Lus10004009 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10004009 pacid=23153350 polypeptide=Lus10004009 locus=Lus10004009.g ID=Lus10004009.BGIv1.0 annot-version=v1.0
ATGGCGCCGAGTAAGAACAGCCGCGGTAAGGCTAAGGGAGAGAAGAAGAAGAAAGACGAGAAGATTCTGCCGACAGTAATGGATATCACCGTGAACCTTC
CCGACGATACTCATGTCATTCTCAAGGGAATATCAACGGACAGAATTATAGACGTTCGTCGTCTTTTGTCCGTCAACACGGAAACTTGCTCCGTCACTAA
TGTCTCCTTCACTCATGAGGTTAGAGGCCCGCGGTTGAAAGATACGGTCGACGTTACGGCTCTCAAGCCTTGCGTCCTCACTTTGATTGAAGAGAATTAC
GATGAAGGTCTTGCCGTCGCGCATGTCCGGCGGCTATTGGACGTCGTCGCCTGTACGACCTGGTTCGGTCCGTCGGACGCTGGCGGAAAGAATGGTGCCA
GCAAGAAGGCTGCTGCTGCTGCTGTTAAGTCTGAAGAAGGTCCTGCTGGCGGAGGAGGGAAGCAGCCGTCGCTTCAGGACGTGCCGGAGGAAGAAGGAGA
GCTGAGCCATTCTTGCCCTAAGCTTGGGCAATTTTATGACTTCTTCTCGCTGTCGCACCTCACTCCTCCTCTTCAGTTTATTAGGAAGGGGGATAAGCGG
CAGGTGGATGAGATTTCAGCTGAGGATCACCTTTTTTCACTGGATGTGAAGCTTTGCAATGGAAAGCTGGTCCAGGTGGAAGCTTGCCGAAAAGGATTCT
TTAGTGTAGGGAAACAGCGGATTCTTTGCCACAGTCTTGTGGACTTGTTGCGACAGCTTAGTAGAGCCTTTGACAATGCTTATAATGAGCTCATGAAAGC
ATTCTCAGACCGCAACAAGTTTGGGAATCTTCCTTATGGATTCAGAGCAAACACATGGCTTATCCCTCCTGCTGCTGCCCAATTTCCGTCATCTTTCCCG
CCTCTTCCTGTTGAGGATGAGAAGTGGGGAGGAAATGGAGGTGGTCTTGGTAGAGATGGGAAACAAGATCTGATACCTTGGGCTGATGAATTTTCATATC
TTGCGTCCATGCCTTGCAAGAATGCAGAGGAAAGGCAGGTCCGAGACAGGAGGGCGTTTCTTCTCCACAGCCTATTTGTTGACATTGCCATTTTCAGGGC
CATTAAAGCTGTACAGTTAGTAAATGAAAAGAAAGAGTTACCTGGTTCAATTAACAATTGTGAAGGTCTATACACAGAGAGAGTAGGGGATTTGATTATT
ACGATTGTGAAAGATGTTTCCAATGCTAGCTCAAAGATGGACACTAAGATCGATGGAATTCAAGCAGCCGGGGTAGACAAAAAAAATTTACTAGAAAGAA
ACTTGCTGAAGGGGATAACCGCAGATGAAAATACTGCTGCACATGATATTGCCACACTTGGGACTGTAAATGTGAGATACTGCGGTTATGTTGCTACTGT
AAAAGTTGAGGAAAGAAAGGAGAGTAACGTCACATCTCAAAGAGTTACACTTGAACAGCCCGACGGAGGAGCCAATGCCCTTAATATTAGCAGTTTGAGA
CTGTTACTTCACAAAACACCCAATTCAGAGCACAGCAAACTAGATTTGTCCTCACGAGCTACAGAACAGGAAGAGTTCAGTGCTTCCCAGGGTTTTGTAG
AGAGACTACTGGAAGAAAGTATTTTACAACTTGAAGAAGAGGAAGCAGAGCAAGTTCCTTTTGTACGGTGGGAGCTTGGCGCTTGTTGGATACAACATTT
