e-112
ARABIDOPSIS THALIANA STEROL O-ACYLTRANSFERASE 1, acyl-CoA sterol acyl transferase 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.009G055100
128 / 2e-36
AT3G51970 330 / 8e-112
ARABIDOPSIS THALIANA STEROL O-ACYLTRANSFERASE 1, acyl-CoA sterol acyl transferase 1 (.1)
Potri.009G054900
125 / 3e-35
AT3G51970 280 / 1e-92
ARABIDOPSIS THALIANA STEROL O-ACYLTRANSFERASE 1, acyl-CoA sterol acyl transferase 1 (.1)
Potri.016G014700
124 / 1e-34
AT5G55340 242 / 1e-77
MBOAT (membrane bound O-acyl transferase) family protein (.1)
Potri.001G106800
118 / 2e-32
AT3G51970 255 / 1e-82
ARABIDOPSIS THALIANA STEROL O-ACYLTRANSFERASE 1, acyl-CoA sterol acyl transferase 1 (.1)
Potri.016G014600
118 / 2e-32
AT3G51970 299 / 1e-99
ARABIDOPSIS THALIANA STEROL O-ACYLTRANSFERASE 1, acyl-CoA sterol acyl transferase 1 (.1)
Potri.006G009800
114 / 4e-31
AT3G51970 244 / 3e-78
ARABIDOPSIS THALIANA STEROL O-ACYLTRANSFERASE 1, acyl-CoA sterol acyl transferase 1 (.1)
Potri.008G145700
109 / 3e-29
AT5G55340 264 / 1e-86
MBOAT (membrane bound O-acyl transferase) family protein (.1)
Potri.009G055000
107 / 1e-28
AT3G51970 314 / 7e-106
ARABIDOPSIS THALIANA STEROL O-ACYLTRANSFERASE 1, acyl-CoA sterol acyl transferase 1 (.1)
Potri.001G260700
103 / 5e-27
AT3G51970 268 / 2e-87
ARABIDOPSIS THALIANA STEROL O-ACYLTRANSFERASE 1, acyl-CoA sterol acyl transferase 1 (.1)
Potri.006G009600
89 / 6e-22
AT5G55360 145 / 3e-41
MBOAT (membrane bound O-acyl transferase) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0517
MBOAT-like
PF03062
MBOAT
MBOAT, membrane-bound O-acyltransferase family
Representative CDS sequence
>Lus10003655 pacid=23171432 polypeptide=Lus10003655 locus=Lus10003655.g ID=Lus10003655.BGIv1.0 annot-version=v1.0
ATGCATCCCCACTTCGTCTTCGCCCTCTATTGCCTACACTTGTATCTCGAATTAGAACTCGTTCTCGCTCTGTGCACGGTCCCGGCTCGAGCTCTGTTTG
GGTTCGAGCTGAAGCCTCAGTTCGACGAGCTGTATCTCTCCACGTCACTTCAGGACTTTTCGGGGCGGAGGTGGAATCTGATGCTCACCAAGATATTGCG
TCCCACCGTCTACGTTCCGATCAGAGGAGTGGCGGGGCGTGTAGTGGGTGGGCAGTGGGCTTCGCTGTCGGCGGTGGTGGCCACGTTCGTGGTGTCGCGG
TTAATGCACGAATTGATGTATTACTATATGAATAGGGTGGGGCCCACATGGGAGGTGGCGTGGTTTTTCGTGGAGTTTGCGTGGCTATGGAAGTGGGGGT
TAAGAAGAAATTTAAAGAGAAAGCATTTTGGTTGCCGTCGGTGA
AA sequence
>Lus10003655 pacid=23171432 polypeptide=Lus10003655 locus=Lus10003655.g ID=Lus10003655.BGIv1.0 annot-version=v1.0
MHPHFVFALYCLHLYLELELVLALCTVPARALFGFELKPQFDELYLSTSLQDFSGRRWNLMLTKILRPTVYVPIRGVAGRVVGGQWASLSAVVATFVVSR
LMHELMYYYMNRVGPTWEVAWFFVEFAWLWKWGLRRNLKRKHFGCRR
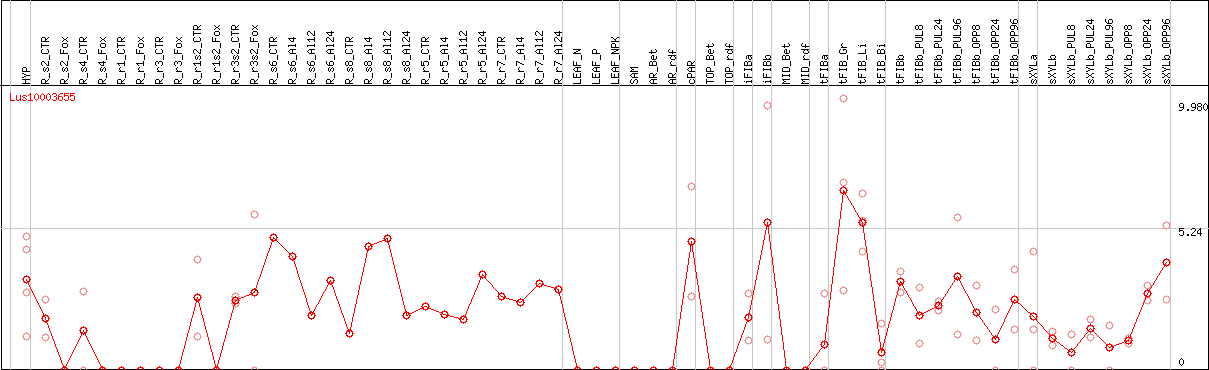
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10003655 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.