External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G03750 310 / 1e-104
SUVR3, SDG20
SET domain protein 20 (.1.2)
AT2G23740 104 / 2e-24
C2H2ZnF
AtCZS
nucleic acid binding;sequence-specific DNA binding transcription factors;zinc ion binding (.1.2)
AT3G04380 96 / 6e-22
SDG31, SUVR4
SET DOMAIN PROTEIN 31, SET-domain containing protein lysine methyltransferase family protein (.1.2)
AT1G04050 93 / 1e-20
SDG13, SUVR1
SET DOMAIN PROTEIN 13, homolog of SU(var)3-9 1 (.1)
AT2G35160 89 / 2e-19
SGD9, SUVH5
SET DOMAIN-CONTAINING PROTEIN 9, SU(VAR)3-9 homolog 5 (.1)
AT2G22740 87 / 8e-19
SDG23, SUVH6
SET DOMAIN PROTEIN 23, SU(VAR)3-9 homolog 6 (.1), SU(VAR)3-9 homolog 6 (.2)
AT1G76710 84 / 1e-17
ASHH1 ,SDG26
ASH1-RELATED PROTEIN 1, ASH1 RELATED PROTEIN 1, SET domain group 26 (.1.2)
AT1G73100 81 / 9e-17
SDG19, SUVH3
SET DOMAIN PROTEIN 19, SU(VAR)3-9 homolog 3 (.1)
AT5G43990 79 / 4e-16
SDG18, SUVR2
SET DOMAIN PROTEIN 18, SET-domain containing protein lysine methyltransferase family protein (.1.2.3.4.5)
AT5G04940 77 / 3e-15
SUVH1
SU(VAR)3-9 homolog 1 (.1), SU(VAR)3-9 homolog 1 (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013623
520 / 0
AT3G03750 320 / 3e-108
SET domain protein 20 (.1.2)
Lus10016788
103 / 6e-24
AT2G23740 1325 / 0.0
nucleic acid binding;sequence-specific DNA binding transcription factors;zinc ion binding (.1.2)
Lus10022483
102 / 1e-23
AT2G23740 1070 / 0.0
nucleic acid binding;sequence-specific DNA binding transcription factors;zinc ion binding (.1.2)
Lus10012513
99 / 9e-23
AT3G04380 419 / 2e-143
SET DOMAIN PROTEIN 31, SET-domain containing protein lysine methyltransferase family protein (.1.2)
Lus10022657
96 / 3e-22
AT3G04380 412 / 2e-142
SET DOMAIN PROTEIN 31, SET-domain containing protein lysine methyltransferase family protein (.1.2)
Lus10024946
97 / 7e-22
AT5G43990 488 / 7e-163
SET DOMAIN PROTEIN 18, SET-domain containing protein lysine methyltransferase family protein (.1.2.3.4.5)
Lus10011011
94 / 8e-21
AT2G35160 620 / 0.0
SET DOMAIN-CONTAINING PROTEIN 9, SU(VAR)3-9 homolog 5 (.1)
Lus10016600
93 / 1e-20
AT1G76710 600 / 0.0
ASH1-RELATED PROTEIN 1, ASH1 RELATED PROTEIN 1, SET domain group 26 (.1.2)
Lus10005476
92 / 1e-20
AT3G04380 503 / 3e-176
SET DOMAIN PROTEIN 31, SET-domain containing protein lysine methyltransferase family protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G064600
342 / 5e-117
AT3G03750 258 / 1e-83
SET domain protein 20 (.1.2)
Potri.005G129800
106 / 3e-25
AT2G23740 1468 / 0.0
nucleic acid binding;sequence-specific DNA binding transcription factors;zinc ion binding (.1.2)
Potri.007G033000
104 / 2e-24
AT2G23740 1436 / 0.0
nucleic acid binding;sequence-specific DNA binding transcription factors;zinc ion binding (.1.2)
Potri.005G195400
99 / 6e-23
AT1G76710 657 / 0.0
ASH1-RELATED PROTEIN 1, ASH1 RELATED PROTEIN 1, SET domain group 26 (.1.2)
Potri.002G257400
99 / 2e-22