GCAAGATCAGAAAAATGCAGATAAGGATAAGAAACCTTCTGATAAGAATAAGAAATCTTCCACAGAAAAAGAAATCAAGGTTGAGGGACTTGGTACGCCT
CTTAAGTCTCTCAAGGCAAAGAAAAAAACGGATGAAAGCGAAAAACCAAGAGTCTCTGCTAATGGTGTAGTCTGGGAACTCGAGGATTCTGCTTCTCCAA
TGTGTCAACTTGAAAGTTCTGCAGATGATGAGCTTTCGCTACAGAGGATTTTGTCGGAAACGGCTTTTGCTCGACTGAAAGAAACAGACACTGGACTTCA
TTCTAAGTCTTTGCCCGAATTGATTGACTTGTCTCAGAAATATTACACTGAGGTAGCTCTTCCAAAATTGGTTGCAGATTTTGGTTCGCTCGAACTTTCT
CCAGTTGATGGCCGAACTCTAACTGATTTTATGCACACTAGAGGTCTTCGTATGCGTTCACTGGGACGTGTTGTCAAGCTTTCAGAGAAGTTGTCACATG
TACAGTCGCTTTGCATTCATGAGATGATTGTACGAGCCTTTAAGCACATTATCCAGGCTGTCATTGCAGCTGTGGCTGATCATGATAAAATGGCTGTATC
CGTAGCTGCTGCTCTAAATCTCATGCTCGGAGTACCTGAAAATCAAGATTCCAAGCCTTCCCATGTCCATTCTCTTGTATGGAGATGGCTAGTGTTATTT
CTAAGAAAACGATTTGATTGGGAACTTAGCATCCCAAACTTCAAAGATGTCCGAAAATTTGCTTTGCTTCGTGGATTGTGCCACAAGGTGGGTATTGAGT
TGGTTCCGAGGGATTTTGACATGGATTCTCCATACCCATTCCAGAAATCAGATGTTGTCAGCCTTGTCCCAGTGCATAAGCAAGCAGCGTGTTCCTCTGC
AGATGGAAGGCAGCTTTTGGAATCATCAAAAACAGCCTTAGATAAAGGAAAACTTGAAGATGCCGTAACATATGGTACAAAGGCACTTGCAAAGCTGGTA
GCTGTTTGCGGCCCTTACCATAGAATGACTGCAGGAGCTTACAGCCTTCTTGCGGTTGTACTATATCACACTGGCGACTTCAACCAGGCCACTATATATC
AGCAGAAAGCACTTGATATCAATGAGAGAGAATTAGGACTTGACCACCCAGATACCATGAAGAGTTATGGAGATCTTGCTGTCTTCTATTACCGACTCCA
ACATACTGAGCTGGCTCTTAAGTATGTGAAGCGTGCCCTGTACCTGTTGCATCTGACATGTGGTCCATCACACCCAAATACTGCTGCGACATACATAAAT
GTGGCTATGATGGAGGAAGGGCTGGGAAATGTGCATGTTGCACTTAGATATCTTCATAAGGCTCTAAAGTGCAACCAAAGGTTGCTTGGCCCAGACCACA
TTCAGACGGCAGCAAGTTACCATGCAATTGCAATCGCACTATCATTGATGGAAGCATACCCCTTGAGCGTACAGCATGAACAAACAACTCTACAAATTCT
CCGAGCAAAGCTGGGTCCAGATGATTTGCGTACTCAGGATGCTGCTGCTTGGCTTGAGTACTTTGAGTCAAAAGCATTTGAGCAGCAAGAAGTTGCAAGG
AATGGTACAAAAAAGCCAGATGCATCAATTGCTAGCAAGGGGCATCTGAGTGTATCTGATTTACTAGATTATATCAATCCGAGCCAGGATCCCAAGGGGA
GAGAAGCAGCAGCTAAAAGAAAAAGCTACTTGACAAAGGTAAAGGGAAAAGCTGAACCTGAACCCATTGTTCCAAGTTTTCCTGAATCTCCAAAAGAGAA
AGTCGAAGAAGCAGCTGATGAAGAGTCAGTTATTCCTATCGTCAAAGAAGAACTCAGTTTGATGGAGAATGAGAATGAACAACCTGACTCAGAAGGGTCT
ATAGATGAAAAATCAGTTATGAAAGAGGAAGTATTACATGAAACACATGCTGAAGGAGATGATGGATGGCAACCGGTTCAAAGACCACGATCAGCTGGCT
