External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G33970 196 / 3e-60
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
AT1G33930 154 / 3e-44
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT1G33960 154 / 5e-44
AIG1
AVRRPT2-INDUCED GENE 1, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT4G09940 155 / 6e-44
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT1G33890 151 / 6e-43
Avirulence induced gene (AIG1) family protein (.1)
AT1G33950 149 / 2e-42
Avirulence induced gene (AIG1) family protein (.1), Avirulence induced gene (AIG1) family protein (.2)
AT1G33900 147 / 2e-41
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT2G26820 149 / 3e-41
ATPP2-A3
phloem protein 2-A3 (.1)
AT4G09930 141 / 4e-39
Avirulence induced gene (AIG1) family protein (.1)
AT4G09950 134 / 1e-36
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005300
260 / 4e-84
AT1G33970 321 / 7e-109
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10028023
236 / 1e-76
AT4G09940 181 / 2e-55
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10005299
233 / 1e-73
AT1G33970 339 / 5e-115
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10009484
182 / 5e-54
AT4G26220 255 / 1e-83
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10009485
164 / 6e-50
AT1G33970 142 / 2e-42
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10026891
149 / 3e-43
AT2G26820 156 / 2e-45
phloem protein 2-A3 (.1)
Lus10009482
100 / 2e-24
AT1G33970 148 / 9e-43
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10009483
94 / 2e-23
AT1G33970 91 / 3e-23
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10005301
68 / 8e-14
AT1G33970 121 / 3e-34
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G077300
249 / 9e-81
AT1G33970 345 / 2e-118
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Potri.013G104800
241 / 2e-77
AT1G33970 314 / 3e-106
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Potri.010G014300
43 / 0.0003
AT4G02510 812 / 0.0
TRANSLOCON AT THE OUTER ENVELOPE MEMBRANE OF CHLOROPLASTS 86, TRANSLOCON AT THE OUTER ENVELOPE MEMBRANE OF CHLOROPLASTS 160, PLASTID PROTEIN IMPORT 2, translocon at the outer envelope membrane of chloroplasts 159 (.1)
Potri.004G171600
43 / 0.0003
AT3G16620 1128 / 0.0
ARABIDOPSIS THALIANA TRANSLOCON OUTER COMPLEX PROTEIN 120, translocon outer complex protein 120 (.1)
Potri.009G131200
43 / 0.0003
AT3G16620 1129 / 0.0
ARABIDOPSIS THALIANA TRANSLOCON OUTER COMPLEX PROTEIN 120, translocon outer complex protein 120 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF04548
AIG1
AIG1 family
Representative CDS sequence
>Lus10003732 pacid=23169620 polypeptide=Lus10003732 locus=Lus10003732.g ID=Lus10003732.BGIv1.0 annot-version=v1.0
ATGGGAGGATGCTCGATTGAAAACGACTGGGAGTTCACCGAGCCATCTAGTGGGACAGCTCGAACAATTGTACTAGTTGGTCGTACTGGTAATGGCAAGA
GTGCGACCGGCAATACCATCCTGTGCAGAAAAGCTTTCAAGTCCAGGAGTAGCTCTCTTGGCGTTACGAGTAGCTGCGAATTGCAGTCAACTGTTTTGAA
CGATGGTCAGATCGTAAATGTCATTGATACTCCAGGCCTATTTGATCCTACAATGGATTCTGATTTCAAGAAGAGAACAGCAGTTCTTAGCCTGCAATCA
TTGTTCGGAACCAACATAACCGACTACATGATTGTTCTTTTCGCCGGAGGAGACGATCTTGAAGACAATGAACAAAGTTTAAACCATTATCTGTGTAATG
ATTCTGAAGTCTGCTCTCCTTTAAAGGAAGTTTTGTCGCTTTGCGACAATCGAATGATCTTGTTCAACAATAAGATCAAATATGAACTTGAGAGAGTTAA
ACAGGTTACCCAGCTTATGAATCTTGTAAGCAGCCTCGTAGCGCAGAATGGTGGGAAACCCTTCACTAGTGACATCTTTCAGATGGTGCAGAAACAGTCT
GAGGTAGATTCCTTCAACATAAAGAAGGCCGAATCTAGTCTGAATGAAGAACAGTTTAACCGCCTCAATGAAATGGTAACTTCTTCCTCTGTAACTAACC
TTCATCTTTTTTCATGTCTGCTGACTAATGCCGTCTCGTTAGATTTCGGAAAAGCTAAGGGAGACAACAGAGAGATGGAGGAGCAGGTGAGAGCGGAGCA
AGCTGCACGTTTGAAGGCTGAGAAGGCTGCAAAAGAGGCTCAAGAGAAATTAGCAAGATCAAAGAAGGATCGTAAAAAGCAGGCTATGAAGGTAAAGGAG
CTATGCGAGCAGCGTGCTAGGGACGCTGAGGAGGTTCGCAAGCAGCGTGCTAGGGTGGCCGAGGAGCGTCACAGGCTGCTTGCTGAGAAAAATTCTTGTT
CAATTCTCTAG
AA sequence
>Lus10003732 pacid=23169620 polypeptide=Lus10003732 locus=Lus10003732.g ID=Lus10003732.BGIv1.0 annot-version=v1.0
MGGCSIENDWEFTEPSSGTARTIVLVGRTGNGKSATGNTILCRKAFKSRSSSLGVTSSCELQSTVLNDGQIVNVIDTPGLFDPTMDSDFKKRTAVLSLQS
LFGTNITDYMIVLFAGGDDLEDNEQSLNHYLCNDSEVCSPLKEVLSLCDNRMILFNNKIKYELERVKQVTQLMNLVSSLVAQNGGKPFTSDIFQMVQKQS
EVDSFNIKKAESSLNEEQFNRLNEMVTSSSVTNLHLFSCLLTNAVSLDFGKAKGDNREMEEQVRAEQAARLKAEKAAKEAQEKLARSKKDRKKQAMKVKE
LCEQRARDAEEVRKQRARVAEERHRLLAEKNSCSIL
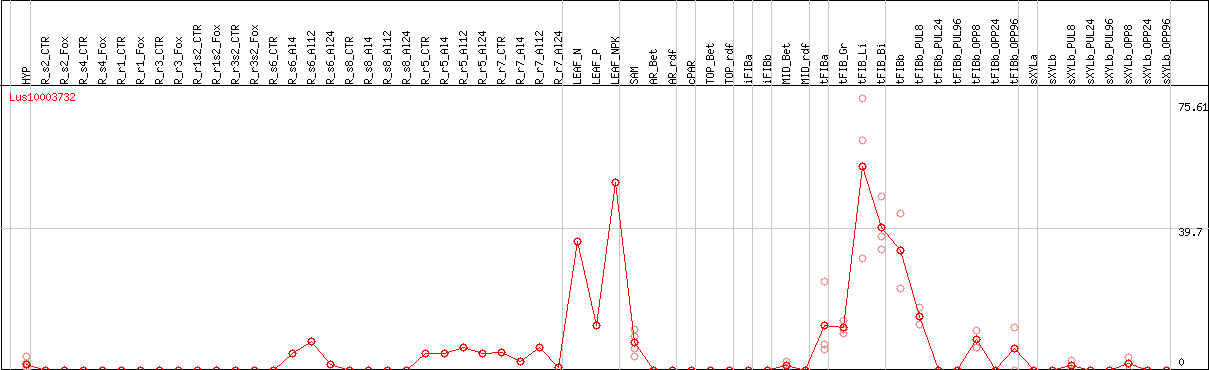
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10003732 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.