External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G54690 240 / 5e-83
HTA3 ,G-H2AX ,GAMMA-H2AX ,H2AXB
histone H2A 3, GAMMA H2AX, gamma histone variant H2AX (.1)
AT1G08880 240 / 6e-83
HTA5 ,G-H2AX ,GAMMA-H2AX ,H2AXA
histone H2A 5, gamma histone variant H2AX, GAMMA H2AX, Histone superfamily protein (.1)
AT5G54640 199 / 7e-67
ATHTA1, HTA1, RAT5
RESISTANT TO AGROBACTERIUM TRANSFORMATION 5, histone H2A 1, Histone superfamily protein (.1)
AT1G51060 198 / 1e-66
HTA10
histone H2A 10 (.1)
AT4G27230 197 / 2e-66
HTA2
histone H2A 2 (.1.2)
AT3G20670 196 / 7e-66
HTA13
histone H2A 13 (.1)
AT5G59870 184 / 8e-61
HTA6
histone H2A 6 (.1)
AT5G02560 177 / 4e-58
HTA12
histone H2A 12 (.1.2)
AT5G27670 174 / 1e-56
HTA7
histone H2A 7 (.1)
AT2G38810 126 / 4e-38
HTA8
histone H2A 8 (.1.2.3)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G028900
234 / 1e-80
AT1G54690 220 / 4e-75
histone H2A 3, GAMMA H2AX, gamma histone variant H2AX (.1)
Potri.005G040800
233 / 2e-80
AT1G54690 221 / 2e-75
histone H2A 3, GAMMA H2AX, gamma histone variant H2AX (.1)
Potri.005G040700
227 / 7e-78
AT1G08880 183 / 2e-60
histone H2A 5, gamma histone variant H2AX, GAMMA H2AX, Histone superfamily protein (.1)
Potri.013G028800
213 / 3e-72
AT1G08880 190 / 2e-63
histone H2A 5, gamma histone variant H2AX, GAMMA H2AX, Histone superfamily protein (.1)
Potri.004G031300
196 / 5e-66
AT1G08880 150 / 6e-48
histone H2A 5, gamma histone variant H2AX, GAMMA H2AX, Histone superfamily protein (.1)
Potri.011G131400
196 / 8e-66
AT1G51060 200 / 1e-67
histone H2A 10 (.1)
Potri.001G415700
196 / 2e-65
AT1G51060 174 / 1e-56
histone H2A 10 (.1)
Potri.006G082300
178 / 2e-58
AT5G02560 143 / 2e-44
histone H2A 12 (.1.2)
Potri.005G026500
175 / 3e-57
AT5G27670 145 / 2e-45
histone H2A 7 (.1)
Potri.013G018200
172 / 2e-56
AT5G27670 160 / 4e-51
histone H2A 7 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0012
Histone
PF00125
Histone
Core histone H2A/H2B/H3/H4
Representative CDS sequence
>Lus10003750 pacid=23169638 polypeptide=Lus10003750 locus=Lus10003750.g ID=Lus10003750.BGIv1.0 annot-version=v1.0
ATGAGTTCAGTCGCCGCCGCTGGTTTAGCCAAGTCCGGCAGAGGCAAGCCGAAATCCAAGGCTGTCTCTAGGTCTCAGAAAGCCGGTCTACAGTTCCCCG
TCGGCCGCATAGCTCGTTTCCTAAAGAGAGGTAGATACGCTGAGCGCGTTGGATCTGGGTCCCCTGTCTACCTCTCCGCCGTCATGGAGTATCTCGCCGC
TGAGGTTTTGGAGCTGGCTGGAAACGCTGCGAGGGATAACAAGAAGACAAGGATCGTTCCGAGGCACATTCAGCTGGCTGTGAGGAACGACGAGGAGTTG
AGCAAGCTGCTCGGAGGTGTCACCATTGCTAACGGAGGCGTGTTGCCGAAGATTCACCAGAGTTTGCTGCCGAAGAAAGCTGGGAAAGATAAGGGAGAGA
TCGGCTCTGCTTCCCAAGAGTTTTGA
AA sequence
>Lus10003750 pacid=23169638 polypeptide=Lus10003750 locus=Lus10003750.g ID=Lus10003750.BGIv1.0 annot-version=v1.0
MSSVAAAGLAKSGRGKPKSKAVSRSQKAGLQFPVGRIARFLKRGRYAERVGSGSPVYLSAVMEYLAAEVLELAGNAARDNKKTRIVPRHIQLAVRNDEEL
SKLLGGVTIANGGVLPKIHQSLLPKKAGKDKGEIGSASQEF
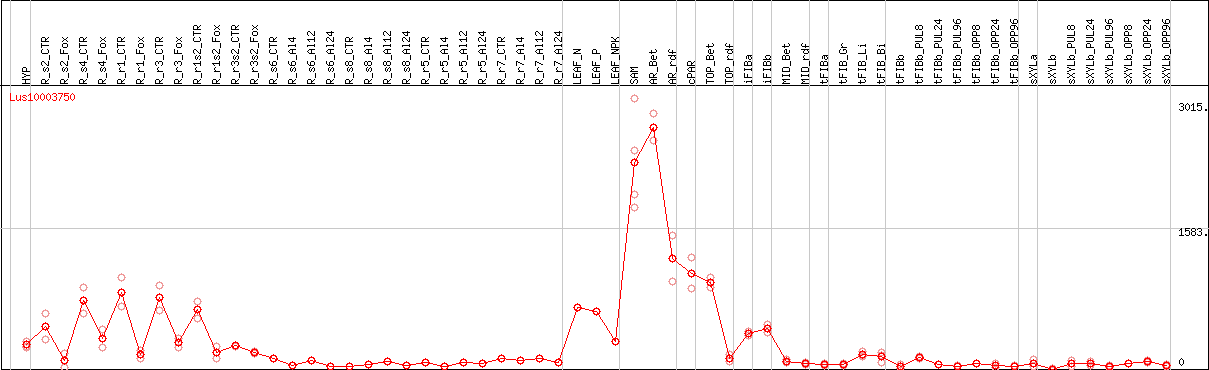
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10003750 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.