CTAATGGTCGACGGAGGAAACAACATCGGGGTATCATTGGCAAAGTATATGTTCATACAAAAAAGGTTACGGATGCTGAAGTGGATTATGCTTCGCCCAA
GAACACCCTTCAGAGTACTAAGCATTACATTGTAAAGAAACGACCCATCTCTCAGGGAACTTATGCAGACCAACAAGGTGCACTTCCTTCTCAGCCAGCC
AAGTATGGGCGGAGGATCACAAAAACCGTAGCATATCGAGTAAAATCTGTTCCTTCGCCTGATAAAGATGCTGCTGCCTCCGAGACTTCTGAACAAGTGG
GCAACGTTTTCTCCTCTGCAACAGAACCTGTTACAATTTCTGCTCCAAGTGAGGGTAACTCGTTAAAGAATCAAATGGTTAGCTTGGGAAAGTCCCCATC
GTACAAGGAAGTCGCATTGGCGCCACCTGGCACTATCTGGTCACCTGTTTCGGATATTGGAGATAAACCAGTTACTGGTGTTCATAACCATGAAGAGACA
CAGGAAGCTAGGGAAATTGCTATTCCAGCTGTATTCGAGAAGAACACAGAAAATGCATCAATCGACTTGGTAGAAGATAGTTTGACGACACCCTCCAGCG
TGCAAGAATCCAAGTCTACTCTTTTAGATCATGAAGAAAACGACATATTGACAGATGGTTCAATCGACATACCCGAAGCATTCCAGAATAATCAGAGTTC
TGCGTCAGATTTCAGACAACATTCAGAAGATGAAATTGTTATCCTTGAAAATATCCTTGAAAACAAAGAAGAAAGCCACCCTATCAGTGAAAGAGAAGGC
TCAGGTCTTGAAGGTAATTCTGATTCCTCATTTCTGGGAGAAGATCTGAAGGAAGTAGCATCTGTACCTAATAAGAAGCTATCTGCCTCTGCTGCTCCAT
TTAACCCTTCGCCATCTGTTATCCGACCCGTACCACTCCCTCTGAACATTCCTCTTCCTTCCGGTCCCGGTCCGGTTCAAGCCGTTGCACCCTGGCCAGT
AACCCTTCACCCGGTGACTGGAACTGTCATATCGACTGTTAGTCCACTCCCCTCTCCCCACCACCCGTTTCCATCGCCTCCTCATACCCCGAACATGATG
CGACCACTGTCCTTCATGTACCCACCTTACAGTCAAGCCCAGGGAATCCCACCAAGTACATATCCCATCACCAGCACTGCATTTCATCCAAGCCATTACT
CTTGGCAGTGCAATGTGAACAGAAATGTTCCGGAGTTTGTTCCTACCACAATGTGGCCTGGTTGTCACATGGTGGAGTTCCCTCCTGCCCCTTCACCTGT
TATGGAGCCCGTTTCTGATCCTGTACCAGACCATAATGAGACAATGGAGTGTCCTGACTCGTTCCTGCCAGCAGATATTGTCAATGTAACTGATTCTGAC
AAGGACGTGAGTCTTCCGGTGGAAGAAAATGGTCGTTTGAACTTGTCCGGTGCCGACACTTCTACTAGCAACAGCATGAAAGAGAACTCGGCTGAGGGGG
AGAGAACTTTCAGTATTTTGATAAGGGGAAGGAGAAACCGAAAGCAGACATTGAGAATGCCAATAAGTTTGCTCAATAAGCCATATGGCTCGCAGTCTTT
CAAAGTTGTTTACAACAGAGTAGTGAGAGGTAGTGAATCTCCAAAGCAGACTGCCGTTTCTTCAACTGGTTAG
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10004009 pacid=23153350 polypeptide=Lus10004009 locus=Lus10004009.g ID=Lus10004009.BGIv1.0 annot-version=v1.0
