External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G02790 324 / 4e-113
GSTL3
Glutathione transferase L3, Glutathione S-transferase family protein (.1)
AT5G02780 305 / 1e-105
GSTL1
glutathione transferase lambda 1 (.1.2)
AT3G55040 288 / 3e-98
GSTL2
glutathione transferase lambda 2 (.1)
AT1G19570 70 / 2e-14
DHAR5, DHAR1, ATDHAR1
DEHYDROASCORBATE REDUCTASE 5, dehydroascorbate reductase (.1.2)
AT1G75270 59 / 3e-10
DHAR2
dehydroascorbate reductase 2 (.1)
AT1G53680 51 / 1e-07
ATGSTU28
glutathione S-transferase TAU 28 (.1)
AT1G78380 50 / 4e-07
GST8, ATGSTU19
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
AT2G02380 48 / 2e-06
ATGSTZ2
glutathione S-transferase (class zeta) 2 (.1)
AT1G78370 47 / 2e-06
ATGSTU20
glutathione S-transferase TAU 20 (.1)
AT2G02390 46 / 4e-06
GST18, ATGSTZ1
GLUTATHIONE S-TRANSFERASE 18, glutathione S-transferase zeta 1 (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015049
349 / 9e-123
AT5G02790 322 / 3e-112
Glutathione transferase L3, Glutathione S-transferase family protein (.1)
Lus10040347
291 / 6e-99
AT3G55040 290 / 8e-98
glutathione transferase lambda 2 (.1)
Lus10019895
274 / 1e-88
AT5G12130 399 / 6e-135
PIGMENT DEFECTIVE 149, TELLURITE RESISTANCE C, integral membrane TerC family protein (.1)
Lus10026199
60 / 5e-10
AT1G17160 496 / 8e-173
pfkB-like carbohydrate kinase family protein (.1.2)
Lus10042468
58 / 6e-10
AT1G78380 306 / 6e-107
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Lus10014022
59 / 7e-10
AT5G12130 402 / 7e-137
PIGMENT DEFECTIVE 149, TELLURITE RESISTANCE C, integral membrane TerC family protein (.1)
Lus10042469
56 / 2e-09
AT1G78380 284 / 5e-98
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Lus10042470
56 / 3e-09
AT1G78380 291 / 1e-100
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Lus10030362
55 / 8e-09
AT1G17170 210 / 7e-69
Arabidopsis thaliana Glutathione S-transferase \(class tau\) 24, glutathione S-transferase TAU 24 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.016G083500
364 / 5e-129
AT5G02790 340 / 2e-119
Glutathione transferase L3, Glutathione S-transferase family protein (.1)
Potri.006G133500
355 / 3e-125
AT5G02790 325 / 1e-113
Glutathione transferase L3, Glutathione S-transferase family protein (.1)
Potri.008G046800