AT5G43990 524 / 8e-176
SET DOMAIN PROTEIN 18, SET-domain containing protein lysine methyltransferase family protein (.1.2.3.4.5)
Potri.013G048100
97 / 4e-22
AT3G04380 471 / 5e-161
SET DOMAIN PROTEIN 31, SET-domain containing protein lysine methyltransferase family protein (.1.2)
Potri.014G192500
97 / 7e-22
AT5G43990 531 / 3e-178
SET DOMAIN PROTEIN 18, SET-domain containing protein lysine methyltransferase family protein (.1.2.3.4.5)
Potri.009G138600
94 / 6e-21
AT3G04380 427 / 2e-144
SET DOMAIN PROTEIN 31, SET-domain containing protein lysine methyltransferase family protein (.1.2)
Potri.003G083100
92 / 4e-20
AT2G35160 676 / 0.0
SET DOMAIN-CONTAINING PROTEIN 9, SU(VAR)3-9 homolog 5 (.1)
Potri.005G182100
86 / 3e-18
AT1G77300 807 / 0.0
LAZARUS 2, CAROTENOID CHLOROPLAST REGULATORY1, ASH1 HOMOLOG 2, histone methyltransferases(H3-K4 specific);histone methyltransferases(H3-K36 specific) (.1), histone methyltransferases(H3-K4 specific);histone methyltransferases(H3-K36 specific) (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF05033
Pre-SET
Pre-SET motif
PF00856
SET
SET domain
Representative CDS sequence
>Lus10003864 pacid=23163356 polypeptide=Lus10003864 locus=Lus10003864.g ID=Lus10003864.BGIv1.0 annot-version=v1.0
ATGCCAAAAATGGGGACGGAGAAGCACCGGAAGCAAAGCGACGGTGAAGAAGACCCAAATGCAACTCCGTTCATCAACTGCGCGGAGCTCATCCTCCCAT
GGCTAACCCCGCAAGAACTCGCCACCGTCGCCTCCACCTGCAAATACCTAGCACAACTCACCCGCATCATCACTCTCCATCGATCCTCCGACGCTTCGAG
ATCGTTGGAGAAACTTCCCATTCCTTTCCATAGCTCCGCGCAATCTCCCCGATACACCTATTTCCTATACTCCCCAACTCAGCTGCTGTCGTCGCAATCC
CCCGATCGACAGCCCTGGGGCTTCCCTAATTTCTACGCCGACTCACCGGCGCTCCATGCGACTGATAGCGGTTGCGGATGCGGTTGCGAAGGATGCGGAA
GAGGCGATGGACGTTTGGAATTGGAAGAGCTGGAATTAGGAGTGCTGAGCGAGTGTGGACCGAGTTGCGGGTGCGGGTTTGAATGCAGGAATCGGTTGAC
TCAAAGAGGGATTAGGGTTAAACTGAAGATCGTTTGGGATGAGAAGAAAGGCTGGGGTTTGTTCGCTGATCAGTTGATTCCACAATGCCAATTCATCTGC
GAGTATACCGGAGAACTCTTGACAACCGCCCAAGCAAGACGACGGCAACAGGTCTACGACGACCAACTCTCTTGTTCTTCTGCTCTTTTGGTTGTGAGAG
AGCATCTGCCGTCGGGAAAGGCTTGTTTGAGGGTGAACATTGATGCGACCAAGATTGGGAATGTGGGAAGATTCATAAACCATTCTTGTGATGGTGGTAA
CCTGTGTACAGCAACAGTGAGGAGCCCCGGAGCTTGCTTTCCTCGTCTCTGTTTCTTTGCTAGTAGAGACATTCAAGTTGGGGAAGAGCTCAGTTTTAGT
TATGGTGAAACTAGGGTGAGGTCTAATGGTCTGCAGTGCTTCTGTGACAGCTCTTCTTGCTCTGGAACTTTACCTTCAGAAGTGACTTAG
AA sequence
>Lus10003864 pacid=23163356 polypeptide=Lus10003864 locus=Lus10003864.g ID=Lus10003864.BGIv1.0 annot-version=v1.0
MPKMGTEKHRKQSDGEEDPNATPFINCAELILPWLTPQELATVASTCKYLAQLTRIITLHRSSDASRSLEKLPIPFHSSAQSPRYTYFLYSPTQLLSSQS
PDRQPWGFPNFYADSPALHATDSGCGCGCEGCGRGDGRLELEELELGVLSECGPSCGCGFECRNRLTQRGIRVKLKIVWDEKKGWGLFADQLIPQCQFIC
EYTGELLTTAQARRRQQVYDDQLSCSSALLVVREHLPSGKACLRVNIDATKIGNVGRFINHSCDGGNLCTATVRSPGACFPRLCFFASRDIQVGEELSFS
YGETRVRSNGLQCFCDSSSCSGTLPSEVT
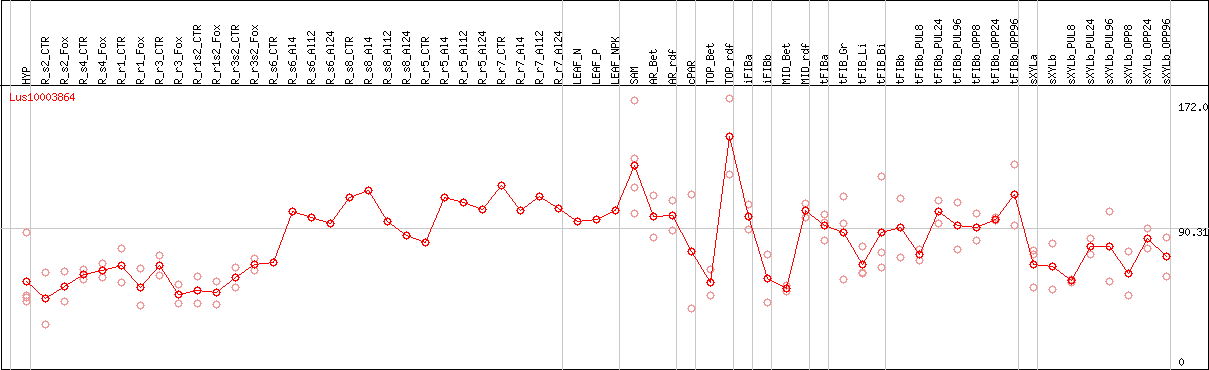
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10003864 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.