MAPSKNSRGKAKGEKKKKDEKILPTVMDITVNLPDDTHVILKGISTDRIIDVRRLLSVNTETCSVTNVSFTHEVRGPRLKDTVDVTALKPCVLTLIEENY
DEGLAVAHVRRLLDVVACTTWFGPSDAGGKNGASKKAAAAAVKSEEGPAGGGGKQPSLQDVPEEEGELSHSCPKLGQFYDFFSLSHLTPPLQFIRKGDKR
QVDEISAEDHLFSLDVKLCNGKLVQVEACRKGFFSVGKQRILCHSLVDLLRQLSRAFDNAYNELMKAFSDRNKFGNLPYGFRANTWLIPPAAAQFPSSFP
PLPVEDEKWGGNGGGLGRDGKQDLIPWADEFSYLASMPCKNAEERQVRDRRAFLLHSLFVDIAIFRAIKAVQLVNEKKELPGSINNCEGLYTERVGDLII
TIVKDVSNASSKMDTKIDGIQAAGVDKKNLLERNLLKGITADENTAAHDIATLGTVNVRYCGYVATVKVEERKESNVTSQRVTLEQPDGGANALNISSLR
LLLHKTPNSEHSKLDLSSRATEQEEFSASQGFVERLLEESILQLEEEEAEQVPFVRWELGACWIQHLQDQKNADKDKKPSDKNKKSSTEKEIKVEGLGTP
LKSLKAKKKTDESEKPRVSANGVVWELEDSASPMCQLESSADDELSLQRILSETAFARLKETDTGLHSKSLPELIDLSQKYYTEVALPKLVADFGSLELS
PVDGRTLTDFMHTRGLRMRSLGRVVKLSEKLSHVQSLCIHEMIVRAFKHIIQAVIAAVADHDKMAVSVAAALNLMLGVPENQDSKPSHVHSLVWRWLVLF
LRKRFDWELSIPNFKDVRKFALLRGLCHKVGIELVPRDFDMDSPYPFQKSDVVSLVPVHKQAACSSADGRQLLESSKTALDKGKLEDAVTYGTKALAKLV
AVCGPYHRMTAGAYSLLAVVLYHTGDFNQATIYQQKALDINERELGLDHPDTMKSYGDLAVFYYRLQHTELALKYVKRALYLLHLTCGPSHPNTAATYIN
VAMMEEGLGNVHVALRYLHKALKCNQRLLGPDHIQTAASYHAIAIALSLMEAYPLSVQHEQTTLQILRAKLGPDDLRTQDAAAWLEYFESKAFEQQEVAR
NGTKKPDASIASKGHLSVSDLLDYINPSQDPKGREAAAKRKSYLTKVKGKAEPEPIVPSFPESPKEKVEEAADEESVIPIVKEELSLMENENEQPDSEGS
IDEKSVMKEEVLHETHAEGDDGWQPVQRPRSAGSNGRRRKQHRGIIGKVYVHTKKVTDAEVDYASPKNTLQSTKHYIVKKRPISQGTYADQQGALPSQPA
KYGRRITKTVAYRVKSVPSPDKDAAASETSEQVGNVFSSATEPVTISAPSEGNSLKNQMVSLGKSPSYKEVALAPPGTIWSPVSDIGDKPVTGVHNHEET
QEAREIAIPAVFEKNTENASIDLVEDSLTTPSSVQESKSTLLDHEENDILTDGSIDIPEAFQNNQSSASDFRQHSEDEIVILENILENKEESHPISEREG
SGLEGNSDSSFLGEDLKEVASVPNKKLSASAAPFNPSPSVIRPVPLPLNIPLPSGPGPVQAVAPWPVTLHPVTGTVISTVSPLPSPHHPFPSPPHTPNMM
RPLSFMYPPYSQAQGIPPSTYPITSTAFHPSHYSWQCNVNRNVPEFVPTTMWPGCHMVEFPPAPSPVMEPVSDPVPDHNETMECPDSFLPADIVNVTDSD
KDVSLPVEENGRLNLSGADTSTSNSMKENSAEGERTFSILIRGRRNRKQTLRMPISLLNKPYGSQSFKVVYNRVVRGSESPKQTAVSSTG
|
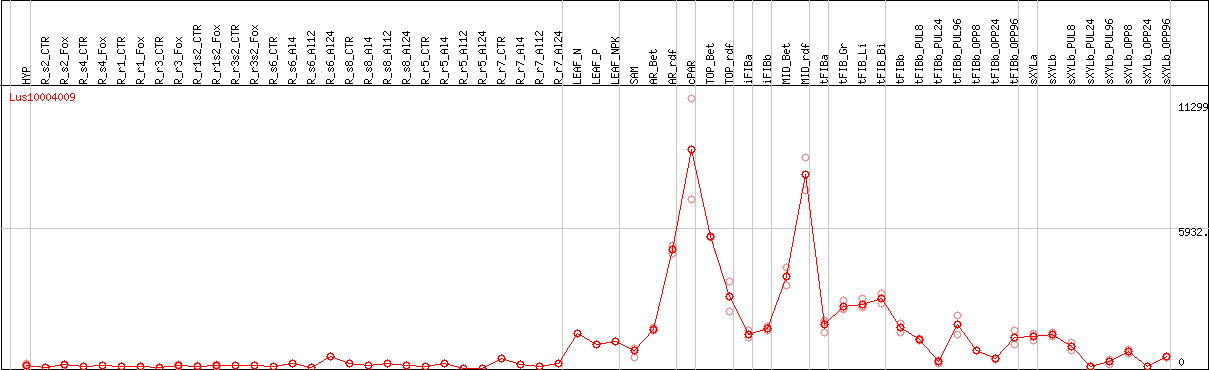
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10004009 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