303 / 5e-104
AT3G55040 318 / 4e-109
glutathione transferase lambda 2 (.1)
Potri.010G214800
100 / 4e-26
AT5G02780 98 / 6e-26
glutathione transferase lambda 1 (.1.2)
Potri.011G140400
62 / 2e-11
AT1G78380 308 / 9e-108
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Potri.001G436500
57 / 1e-09
AT1G78380 303 / 2e-105
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Potri.011G140800
55 / 5e-09
AT1G78380 318 / 1e-111
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Potri.001G437000
55 / 5e-09
AT1G78380 308 / 1e-107
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Potri.011G114000
54 / 8e-09
AT1G17180 265 / 2e-90
glutathione S-transferase TAU 25 (.1)
Potri.001G437200
53 / 3e-08
AT1G78380 308 / 2e-107
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0497
GST_C
PF00043
GST_C
Glutathione S-transferase, C-terminal domain
Representative CDS sequence
>Lus10003994 pacid=23157486 polypeptide=Lus10003994 locus=Lus10003994.g ID=Lus10003994.BGIv1.0 annot-version=v1.0
ATGGCTGCCACTGCTTTGGACCGAACCGTGCCGGAGAATTTGCCTCCTTCATTGGATTCCACGGCGGCACAGCCTCCTCTCTTCGACGGAACTACCAGGC
TGTACACGGCTTATATCTGTCCCTATGCTCAACGAGTGTGGATCACTAGGAATTTCAAGGGGTTGCAAAATGAGATAAAGTTGGTTCCTTTGAATCTTCA
GGACAGGCCAACTTGGTATAAAGACAAGGTTTATCCTCTCAATAAGGTCCCAGCCCTTGAACACAATGGAAAAATCATTGGGGAGAGTCTAGACTTGATT
AAATACTTGGATAGCAACTTTGAAGGACCTTCTCTTACTCCTGGTGATCGTGTTAAGAGACAGTTTGCTGATGAGTTGCTGTCATCCTGCGACACCTTTG
TTGGATCTGTGTATTCTTCTTTCAAGGGAGATCTGGCCACAGAAGTTGATCCTGTTTTTGATTATCTGGAAAATGCGCTTTCAAGGTTCCATGATGGGCC
CTTCTTCCTTGGCCAATTCAGTCTAGTGGACATAGCCTACATTCCTTTTATTGAAAGATTCCAGGTGTTCTTATCTGAAGTGTTCAAGTATGATATTACT
TCAGGCAGGCCTGGACTTGCAGCTTTGATTGAGGAGTGTAACAAAATTGAGGCTTACAAGCCAACAAAAGTTGATCCTAAGGAGACAGTTCTGAACTTGA
GCAAGCGTTTCCTGGTAACCGATGTTCATTTACTGTAA
AA sequence
>Lus10003994 pacid=23157486 polypeptide=Lus10003994 locus=Lus10003994.g ID=Lus10003994.BGIv1.0 annot-version=v1.0
MAATALDRTVPENLPPSLDSTAAQPPLFDGTTRLYTAYICPYAQRVWITRNFKGLQNEIKLVPLNLQDRPTWYKDKVYPLNKVPALEHNGKIIGESLDLI
KYLDSNFEGPSLTPGDRVKRQFADELLSSCDTFVGSVYSSFKGDLATEVDPVFDYLENALSRFHDGPFFLGQFSLVDIAYIPFIERFQVFLSEVFKYDIT
SGRPGLAALIEECNKIEAYKPTKVDPKETVLNLSKRFLVTDVHLL
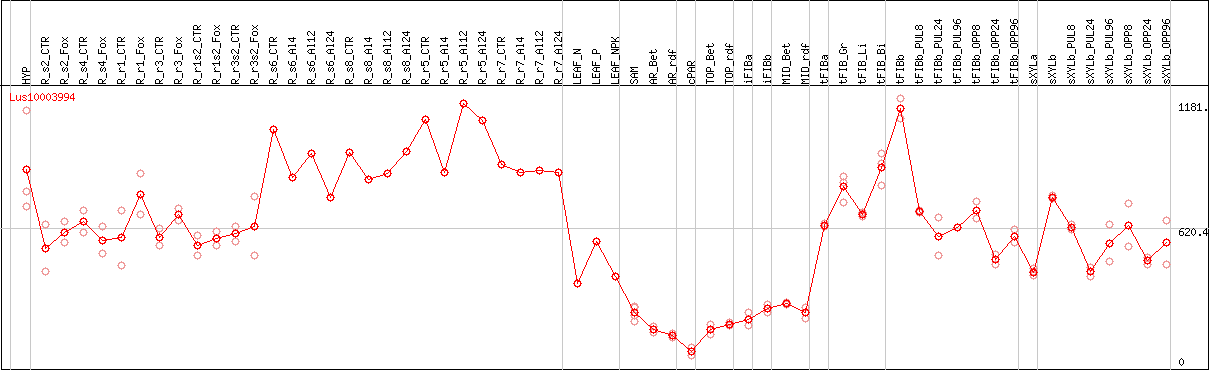
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10003994 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.